Abstract
p55PIK, a regulatory subunit of phosphatidylinositol 3-kinases, promotes cell cycle progression by interacting with cell cycle modulators such as retinoblastoma protein (Rb) via its unique amino-terminal 24 amino-acid residue (N24). Overexpression of N24 specifically inhibits these interactions and leads to cell cycle arrest. Herein, we describe the generation of a fusion protein (Tat transactivator protein (TAT)–N24) that contains the protein transduction domain and N24, and examined its effects on the proliferation and differentiation of leukemia cells. TAT–N24 not only blocks cell proliferation but remarkably induces differentiation of leukemia cells in vitro and in vivo. Systemically administered TAT–N24 also significantly decreases growth of leukemia cell tumors in animal models. Furthermore, overexpression of p55PIK in leukemia cells leads to increased proliferation; however, TAT–N24 blocks this effect and concomitantly induces differentiation. There is significant upregulation of p55PIK mRNA and protein expression in leukemia cells from patients. TAT–N24 inhibits cell cycle progression and induces differentiation of bone marrow cells derived from patients with several different types of leukemia. These results show that cell-permeable N24 peptide induces leukemia cell differentiation and suggest that p55PIK may be a novel drug target for the treatment of hematopoetic malignancies.
Similar content being viewed by others
Main
Class IA phosphatidylinositol 3-kinases (PI3Ks), composed of p110 catalytic and regulatory subunits, are involved in many important physiological and pathological processes.1 Three genes encode the p110 catalytic subunits and there are three genes encoding regulatory subunits (PIK3R1, 2 and 3) that give rise to at least six products: p85α, p55α, p50α, p85β, p55PIK (p55γ) and p50PIK (p50γ). The three p110 catalytic subunit isoforms can interact with each of the regulatory subunits, suggesting that different regulatory subunits may mediate signaling of specific PI3K pathways.1, 2 However, the mechanisms for specific pathway signaling by regulatory subunits are not fully understood.
PI3K-mediated signaling is critical for tumor growth and progression. In human cancers, genetic dysregulation of its signaling components can occur at several levels, including deletion and mutation of PTEN, which negatively regulates PI3K-mediated pathways as well as gene amplification of PIK3CA (encoding p110α) and Akt, a major downstream effector.1 Additionally, somatic mutations of PIK3CA have been reported in more than 25% of colorectal gastric, breast and brain tumors, making it one of the most commonly mutated oncogenes that occur in human cancer.3 As PI3K signaling is important for tumor growth and progression, and is activated in many cancers, the PI3Ks have become important targets for the development of new anticancer drugs and several PI3K catalytic subunit inhibitors have been developed and shown promise against a variety of cancers.4 However, these inhibitors can lead to serious side effects by their broad blockade on all PI3K signaling pathways and thereby limit their clinical use.
Previously, we reported that the p55PIK subunit of PI3K is associated with the important cell cycle regulator, retinoblastoma protein (Rb), via the amino-terminal 24 amino-acid residues (N24) of p55PIK.5 The p55PIK interaction with other proteins appeared to be essential for cell proliferation. In support of this notion, ectopically expressed N24 peptide, which competes with endogenous p55PIK binding to Rb, inhibited cell proliferation in several cancer cell lines.5 Recently, studies showed increased p55PIK mRNA expression and protein levels in majority of ovarian cancer. Knockdown of p55PIK proteins by siRNA leaded to decreased cell proliferation in ovarian cancer cell lines.6
To examine N24 as a potential cancer therapy, we developed an N24 expression system in replication-deficient adenovirus (Ad–N24), which inhibited cell proliferation and tumor growth of colon and thyroid cancers in vivo. Moreover, Ad–N24 also inhibited the growth of cancer cells, which do not express Rb, indicating that N24 inhibits tumor growth by Rb-independent mechanisms as well.7 While Ad–N24 is effective when injected locally into tumors, inefficient systemic delivery and safety concerns limit Ad–N24’s clinical utility. Also, due to lack of adenovirus receptors, most leukemia cells are not permissive to Ad–N24. Recent identification of the protein transduction domains (PTDs), which facilitate cell penetration of peptides, provides a potential strategy to overcome this problem.8 Fusion proteins containing PTDs have been used in cell culture systems; however, there are only a few examples of their application in vivo.9, 10 Given these issues, we decided to design a fusion protein composed of the PTD from HIV Tat transactivator protein (TAT) and N24 (TAT–N24) to investigate the effects of N24 on the proliferation and differentiation of leukemia cells and whether systemic administration of TAT–N24 could inhibit leukemia proliferation and growth in vivo.
Results
Generation and purification of TAT–N24
We generated TAT–N24 by cloning cDNA encoding N24 and TAT into a bacterial expression vector. This expression vector encoded 6xHis and hemagglutinin (HA) tags to enhance purification and detection of TAT–N24. We also generated a control fusion protein (TAT–N24M), in which the cDNA encoding sequence for N24 was replaced by the cDNA encoding the shuffled N24 in the same expression vector. Expression and affinity-purification of TAT–N24 and TAT–N24M control proteins were performed according to standard protocols under denaturing conditions (6 M urea). Purified TAT–N24 protein was dialyzed against PBS prior to its use. The uptake and accumulation of TAT–N24 protein in cultured cells were confirmed by western blotting and immunofluorescence staining using anti-HA antibody (Supplementary data 1).
Effects of TAT–N24 on the proliferation and differentiation of human leukemia HL60 and K562 cells
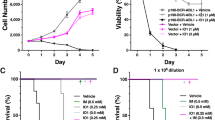
We examined the effects of purified TAT–N24 and TAT–N24M on cellular proliferation of HL60 cancer cells by counting cell number daily after peptide addition (Figure 1a). After 4 days, TAT–N24 (120 μg/ml) decreased cell number by 45% compared with TAT–N24M (4.1 × 104 versus 7.5 × 105 cells). TAT–N24M (120 μg/ml) had no effect on cell number compared with vehicle alone. TAT–N24 also induced cell cycle arrest at G0/G1 transition and significantly decreased DNA synthesis (Figures 1b and c). These data show that TAT–N24 protein was still bioactive and inhibited cell proliferation even when purified under denaturing conditions.
