Abstract
p63, an ancestral transcription factor of the p53 family, has three C-terminal isoforms whose relative in vivo functions are elusive. The p63 gene is essential for skin and limb development, as vividly shown by two independent global knockout mouse models. Both strains, although constructed differently, have identical and severe phenotypes, characterized by absent epidermis and hindlimbs and only rudimentary forelimbs at birth. Here we show that mice from one model, Brdm2, express normal levels of truncated p63 proteins that contain the DNA binding and oligomerization domain but lack the long carboxy-terminal SAM (sterile α-motif) and post-SAM domains that are specific for the α and β isoforms. As such, transcriptionally active p63 proteins from Brdm2 mice resemble the naturally occurring p63γ isoforms, which of all the p63 isoforms most closely resemble p53. Thus, Brdm2 mice are p63α/β isoform-specific knockout mice, gaining unexpected new importance. Our studies identify that p63α/β but not p63γ are absolutely required for proper skin and limb development.
Similar content being viewed by others
Main
The transcription factor p63, a member of the p53 family and in parts a structural homolog, plays a pivotal role in epidermal and limb development, as shown by two mouse models. The p63 gene consists of 14 exons and produces 6 well-described isoforms, derived from two promoters (TA and ΔN) that each give rise to three different carboxy-termini through alternative splicing. The longest isoform α skips exon 10′ (originally called exon 15, see Genbank Accession No. AF533892), the β isoform skips exon 13 and the γ isoform ends with alternatively spliced exon 10′.1
Soon after the gene's discovery, two groups generated p63 knockout models but used different embryonic stem-cell ablation technologies for p63 exon disruption.2, 3 McKeon and colleagues replaced exons 6–8 in the DNA-binding domain (which is common to both TA and ΔN isoforms) with a Neo cassette, leading to a complete loss of all p63 proteins. In contrast, Mills et al. employed a gap-repair mechanism for p63 targeting, using an insertional vector from a pre-existing library.4 In the Mills et al. mutant allele named Brdm2,2 the target vector insertion interrupted the gene only after exon 10, followed by a net duplication of the region of homology (exons 5–10), plus the endogenous exons 10′–14 (Figure 1a). Importantly, the phenotypes of both homozygous mutant models are identical and dramatic: newborns with essentially no epidermis or epidermal appendages, absence of hind limbs, only rudimentary forelimbs, craniofacial abnormalities and profound defects in other stratified epithelial tissues (urothelium, mammary and prostate).2, 3 In contrast, heterozygous mice for either of these mutations showed none of these phenotypes.2, 3
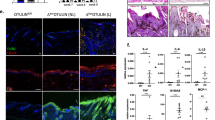
Brdm2 mutant squamous epithelium expresses p63 protein(s). (a) The targeted Brdm2 allele is shown.2 The pTV12E(60) vector used for the gap repair method of gene ablation of Trp63 in Brdm2 mice contains an Agouti minigene, the 3′ portion of an hprt cassette followed by its 3′UTR, a loxP site, a puromycin resistance gene, and a 129S5-derived DNA fragment containing Trp63 exons 5, 6 and 10.4 Insert genotyping of our Brdm2/Brdm2 (B/B) mice across the Trp63 genome/vector border confirmed the insertion of the targeting vector between exons 10 and 10′ (red line, mutant-specific PCR product). Note: exon10′ was formerly called exon15 (see Genbank AF533892). (b) WT and p63 Brdm2/Brdm2 littermate embryos at day E18. (c, d) WT embryo at E18. p63 expression in basal and suprabasal cells of skin and hair follicles. (c) H&E section. (d) Immunoperoxidase staining with p63-specific antibody 4A4. (d, f, h, k–n) All 4A4 immunostainings were carried out side-by-side, using identical conditions. (e, f) Microscopically, p63B/B embryos at day E18 show essentially absent epidermis, leaving only underlying compressed dermis. (e) H&E. However, rare remnant epidermal patches with relatively normal squamous stratification and hair follicles are present, which (f) express p63 in basal and suprabasal cells. (g, h) In mechanically protected anatomic niches like the anterior nasal cavity, p63B/B embryos at day E18 show normal