Effects of TAT–N24 on cell proliferation of HL60 cells. (a) TAT–N24 effects on cell proliferation. TAT–N24 or TAT–N24M (120 μg/ml) was added to HL60 cells in culture and cell number was counted daily for the next 1–4 days. The mean±S.D. of triplicate samples are shown. (b) TAT–N24 effects on cell cycle progression in HL60 cells. Cells were collected 48 h after incubation with TAT–N24 or TAT–N24M (120 μg/ml), PBS solvent was added to the control cells and cell cycle analyses were performed as described.7 Shown are representative histograms from an individual experiment. Similar results were obtained in three independent experiments. (c) Effects of TAT–N24 on DNA synthesis. Cells were incubated with TAT–N24 (120 μg/ml), TAT–N24M (120 μg/ml) or PBS (Control) for 48 h. BrdU was added 3 h prior to harvest of cells. Incorporation of BrdU into DNA and the DNA content in nuclei was determined by flow cytometry analysis.7 Shown here are the representative data from three independent experiments. Note that % cells in DNA synthesis is defined as BrdU-positive cells (R2)/total cells. BrdU intensity per cells in the R2 area is shown in arbitrary units with the mean of solvent-treated cells (Control) defined as 1
As TAT–N24 induces cell cycle arrest and blocks cell proliferation, we next investigated whether TAT–N24 could affect cell differentiation after cell cycle arrest. To address this issue, we studied the effects of TAT–N24 on differentiation of HL60 leukemia cells, which have been extensively studied as an experimental model to elucidate differentiation pathways.11 As all-trans retinoic acid (ATRA) induces HL60 cells to mature and express granulocytic markers such as CD11b,12, 13 it was used as a positive control in our experiments. As expected, ATRA increased the number of cells expressing CD11b (17.3% ATRA treated versus 4.8% in the control; Figure 2a). Surprisingly, TAT–N24 (120 μg/ml) also increased the number of cells expressing Cd11b (26.2% in the TAT–N24 treated versus 5.4% in the TAT–N24M treated), suggesting that TAT–N24 not only inhibited cell proliferation, but also induced differentiation. Quantitative real-time RT-PCR also showed that TAT–N24 increased CD11b mRNA expression in HL60 cells by 6.8-fold (data not shown).
Effects of TAT–N24 on expression of differentiation markers Cd11b and Cd14 in HL60 cells. (a) Effects on the expression of Cd11b in HL60 cells. TAT–N24 or TAT–N24M (120 μg/ml), ATRA (1 μg/ml) or PBS alone (Control) was added and the cells were grown in culture for 3 days. Immunostaining with FITC-labeled anti-Cd11b antibody was performed as described.26 Shown are representative histograms from an individual experiment. Number shown in each box is the percentage of cells with Cd11b+ expression (M1). Similar results were obtained in three independent experiments. (b) TAT–N24 effects on the expression of Cd14 in HL60 cells. The cells were incubated with TAT–N24 (120 μg/ml), TAT–N24M (120 μg/ml), ATRA (1 μg/ml) or PBS (Control) for 3 days. Immunostaining with PE-labeled anti-Cd14 antibody was done as described.26 Shown are representative histograms from an individual experiment. Number shown is the percentage of cells with Cd14+ expression. Similar results were seen in at least three independent experiments. (c) TAT–N24 effects on the expression of Cd11b and Cd14 in HL60 cells. The cells were incubated with TAT–N24 (120 μg/ml), TAT–N24M (120 μg/ml), ATRA (1 μg/ml), TAT–N24 (120 μg/ml) and ATRA (1 μg/ml) (TAT-N24+ATRA) or PBS (Control) for 3 days. Immunostaining with FITC-labeled anti-Cd11b and PE-labeled anti-Cd14 antibodies were performed as described.26 Shown are representative histograms from an individual experiment. Number shown is the percentage of cells with Cd11b+ (event numbers in upper right and upper left area/total events) and Cd14+ expression (event numbers in upper right and lower right areas/total events). Similar results were seen in at least three independent experiments. (d) TAT–N24 effects on the expression of cell lineage markers in HL60 cells. The cells were incubated with TAT–N24 (120 μg/ml), TAT–N24M (120 μg/ml), ATRA (1 μg/ml) or PBS (Control) for 3 days. mRNA levels of CCL2, CCR, CD163 and CD38 are quantified by real-time reverse transcription-PCR. Shown is summarized result from three independent experiments (mean±S.D., *P<0.05 compared with the mRNA level in cells treated with PBS defined as 1)
It is reported that ATRA treatment induced the differentiation of HL60 into granulocyte lineage, and phorbol myristate acetate (PMA) induces the differentiation of HL60 into monocyte-like cells, which expresses CD14 differentiation marker in HL60 cells.14 To explore which mature lineage of differentiation cells were induced by TAT–N24, we next examined the expression of Cd14 in HL60 cells treated with TAT–N24, TAT–N24M or ATRA for 3 days. ATRA slightly increased the expression of Cd14 (10.6% ATRA treated versus 6.8% in the control). In contrast, TAT–N24 strongly induced the expression of Cd14 (46.5% TAT–N24 treated versus 7.3% TAT–N24M treated) in HL60 cells (Figure 2b). These findings suggest that different signaling pathways may be activated by ATRA and N24 to regulate the differentiation of HL60 cells.14 Interestingly, the combination of ATRA and TAT–N24 synergistically induced Cd11b expression (70.4% ATRA+TAT-N24-treated group versus 18.1% TAT–N24 alone and 31.7% ATRA alone) but did not further increase the expression of Cd14 (37.2% in ATRA+TAT-N24-treated group versus 32.5% in TAT–N24-treated group) in HL60 cells (Figure 2c), suggesting that signaling pathways mediated by TAT–N24 in the differentiation of HL60 did not overlap with the signaling pathways by ATRA. To examine which differentiation cell lineage was induced by TAT–N24 in HL60, we have determined the expression of more cell lineage markers (CCL2, CCR and CD163 for monocyte, CD38 for granulocyte lineage) in TAT–N24 and ATRA-treated HL60 cells. Data shown in Figure 2d indicated that TAT–N24 induced HL60 to differentiate into monocyte lineage.
Next, we examined the effects of TAT–N24 on K562, a cell line derived from a chronic myeloid leukemia (CML) patient, which has been used as a model for megakaryocytic differentiation.15 PMA induces K562 cells to differentiate in a manner that resembles the cytological and biochemical events observed during megakaryocytic differentiation.15 As expected, TAT–N24 (120 μg/ml) blocked the cell proliferation by inhibiting the DNA synthesis of K562 cells (Figure 3a). We also observed significant megakaryocyte-like morphological changes in TAT–N24-treated cells (33.8% cells treated with TAT–N24 versus 3% cells with TAT–N24M), indicating that TAT–N24 induced differentiation of K562 cells (Figure 3b). The expression of Toll-like receptors 2 and 4 (TLR2/4), which are markers for K562 differentiation, also increased in a dose-dependent manner when K562 cells were treated with TAT–N24 for 3 days (Figure 3c). Similarly, TAT–N24 increased TLR2 and TLR4 mRNA expression as measured by quantitative real-time RT-PCR assays (data not shown). Furthermore, we have examined the expression of more megakaryocytic differentiation markers such as CD41 and CD61, and data showed that the expression of CD41 and CD61 significantly increased when K562 cells were treated with TAT–N24, suggesting that TAT–N24 treatment in K562 lead to differentiate into megakaryocyte lineage cells (data not shown).
Effects of TAT–N24 on cell proliferation and differentiation of K562 cells. (a) Effects of TAT–N24 on DNA synthesis. Cells were incubated with TAT–N24 (120 μg/ml), TAT–N24M (120 μg/ml), PMA (20 ng/ml) or PBS (Control) for 48 h. Incorporation of BrdU into DNA and the DNA content in nuclei was determined.7 Shown here are the representative data from three independent experiments. Note that % cells in DNA synthesis is defined as BrdU-positive cells/total cells. BrdU intensity per cells in the R2 area is shown in arbitrary units. (b) TAT–N24 effects on morphological changes in K562 cells. TAT–N24 (120 μg/ml) or TAT–N24M (120 μg/ml), PMA (20 ng/ml) or PBS (Control) were added to cultured K562 cells for 3 days. Cell morphology was evaluated by microscopy and the % cells with megakariocytic morphology was calculated by counting 400 cells (mean±S.D. of triplicate samples is shown). (c) TAT–N24 effects on the expression of TLR2 and TLR4 proteins in K562 cells. K562 cells were incubated with various amount of TAT–N24 or TAT–N24M (120 μg/ml), PMA (20 ng/ml) or PBS (Control) for 3 days. Cellular lysates were prepared and the expression of TLR2, TLR4, Akt, GAPDH in lysates was analyzed. Similar results were seen in at least three independent experiments. (d) TAT–N24 effects on the DNA synthesis in K562 cells overexpressing p55PIK. K562 cells were infected with adenovirus expressing p55PIK and GFP (Ad-p55PIK) or GFP alone (Ad-GFP). After 24 h infection, the cells were treated with TAT–N24 (120 μg/ml), TAT–N24M (120 μg/ml) or solvent for 48 h and BrdU incorporation assay was performed. Shown are the percentages of cells with significant BrdU incorporation (R2). (e) TAT–N24 effects on the expression of differentiation markers in K562 cells overexpressing p55PIK. Cells were infected with adenovirus expressing p55PIK and GFP (Ad-p55PIK) or GFP alone (Ad-GFP). After 24 h infection, the cells were treated with TAT–N24 (120 μg/ml), TAT–N24M or solvent for 72 h. Cell lysates were collected and the protein levels of TLR2, TLR4, p55PIK and GAPDH was determined
Increased cell proliferation in K562 cells overexpressing 55PIK and TAT–N24 blockade of the effects induced by overexpressing p55PIK in K562 cells
To further examine whether p55PIK plays a critical role in cell proliferation, we constructed an adenovirus that overexpressed p55PIK (Ad-p55PIK) in K562 cells as these cells are permissive for adenovirus infection (Figure 3d, data not shown).16
After infection of K562 with Ad-p55PIK, more than 95% cells achieved increased expression of p55PIK determined by the coexpression of GFP, as the cloning vector for Ad-p55PIK, pTrack-CMV-EGFP from Clontech (Mountain View, CA, USA), included coding sequence of GFP under the control of CMV promoter (data not shown).
Overexpression of p55PIK modestly increased DNA synthesis in K562 cells (66.4% positive for DNA synthesis in Ad-p55PIK-infected K562 cells versus 50.9% positive in Ad-GFP-infected control cells, Figure 3d), whereas TAT–N24 inhibited DNA synthesis, even in the cells overexpression of p55PIK, (39.4% positive in DNA synthesis in TAT-N24-treated cells versus 63.0% in TAT-N24M-treated cells, Figure 3d). Overexpression of p55PIK did not change the expression of TLR2 and TLR4 in K562 cells; however, TAT–N24 (120 μg/ml) still was able to induce TLR2 and TLR4 proteins expression, albeit to a lesser degree than it did in untransformed K562 cells (Figure 3e). These data provide strong evidence to show that the TAT–N24 directly targets p55PIK, and the p55PIK signaling pathway may play major roles in the development and progression of leukemia.