appearing squamous epithelium that expresses p63. (g) H&E section. (h) p63-expressing squamous epithelium lining the vestibulum and lateral nasal cavity. Enlarged view of boxed area in (g). Respiratory epithelium covering the nasal septum is p63-negative (not shown). (i) WT and p63B/B littermate embryos at day E15. In contrast to mutant E18 embryos which have all but lost their epidermis, mutant E15 embryos do show an — although thin and translucent — but completely intact epidermal integument which is visible by eye under 4 × magnification. (j–n) Mutant E15 epidermis consists of a contiguous 3–5 cell layer terminally differentiated squamous epithelium with focal thickening and patchy underlying hair follicles, expressing p63 in basal and suprabasal cells. (j) H&E sections. (k–n) 4A4 immunoperoxidase-stained sections. (c, d, f, j–n) 10 × magnification, (e, h) 40 × magnification, (g) 4 × magnification
Nevertheless, over the ensuing years both the KO models have generated a vexing degree of disagreement that remains unresolved.5 Examples include the role of p63 in skin epithelial commitment and terminal differentiation versus stem cell proliferation5, 6, 7 and in oncogenic versus tumor suppressor function in tumorigenesis.8, 9, 10 It should be noted that by far most studies on p63 function were carried out on the Brdm2 strain,2, 6, 8, 9, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24 from which major conclusions were drawn about its role in developmental neuronal apoptosis,14 in germ cell apoptosis during testicular development and irradiation,19, 22, 23 and in organismal senescence during aging.15
Results
Brdm2 embryos express normal levels of truncated p63 proteins that contain the DNA binding and oligomerization domain but lack the long carboxy-terminal SAM and post-SAM domains
For unrelated studies we needed p63 null tissues. Genotyping of Brdm2/Brdm2 mice (gift of A Mills) across the p63 genomic/vector border invariably confirmed the reported insertion of the targeting vector between exon 10 and 10′ in all embryos studied (Figure 1a).2 A total of 26 Brdm2/Brdm2 embryos were examined, ranging from day E13 to P1 newborns. Phenotypes were invariant. As expected,2, 3 by macroscopic inspection, late stage E18 Brdm2/Brdm2 embryos (called B/B in Figures) exhibited a profound lack of epidermis, absent hindlimbs and severely underdeveloped forelimbs (Figure 1b). In contrast, p63 wild-type (WT) and heterozygous (WT/Brdm2, data not shown) littermates showed well-developed limbs and epidermis (Figure 1b), with p63 expression in the nuclei of basal and suprabasal cells of skin and hair follicles (Figure 1c and d). This was readily detectable by the classic monoclonal pan-p63 antibody 4A4 that detects all isoforms. The apical ectodermal ridge in WT, but not p63−/−, limb buds also expresses high levels of p63 protein.3 Microscopically, mutant E18 Brdm2/Brdm2 embryos were essentially devoid of epidermis, exposing only compressed dermis (Figure 1e and f). However, close inspection showed rare patches of keratinizing epidermis with relatively normal squamous stratification overlying hair follicles (Figure 1e and f). Unexpectedly, these patches expressed high levels of nuclear p63 in basal and suprabasal cells, comparable with WT tissue (Figure 1f). Moreover, in mechanically protected anatomic niches such as the anterior nasal cavity (Figure 1g and h) and the buccal mucosa of the oral cavity of mutant mice (Figure 2a and b), well-developed fragments of normal appearing non-keratinizing squamous epithelium were present. These areas also expressed p63 at WT levels (Figures 1h and 2b). Furthermore, p63 was strongly expressed in cell nuclei of the anterior pituitary as well as in the gonocytes of testis and ovary of Brdm2/Brdm2 mice (Figure 2c and d and data not shown). To confirm this completely, unexpected p63 expression in mutant Brdm2/Brdm2 mice, we validated the p63 specificity of the 4A4 antibody which was raised against aa 1–205 of ΔNp63, spanning the end of exon 3′ to the start of exon 7 within the DNA-binding domain.1 First, remnant epidermal patches and internal squamous and stratified mucosa in McKeon p63 KO mice3 at day E17 did not yield any 4A4 staining (Figure 3a and b and data not shown). Moreover, specific pre-absorption with lysates of H1299 cells transfected with a 1 : 1 mixture of TAp63α and ΔNp63α expression plasmids abolished or severely reduced the nuclear 4A4 staining (Figure 3c and d), whereas pre-absorption with empty vector lysates did not (Figure 3e). In addition, when 4A4 was omitted, immunostaining was lost (data not shown). Finally, 4A4 recognizes the same specific cell types (basal keratinocytes, squamous carcinoma cells, neurons, urothelium, breast and prostate) — and no other — in mouse and human tissues, where it has become a clinically validated diagnostic antibody for p63 (e.g., Ref24) (Figure 3f and g and data not shown). Together, these data confirm the exclusive specificity of the 4A4 antibody for p63 and further exclude any possible cross-reactivity with p53 family members in skin and other stratified epithelia. To further confirm p63 expression in Brdm2/Brdm2 embryos, we used a second independent anti-p63 antibody, polyclonal H-137 raised against aa 15–151 of ΔNp63. As expected, H-137 immunostained epidermis and internal epithelia in WT E18 embryos (Figure 3h and data not shown). It should be noted that, H-137 also immunostained epidermis and internal epithelia in Brdm2/Brdm2 embryos in a pattern identical to the one seen with 4A4 (Figure 2l–m and data not shown).
p63 expression in squamous epithelium, pituitary and internal stratified squamous mucosa of Brdm2/Brdm2 embryos. (a, b) At day E18, p63 Brdm2/Brdm2 embryos show remnants of normal squamous epithelium that expresses normal levels of p63, particularly in mechanically protected anatomic niches. Although cheeks and much of the oral cavity and tongue are denuded at this stage, squamous epithelium is still lining parts of the lip and buccal mucosa. (c, d) p63 Brdm2/Brdm2 embryo at day E18 with normal anterior lobe of the pituitary gland. This glandular epithelium, located in the sella turcica of the skull base and embryologically derived from the oral ectoderm of Rathke's pouch, also strongly expresses p63 in nuclei. (a, c) H&E sections. (b, d) 4A4 immunoperoxidase staining of areas within the framed H&E sections. (e–k) At day E15, p63 Brdm2/Brdm2 embryos show internal stratified squamous mucosa covering the tongue, buccal areas, oropharynx and nasopharynx, esophagus, forestomach and bladder which appears intact by H&E (e, h) and expresses p63 (f, g, i–k). 4A4 immunoperoxidase staining. (l, m) Immunoperoxidase staining with pan p63-specific polyclonal H137 of esophagus and epidermis. E15 p63 Brdm2/Brdm2 embryos. (a, e, f) 4 × magnification, (b, d, g, h–m) 40 × magnification, (c) 10 × magnification
Validation of p63 specificity of classic monoclonal antibody 4A4. (a, b) Remnant squamous epithelium from skin of p63−/− mice (Yang A et al.3) at day E17 is devoid of any 4A4 staining. Only non-specific background is detectable. (c–e) Nuclear staining by monoclonal p63 antibody 4A4 is specifically eliminated by p63 proteins. Skin of a WT embryo at day E19, (c) stained with 4A4 without pre-absorption, (d) 4A4 specifically pre-absorbed with H1299 lysates transfected with a 1 : 1 mixture of TAp63α and ΔNp63α plasmids (SDS-free, 50-fold excess by total protein w:w), and (e) 4A4 that had been control pre-absorbed with empty vector-transfected H1299 lysates (SDS-free, 50-fold excess by total protein w:w). The same amount of 4A4 IgG (2 μg) and conditions were used in all cases. Note that 4A4 staining becomes even sharper with non-specific control pre-absorption. Omitting the primary antibody also eliminated all staining (not shown). (f, g) Specificity validation of 4A4 on human tissue. The antibody stains the same cell types (basal keratinocytes and squamous carcinoma cells) in mouse and human tissues. 4A4 immunoperoxidase staining on sections of normal human skin and invasive squamous cell carcinoma. (h) Specificity validation of H137 on E18 WT mouse skin. (a, b) 40 × magnification, (c–h) 10 × magnification