TAT–N24 effects on tumor xenograft growth of HL60 and K562 cells
We examined the effects of TAT–N24 peptide on the tumor growth of HL60 cells in athymic nude mice. HL60 cells were injected into mice s.c. and then 200 μl lipid emulsions containing 2 mg TAT–N24 or TAT–N24M were injected i.v. into mouse tail veins. TAT–N24 and TAT–N24M injections were administrated as described in the Materials and methods section, and the tumor xenograft growth and tumor weights were monitored. TAT–N24 decreased the tumor growth in mice and the mean xenograft tumor weight decreased by 54% in mice receiving TAT–N24 compared with mice receiving TAT–N24M (Figures 4a and b). Of note, we found significant uptake of both TAT–N24 and TAT–N24M into xenograft tumors suggesting that inhibition of tumor growth is likely mediated by TAT–N24 (Figure 4c). Additionally, TAT–N24 significantly decreased DNA synthesis in xenograft tumors (Figure 4c). Consistent with results from cell culture, TAT–N24 induced the expression of Cd11b in tumor xenografts (Figure 4c). These data showed that TAT–N24 decreased cell proliferation and induced differentiation of HL60 cells in vivo. Importantly, mice receiving TAT–N24 did not exhibit significant changes in appearance or weight, nor in serum glucose levels or hepatic enzyme activities (bilirubin, albumin, alkaline phosphatase, serum glutamic oxaloacetic transaminase and serum glutamic pyruvic transaminase; data not shown).
Effects of systemically delivered TAT–N24 on growth of HL60 tumor xenografts in athymic nude mice. (a) Time course comparing HL60 tumor xenograft growth after i.v. injection of TAT–N24, TAT–N24M or solvent control. Mice inoculated with HL60 cells (107/site) were divided into three groups receiving either 200 μl solvent–lipid emulsion, TAT–N24 (10 mg/ml emulsion) or TAT–N24M (10 mg/ml emulsion) via tail vein. The injection was repeated every 2 days. The size of tumors were measured (mean±S.D., n=5, *P<0.05, TAT-N24-treated group compared with TAT-N24M-treated group). (b) TAT–N24 effects on the xenograft tumor weight in vivo. Shown are representative tumor xenografts and mean tumor weights after necropsy (mean±S.D., n=5, **P<0.01, TAT-N24-treated group compared with TAT–N24M group). (c) Uptake of TAT–N24 or TAT–N24M and their effects on DNA synthesis and expression of Cd11b in tumor xenografts. The uptake of TAT–N24 and TAT–N24M was determined by immunohistochemistry analysis using anti-HA antibodies.26 BrdU incorporation into DNA (DNA synthesis) in tumor samples was determined as described.7 Shown here are representative images of slides. Expression of Cd11b in tumor samples was determined with PE-labeled anti-Cd11b antibodies (red), and nuclear DNA was stained with DAPI (green) as described.26 Shown here are representative images of slides
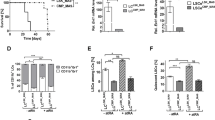
We then examined the effects of TAT–N24 emulsion on the tumor growth of K562 cells in nude mice. Results indicated that injection of TAT–N24 inhibited the growth of K562 tumor xenografts and induced the expression of differentiation markers such as TLR2 and TLR4 in tumor xenografts (Figures 5a and b). Imatinib (IM) is a selective inhibitor of BCR/ABL tyrosine kinase activity and a drug currently used for CML treatment.17 Although IM inhibited the growth of K562 tumor xenografts, it did not induce the expression of differentiation markers TLR2 and TLR4 (Figures 5a and b), consistent with the notion that TAT–N24 inhibition of tumor growth involves both the inhibition of cell proliferation and the induction of tumor cell differentiation.
Effects of systemically delivered TAT–N24 on growth of K562 tumor xenografts in nude mice. (a) TAT–N24 effects on the tumor xenograft weight. Mice inoculated with K562 cells (107/site) received 200 μl solvent–lipid emulsion (Control), TAT–N24 (10 mg/ml emulsion), TAT–N24M (10 mg/ml emulsion), IM (20 mg/kg per day) or TAT-N24+IM (IM 20 mg/ml per day, 2 mg TAT–N24 peptide) via tail vein or i.p. (for IM). Shown are mean tumor weights (mean±S.D., n=5, **P<0.01, *P<0.05, compared with control group). BrdU incorporation into DNA (DNA synthesis) in tumor samples was determined as described.7 Shown here are representative images of slides. (b) Lysates from tumor xenografts were prepared and the expression of TLR2, TLR4, Akt and GAPDH was analyzed
Expression of p55PIK mRNA and p55PIK protein in mononuclear cells of leukemia patients
Previously, we showed that N24 effects depended on its ability to block p55PIK- mediated signaling pathways by disrupting p55PIK interaction with other proteins. As p55PIK is highly expressed in several cancers,6 we examined the expression of p55PIK in leukemia patients. The mononuclear cells from patients with various types of leukemia and healthy volunteers were collected and the expression of p55PIK in these samples was determined by quantitative real-time RT-PCR. The expression of p55PIK mRNA in mononuclear cell samples from 20 leukemia patients was 6.5 times higher than that from 5 normal controls (P<0.01). In particular, the expression of p55PIK mRNA was 2.4 times in acute myeloid leukemia (AML) M7 subgroup patients (P<0.05), 1.8 times higher in AML M5 subgroup patients (P<0.05), 13.6 times higher in AML M3 subgroup patients (P<0.01) and 9.4 times higher in CML patients (P<0.01) (Figure 6a). Cell lysates from one normal control, two AML M7, M5 and M3 patients, two CML patients as well as K562 cells were prepared, and the p55PIK protein expression in the samples were analyzed using an antibody against p55PIK (Santa Cruz Biotech, Delaware, CA, USA). Consistent with the data from quantitative RT-PCR analysis, the protein level of p55PIK was higher in mononuclear cells from these leukemia patients than in cells from normal controls (Figure 6a, data not shown).