We rationalized that the squamous patches in mutant E18 Brdm2/Brdm2 embryos might be surviving remnants of a better developed epidermis from an earlier stage. E15 is the developmental stage when commitment to terminal differentiation occurs.16 Indeed, while the limb truncation phenotype of E15 Brdm2/Brdm2 embryos was already fully evident, they exhibited a thin and translucent but widely intact epidermal integument when closely inspected with a dissecting microscope (Figure 1i; note: the photograph does not capture the image seen by eye). The presence of an although thin but intact integument at E15 was confirmed microscopically (Figure 1j and n). The E15 mutant epidermis consists of a largely contiguous, terminally differentiated keratinizing squamous epithelium that is 3–5 cell layers thick, with focal thickening (Figure 1j and m) and patchy underlying hair follicles (Figure 1k and l). Importantly, the mutant epidermis strongly expresses the nuclear p63 in an orderly row of basal progenitors and suprabasal keratinocytes, with a p63 distribution and staining intensity similar to that of WT epidermis (Figure 1k–n). Moreover, the internal stratified squamous mucosa covering the tongue, buccal areas, oropharynx and nasopharynx, esophagus, forestomach and bladder was invariably well developed by H&E and stained strongly positive for p63 with both monoclonal 4A4 and polyclonal H137 antibodies (Figure 2e–m). Thus, the epithelializations of E18 Brdm2/Brdm2 embryos appear to be the remnants of a better developed epithelium at stage E15, which, importantly, is of a transient nature. Future study focusing on the role of p63 in epithelial stemness and/or adhesion will clarify why the embryonic epidermis is well formed at day E15 but disappears at the later stages in these mutant embryos.
To further support the p63 protein expression in mutant Brdm2/Brdm2 animals, we carried out quantitative RT-PCR analysis of the E15 embryos and mouse embryo fibroblasts (MEFs), derived from E13 embryos. Multiple p63 transcripts derived from exons 1–10 (including TA and ΔN variants) were readily detectable in amounts comparable with that of WT mice when normalized for either GAPDH, β-actin or the ribosomal protein Rplp0, present in all tissues (Figure 4b–d and data not shown, summarized in Figure 4a). p63 expression was also normalized to skin markers. These were present at 6–8-fold lower levels in the mutant embryos, confirming the visual skin impression (see Figure 1i). Indeed, when p63 transcript levels were normalized for multiple epidermal differentiation markers such as cytokeratin K5 and K14 (basal cell layer), K10 (intermediate layer), loricrin (top layer) and desmosomal marker PERP (all layers), on the whole, these broad-based normalizations showed that the mutants expressed levels of p63 exons 1–10 that were comparable with those of WT (Figure 4b, Supplementary Figure 1 and data not shown). In contrast, p63 exons 4–10′ and exons 4–11 transcripts were only detected in WT but never in Brdm2/Brdm2 embryos (Figure 4b). Thus, transcription from the Brdm2 allele occurs from exons 1–10 but does not cross the vector insert. Instead, two new transcripts extending from exon 10 are readily detectable in Brdm2/Brdm2 embryos at day E15. Both are without the 37 amino acid tail of p63γ encoded by exon 10′ and the extensive carboxy-termini of p63α and β encoded by exons 11–14. One transcript is a read-through from exon 10 into the adjacent intron 10, forming an open reading frame that is then immediately terminated by a stop codon after only 1 nucleotide into intron 10. The other is the result of splicing exon 10 in frame onto the non-functional 3′ portion of the hprt minigene (Figure 4a). The 3′hprt cassette has a splice acceptor site at its 5′ end and is followed by its 3′UTR (Figures 1a and 4a). 3′ RACE and sequencing further confirmed the presence of both new transcripts exclusively in Brdm2/Brdm2 embryos. These truncated transcripts were apparently missed in the original study using northern blots that detected long 5 kb mRNA species from the p63 WT allele,2 possibly because they are much smaller than the bona fide p63α mRNAs. Taken together, the invariant presence of the targeting vector after exon 10 at the reported location (Figure 1a, Mills et al., 1999), the absence of WT RT-PCR products that span the vector insertion (exons 4–10′ and exons 4–11) in Brdm2/Brdm2 embryos (Figure 4b) and the reproducibility of the Brdm2/Brdm2 phenotype in random litters exclude the proforma possibility of an accidental reversion to a WT p63 gene by spontaneous excision of targeting vector plus its duplicated exons 5–10.