Expression of p55PIK mRNA and p55PIK proteins in human leukemia cells. (a) p55PIK mRNA levels in isolated mononuclear cells from 5 healthy volunteers and 20 leukemia patients (among the patients, five diagnosed as AML M7 subgroup, five as AML M5 subgroup, five as AML M3 subgroup and five as CML) are quantified by real-time reverse transcription-PCR. Shown is summarized result (mean±S.D., *P<0.05, **P<0.01 compared with the p55PIK mRNA level in cells from healthy volunteers). (b) p55PIK protein levels in isolated mononuclear cells from one healthy volunteer (N) and ten leukemia patients (among the patients, two diagnosed as AML M7 subgroup, two as AML M5 subgroup, two as AML M3 subgroup and two as CML) are analyzed in western blotting. p55PIK level in K56 cells was also analyzed
TAT–N24 effects on cell proliferation and differentiation of bone marrow cells from leukemia patients
We next studied TAT–N24 effects on cell proliferation and differentiation of leukemia cells using CD34+ cells isolated from bone marrow of affected patients. TAT–N24 (120 μg/ml for 4 days) induced the cell cycle arrest at G0/G1 phase (91.4% in TAT–N24-treated group versus 78.9% in TAT–N24M-treated group) of bone marrow cells obtained from a 34-year-old patient diagnosed with AML M3 subgroup (Figure 7a). PMA (1 ng/ml for 4 days) slightly inhibited cell cycle progression in the bone marrow cells (82.4 versus 76.0%). PMA also modestly increased the cell population expressing Cd14 (68.6 versus 58.0%). On the other hand, TAT–N24 strongly increased both the number of cells in the cell population expressing Cd14 (74 versus 57.4%) and the average amount of total expression (5.2 times higher in TAT-N24-treated cells versus TAT-N24M-treated cells; Figure 7a). The expression of CD14 in TAT-N24-treated bone marrow cells from four AML M3 leukemia patients significantly increased more than three times (4.6±1.1, P<0.05). Next, CD34+ cells were isolated from bone marrow cells obtained from four CML patients. TAT–N24 inhibited the cell cycle progression and induced differentiation (measured by expression of TLR2 on bone marrow cells) in all the patients tested (data from one CML patient (a 43-year-old male) shown in Figure 7b, and data not shown). Furthermore, the induction of TLR2 by TAT–N24 in bone marrow cells from four CML patients was significant as well (5.8±1.3, P<0.05).
Effects of TAT–N24 on cell cycle progression and differentiation of the bone marrow cells from leukemia patients. (a) Effects of TAT–N24 on the cell cycle progression and expression of differentiation marker Cd14 on bone marrow cells from a leukemia patient (diagnosed as AML M3 subgroup). CD34+ cells were isolated from bone marrow, cultured in 10% FCS medium and incubated with PBS (Control), TAT–N24M (120 μg/ml), TAT–N24 (120 μg/ml) and PMA (20 ng/ml) for 4 days. Cell cycle analysis and the expression of Cd14 on the cell membrane were performed as described.26 Shown are histograms of cell cycle analysis and immunostaining with anti-Cd14 antibody. Number shown is the percentage of cells with Cd14+ expression (M1). Cd14 signal intensity per cells is shown in arbitrary units with the mean of solvent-treated cells (Control) defined as 1. (b) Effects of TAT–N24 on the cell cycle progression and expression of differentiation marker TLR2 on bone marrow cells from a leukemia patient (diagnosed as CML). CD34+ cells were isolated from bone marrow, cultured in 10% FCS medium and incubated with PBS (Control), TAT–N24M (120 μg/ml) and TAT–N24 (120 μg/ml) for 4 days. Cell cycle analysis and the expression of TLR2 on the cell membrane were performed as described. Shown are histograms of cell cycle analysis and immunostaining with anti-TLR2 antibody. Number shown is the percentage of cells with TLR2+ expression (M1). TLR2 signal intensity per cells is shown in arbitrary units with the mean of solvent-treated cells (Control) defined as 1
Discussion
There has been strong interest in using differentiation reagents such as ATRA to convert the malignant phenotype of leukemia cells to a more differentiated one.18, 19 We thus developed TAT–N24, a cell-permeable fusion peptide containing the N-terminal 24 amino acids of p55PIK regulatory subunit of PI3K and the PTD from HIV Tat protein. Our previous studies demonstrated that N24 inhibits cell cycle progression and tumor development of colon and thyroid cancers. In our present studies with HL60 and K562 human leukemia cell lines, and bone marrow cells from leukemia patients, TAT–N24 not only blocked cell proliferation, but also remarkably induced differentiation of leukemia cells.
K562 as well as leukemia cells from many patients with CML harbor the t(9;22) chromosomal translocation that leads to the expression of the 210-kDa BCR/ABL fusion protein with aberrant protein tyrosine kinase activity. IM selectively inhibits the activity of BCR/ABL and is widely used for CML patients.20 Resistance to IM is commonly observed in patients in the accelerated phase of CML; it also can occur during the chronic phase after long-term exposure to this drug.20 Thus, the use of IM in combination with other novel drugs may not only improve response rates but also decrease drug-resistance.21 Our findings show that the combination of IM and TAT–N24 showed enhanced inhibition on the xenograft tumor growth of leukemia cells (Figures 5a and b), suggesting that the efficacy of TAT–N24 may be further improved when used in combination with other drugs. In the past few years, ATRA and its derivatives have become a mainstay in the treatment of certain types of leukemia.22 Our data showed that the mechanism for N24 induction on leukemia differentiation is different from ATRA. Thus, TAT–N24 by itself or in combination with ATRA may be useful as an antileukemic differentiation therapy, particularly given its unique target and low toxicity in vivo. Moreover, such differentiation therapies may also be less toxic than traditional chemotherapies, which often are cytotoxic.23
N24 blocks p55PIK signaling pathways by disrupting the interaction of p55PIK with other proteins such as Rb.5, 7 Thus, N24 likely has little or no effect on the enzymatic activities of the PI3K catalytic subunits. The mechanism(s) for how cell cycle arrest by N24 leads to the cell differentiation remains to be elucidated. The PI3K p110 catalytic activity inhibitor, LY294002, inhibits the cell proliferation without significant effects on the differentiation of K562 or HL60 cells (data not shown),24 suggesting that the enzymatic activities of PI3K are necessary for both cell proliferation and cell differentiation.
There also is extensive evidence for PI3K involvement in tumor development and progression.25 Several inhibitors targeting PI3K catalytic subunit isoforms are effective in decreasing cell proliferation in cancer cell lines and animal models.3 However, lack of specificity limits their clinical application, particularly as only a subset of the many cellular processes regulated by the PI3K pathway are directly involved in dysregulation of cell proliferation.2 Thus, a critical issue for the clinical application of PI3K inhibitors is the development of therapeutics that can specifically block pathways involved in tumor growth without the inhibition of other PI3K-dependent cellular processes. In contrast, our studies showed that TAT–N24 can be delivered systemically and is effective in vivo without significant side effects, suggesting that p55PIK may be an excellent drug target for leukemia and cancer.
In summary, we have generated a fusion peptide containing the p55PIK amino terminus, TAT–N24, which not only inhibited cell proliferation, but also stimulated differentiation in HL60 and K562 human leukemia cells and in mononuclear cells from patients with different types of leukemia. Additionally, there were significant increases in p55PIK mRNA expression in leukemic cells from patients, suggesting that it may have an etiological role in oncogenesis and cell proliferation. Thus, our findings suggest that TAT–N24 is a useful tool to study the signaling pathways involved in cell proliferation and differentiation mediated by p55PIK and may be an effective clinical therapy for leukemia. Finally, our studies with TAT–N24 demonstrate the feasibility of using PTDs for systemic peptide delivery and thus provide the basis for the development of other novel rationally designed peptide therapies for cancer and other systemic diseases.
Materials and Methods
Ethics statement
All research involving human participants had been approved by the Huazhong University of Science and Technology Ethics committee, and we had obtained informed consent that was written from all participants involved in this study. All animal work had been conducted according to Huazhong University of Science and Technology animal study guidelines.
Expression and purification of TAT–N24 and control TAT–N24M fusion proteins
cDNA fragments encoding 6xHis tag, PDT in Tat protein (YGRKKRRQRRR), HA tag (YPYDVPDYA) and N24 (MDRDDWDWEVVMMPYSTELIPYIT) were introduced into pMD18-T Easy vector (Invitrogen, Grand Island, NY, USA). His–TAT–HA–N24 cDNA fragment then was subcloned into the vector pET32a (Invitrogen). To generate the control fusion protein (TAT–N24M), the N24 cDNA sequence was replaced by cDNA encoding the shuffled N24 (MDRDDWDWEVIMMPYSEPLYTIVT). The TAT–N24 and TAT–N24M fusion proteins have 96 amino-acid residues. The obtained recombinant vectors (designated pHisTAT–N24 and pHisTAT–N24M) were transformed into E. coli BL21 (DE3) for fusion protein expression.
For the purification of TAT–N24 and TAT–N24M fusion proteins, bacteria were lysed in 6 M urea and the affinity purification of TAT–N24 and TAT–N24M was done using Ni++ agarose beads according to the standard protocols. The purified fusion proteins were dialyzed against PBS (pH 9.5), filtered, and stored in aliquots at 4°C.
Western blotting and immunofluorescence and immunohistochemical analysis
Lysates from cells and tumor xenografts were prepared as previously described.7 Immunoblot assays with various antibodies and nuclear staining with propidium iodide were performed as previously described.7 Intensity of immunofluorescence signal was determined by flow cytometry analysis. Immunohistochemical staining of tumor sections with various antibodies was performed as previously described.7 For the immunohistochemical staining of tumor section with CD11b antibody, the frozen tumor samples were cut and prepared on glass slides. The samples were fixed in 95% ethanol for 5 min. Immunostaining using phosphatidylethanolamine (PE)-labeled anti-CD11b antibody (BD Biosciences, San Jose, CA, USA) and nuclear staining with DAPI was done as described.26
Analysis of cell cycle profile and evaluation of morphological changes
Cell cycle profile was analyzed as described before.7 The percentage of positive cells and the mean associated fluorescence were quantitated using a FACScan analyzer.7 HL60 or K562 cells were fixed by methanol and stained with Wright-Giemsa (Sigma, St. Louis, MO, USA). Morphology was evaluated using a Zeiss (Guangzhou, China) Axioplan microscope.24 Images were color balanced in Adobe Photoshop.
Preparation of adenovirus construc expressing p55PIK
The cDNA encoding the full-length human p55PIK obtained by PCR amplification from K562 cells was cloned into shuttle vector pTrack-CMV-EGFP (Clontech). The construction of adenoviruses expressing p55PIK and GFP or GFP alone (control adenovirus) was performed as described.7 The preparation of adenovirus particles and its application in cell culture and animal study was done as described.7
RNA isolation and analysis
Total RNA was isolated from cells or tumors using TRIsol reagent (Invitrogen). Total RNA was reverse transcribed using a kit (One-Step PCR, Qiagen, Hilden, Germany) and then subjected to RT-PCR analysis. Primers used included for Cd11b (forward: 5′-GCTGCCGCCATCATCTTACG-3′; reverse: 5′-CCCTCGGGTCTGCTCGTAG-3′), Cd14 (forward: 5′-ACTTGCACTTTCCAGCTTGC-3′; reverse: 5′-TCGCCCAGTCCAGGATTGTC-3′), TLR4 (forward: 5′-AATGGATCAAGGACCAGAGGC-3′; reverse: 5′-GCAGCCAGCAAGAAGCATCAG-3′), p55PIK (forward: 5′-GAGTATGGACCGCGATGACG-3′; reverse: 5′-GCTTAGGTGGCTTTGGTGGAA-3′), CD38 (forward: 5′-TGCCAAAGTGTATGGGATG-3′; reverse: 5′-GTAGCCTAGCAGCGTGTC)-3′, CD163 (forward: 5′-ATGTGGAGTTGCCCTTTC-3′; reverse: 5′-TTTACCAGGCGAAGTTGA-3′), CD41 (forward: 5′-TCCTCAGAAGCTATCCCTAAAT-3′; reverse: 5′-GGAAGTCTGCCTCATCTCG-3′), CD61 (forward: 5′-TGTCACGGAACCGAGATG-3′; reverse: 5′-GGTAGTGGAGGCAGAGTAATG-3′), CCL2 (forward: 5′-CTTCTGTGCCTGCTGCTC-3′; reverse: 5′-TGCTGCTGGTGATTCTTCT-3′), CCR (forward: 5′-TGGTCCTGCCGCTGCTCAT-3′; reverse: 5′-TGTCACCTGCGTGGCTTGG-3′), Actin (forward: 5′-GTGGACATCCGCAAAGA-3′; reverse: 5′-CTCGTCATACTCCTGCTTG-3′) and GAPDH (forward: 5′-GGTGTGAACCATGAGAAGTATGACAAC-3′; reverse: 5′-CCAGTAGAGGCAGGGATGATGTTC-3′).
Primary human leukemia samples
All leukemia patients involved in this study was diagnosed and classified when receiving treatment in Tongji Hospital, Wuhan, China. Heparinized peripheral bloods were obtained from healthy volunteers and leukemia patients after their informed consent during the course of routine clinical care or as part of an Institutional Review Board–approved specimen bank protocol. Mononuclear cells were then isolated by density gradient centrifugation, suspended in 10% FCS medium. For analyzing the expression of p55PIK mRNA, total RNA was isolated from mononuclear cells and analyzed in RT-PCR. Bone marrow aspirates were obtained from leukemia patients after their informed consent. Mononuclear cells were then isolated by density gradient centrifugation, suspended in 10% FCS medium. CD34+ cells were separated from bone marrow mononuclear cells of eight patients (four CML, four AML M3 subgroup) by using positive immunomagnetic column separation kit (Miltenyi Biotech, Auburn, CA, USA) as described.27 Their use for these studies was approved by the Huazhong University of Science and Technology Institutional Review Board. Cells were treated with TAT–N24 and other agents for 3 days and harvested for cell cycle analysis and determination of differentiation markers expression.
Animal studies
Athymic male nude mice at 8–10 weeks of age weighing 22–24 g were used in this study. All animal experiments were done in accordance with animal research guidelines from Huazhong University of Science and Technology.
Lipid emulsion injection was formulated from equal volume of TAT–N24 or TAT–N24M proteins or 6 M urea (pH 9.5) and injectable lipid emulsion (30% Intralipid, Sino-Swed Pharmaceutical Corp, Ltd, Beijing, China). For experiments investigating the TAT–N24 protein effect on the xenograft tumor growth, mice were injected with 200 μl TAT–N24 or TAT–N24M lipid emulsion or equal amount of solvent–lipid emulsion via tail vein on the same day shortly after tumor inoculation. HL60 or K562 (107 ) cells were injected s.c. into mice. The mice in each group, consisting of 4–6 animals, received injections of 200 μl protein–lipid emulsion or solvent–lipid emulsion. Injections were repeated every 2 days for 10–14 days. In animals treated with ATRA (20 μg in 2 μl ethanol) or IM was injected i.p. once per day after inoculation. IM (kindly provided by Novartis Pharma (Basel, Switzerland) was injected intraperitoneally at 20 mg/kg per day at day 7 after inoculation. The animals were monitored for the tumor growth after inoculation and tumors were collected and analyzed.
Statistical analysis
The data are presented as mean±S.D. of the mean. P-values were calculated using an unpaired Student’s t-test when only two groups were compared. P-values ≤0.05 were considered to be statistically significant (designated by a single asterisk; double asterisk, P≤0.01). The number of animals in each group is indicated by n.
Abbreviations
- PTD:
-
protein transduction domain
- PI3K:
-
phosphatidylinositol 3-kinases
- CML:
-
chronic myeloid leukemia
- AML:
-
acute myeloid leukemia
- ATRA:
-
all-trans retinoid acid
- PMA:
-
phorbol myristate acetate
- TLR:
-
Toll-like receptor
- IM:
-
imatinib
References
Fruman DA, Meyers RE, Cantley LC . Phosphoinositide kinases. Annu Rev Biochem. 1998; 67: 481–507.
Vanhaesebroeck B, Waterfield MD . Signaling by distinct classes of phosphoinositide 3-kinases. Exp Cell Res 1999; 253: 239–254.
Samuels Y, Diaz LA, Schmidt-Kittler O, Cummins JM, Delong L, Cheong I et al. Mutant PIK3CA promotes cell growth and invasion of human cancer cells. Cancer Cell 2005; 7: 561–573.
Ward SG, Finan P . Isoform-specific phosphoinositide 3-kinase inhibitors as therapeutic agents. Curr Opin Pharmacol 2003; 3: 426–434.
Xia X, Cheng A, Akinmade D, Hamburger AW . The N-terminal 24 amino acids of the p55 gamma regulatory subunit of phosphoinositide 3-kinase binds Rb and induces cell cycle arrest. Mol Cell Biol 2003; 23: 1717–1725.
Zhang L, Huang J, Yang N, Greshock J, Liang S, Hasegawa K et al. Integrative genomic analysis of Phosphatidylinositol 3′-kinase family identifies PIK3R3 as a potential therapeutic target in epithelial ovarian cancer. Clin Cancer Res 2007; 13: 5314–5321.
Hu J, Xia X, Cheng A, Wang G, Luo X, Reed MF et al. A peptide inhibitor derived from p55PIK PI3K regulatory subunit is a novel cancer therapy. Mol Cancer Ther 2008; 7: 3719–3728.
Schwarze SR, Ho A, Vocero-Akbani A, Dowdy SF . In vivo protein transduction: delivery of a biologically active protein into the mouse. Science 1999; 285: 1569–1572.
Dietz GP, Bahr M . Synthesis of cell-penetrating peptides and their application in neurobiology. Methods Mol Biol 2007; 399: 181–198.
Hotchkiss RS, McConnell KW, Bullok K, Davis CG, Chang KC, Schwulst SJ et al. TAT-BH4 and TAT-Bcl-xL peptides protect against sepsis-induced lymphocyte apoptosis in vivo. J Immunol 2006; 176: 5471–5477.
Luo P, Wang A, Payne KJ, Peng H, Wang JG, Parrish YK et al. Intrinsic retinoic acid receptor alpha-cyclin-dependent kinase-activating kinase signaling involves coordination of the restricted proliferation and granulocytic differentiation of human hematopoietic stem cells. Stem Cells 2007; 25: 2628–2637.
Nishioka C, Ikezoe T, Yang J, Gery S, Koeffler HP, Yokoyama A . Inhibition of mammalian target of rapamycin signaling potentiates the effects of all-trans retinoic acid to induce growth arrest and differentiation of human acute myelogenous leukemia cells. Int J Cancer 2009; 125: 1710–1720.
Zhao KW, Li X, Zhao Q, Huang Y, Li D, Peng ZG et al. Protein kinase Cdelta mediates retinoic acid and phorbol myristate acetate-induced phospholipid scramblase 1 gene expression: its role in leukemic cell differentiation. Blood 2004; 104: 3731–3738.
Altucci L, Leibowitz MD, Ogilvie KM, de Lera AR, Gronemeyer H . RAR and RXR modulation in cancer and metabolic disease. Nat Rev Drug Discov 2007; 6: 793–810.
Alitalo R . Induced differentiation of K562 leukemia cells: a model for studies of gene expression in early megakaryoblasts. Leuk Res 1990; 14: 501–514.
Hawk N, Sun T, Xie S, Wang Y, Wu Y, Liu J, Arlinghaus RB . Inhibition of the Bcr-Abl oncoprotein by Bcr requires phosphoserine 354. Cancer Res 2002; 62: 386–390.
Hehlmann R, Hochhaus A, Baccarani M . Chronic myeloid leukaemia. Lancet 2007; 370: 342–350.
Sachs L . The control of hematopoiesis and leukemia: from basic biology to the clinic. Proc Natl Acad Sci USA 1996; 93: 4742–4749.
Lotem J, Sachs L . Epigenetics wins over genetics: induction of differentiation in tumor cells. Seminars Cancer Biol 2002; 12: 339–346.
Shah NP, Nicoll JM, Nagar B, Gorre ME, Paquette RL, Kuriyan J et al. Multiple BCR-ABL kinase domain mutations confer polyclonalresistance to the tyrosine kinase inhibitor imatinib (STI571) in chronic phase and blast crisis chronic myeloid leukemia. Cancer Cell 2002; 2: 117–125.
Schiffer CA . BCR-ABL tyrosine kinase inhibitors for chronic myelogenous leukemia. N Engl J Med 2007; 357: 258–265.
Spira AI, Carducci MA . Differentiation therapy. Curr Opin Pharmacol. 2003; 3: 338–343.
Sell S . Stem cell origin of cancer and differentiation therapy. Crit Rev Oncol Hematol 2004; 51: 1–28.
Matkovic K, Brugnoli F, Bertagnolo V, Banfic H, Visnjic D . The role of the nuclear Akt activation and Akt inhibitors in all-trans-retinoic acid-differentiated HL-60 cells. Leukemia 2006; 20: 941–951.
Vogt PK, Hart JR, Gymnopoulos M, Jiang H, Kang S, Bader AG et al. Phosphatidylinositol 3-kinase: the oncoprotein. Curr Top Microbiol Immunol. 2010; 347: 79–104.
Cheng A, Wang S, Cai J, Rao MS, Mattson MP . Nitric oxide acts in a positive feedback loop with BDNF to regulate neural progenitor cell proliferation and differentiation in the mammalian brain. Dev Biol 2003; 258: 319–333.
Yin T, Wu YL, Sun HP, Sun GL, Du YZ, Wang KK et al. Combined effects of As4S4 and imatinib on chronic myeloid leukemia cells and BCR-ABL oncoprotein. Blood 2004; 104: 4219–4225.
Acknowledgements
This work was supported by grants from the National Natural Science Foundation of China (30872472, 30973496, JH), (30800569, XL); National Basic Research Program of China (973-2009CB521802, JH). The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
Author contributions
GW, YD, XC, SL, YT, XL and YF performed experiments; JH, GW and XX analyzed results and made the figures; JH, XX and JG designed the research and wrote the paper.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Edited by M Piacentini
Supplementary Information accompanies the paper on Cell Death and Differentiation website
Supplementary information
Rights and permissions
About this article
Cite this article
Wang, G., Deng, Y., Cao, X. et al. Blocking p55PIK signaling inhibits proliferation and induces differentiation of leukemia cells. Cell Death Differ 19, 1870–1879 (2012). https://doi.org/10.1038/cdd.2012.70
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/cdd.2012.70
Keywords
This article is cited by
-
Comparative Transcriptome Analysis of Spleen Reveals Potential Regulation of Genes and Immune Pathways Following Administration of Aeromonas salmonicida subsp. masoucida Vaccine in Atlantic Salmon (Salmo salar)
Marine Biotechnology (2022)
-
LncRNA JHDM1D-AS1 Suppresses MPP + -Induced Neuronal Injury in Parkinson’s Disease via miR-134-5p/PIK3R3 Axis
Neurotoxicity Research (2021)
-
PIK3R3 inhibits cell senescence through p53/p21 signaling
Cell Death & Disease (2020)
-
PIK3R3 regulates PPARα expression to stimulate fatty acid β-oxidation and decrease hepatosteatosis
Experimental & Molecular Medicine (2018)
-
Loss of BRG1 induces CRC cell senescence by regulating p53/p21 pathway
Cell Death & Disease (2017)