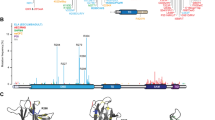
Expression of p63Ex10-truncated transcripts from the Brdm2 allele. (a) Summary of RT-PCR products (red lines) obtained from p63 Brdm2/Brdm2 and WT embryos at day E15. Products obtained only in WT embryos are shown separately (bottom). (b) Quantitative real-time RT-PCR analysis of p63 transcripts in p63 Brdm2/Brdm2 and WT embryos at day E15. Columns 1–3 show common transcripts, columns 4–6 show unique transcripts. Relative amounts of p63 normalized to GAPDH or loricrin, an epidermal differentiation marker, with WT levels set to 1.0 whenever above zero. Additional normalizations to cytokeratin K5, K10, K14, and desmosome marker PERP confirmed comparable p63 transcript levels between mutant and WT embryos (Supplementary Figure 1 and data not shown). Error bars indicate standard error from three independent experiments. Columns 1 and 4 qualitatively show the corresponding amplicons after 40 cycles. For controls, PCR reactions on genomic templates not reverse transcribed (-RT) were run. Individual amplicons are indicated. (c, d) RT-PCR analysis of mouse embryo fibroblasts (MEFs) (passage 5) derived from WT and p63 Brdm2/Brdm2 (B/B) embryos. Transcripts are shown after 40 cycles. GAPDH as input control. Individual amplicons are indicated
Transcriptionally active p63 proteins from Brdm2 mice resemble the naturally occurring p63γ isoforms
Immunoblots showed that the recombinant Brdm2 exon 10-truncated p63 proteins, TA(1–10) and ΔN(3′–10), have the same steady-state levels as the respective native mouse p63γ. In contrast, Hprt fusion proteins of TA(1–10) and ΔN(3′–10) are unstable (Figure 5a). To test the function of Brdm2 p63 proteins, we performed luciferase reporter assays using the p53/p63-responsive human Mdm2 promoter P2 as driver13 and compared their activities with bona fide mouse p63γ. In sum, TA(1–10) and ΔN(3′–10) behave indistinguishably from genuine p63γ. Specifically, in transactivation assays TA(1–10) was at least as active as native TAp63γ (Figure 5b), whereas both ΔN(3′–10) and ΔNp63γ exhibited only minimal transactivation (Figure 5c). The respective p63-Hprt fusions were inactive in transactivation (Figure 5b and c bottom). Likewise, in competitive transrepression assays using either p53 or TAp63γ as the activity to be suppressed, ΔN(3′–10) and ΔNp63γ again behaved indistinguishably (Figure 5d). On the other hand, ΔN(3′–10)Hprt fusion was equally repressive towards p63, but less so towards p53 (Figure 5d). Given that mixed tetramers between WT p53 and p63 proteins do not form,12 but p63 homotetramers do, we speculate that this reflects the fact that the bulky Hprt tag might impair direct DNA binding of ΔN(3′–10)Hprt homotetramers, whereas it still allows to engage in mixed tetramers with TAp63γ, thereby acting as a stronger dominant negative. At any rate, since it is unclear whether Hprt fusions are stable in vivo, it suggests that the latter Brdm2 protein plays either no or only a minor role.
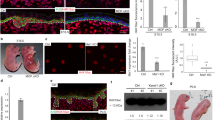
Transcriptional functions of p63exon10-truncated proteins generated from the Brdm2 allele. (a) Steady-state levels of recombinant p63 proteins from the Brdm2 allele. Native mouse p63γ is shown for comparison. All cDNAs were expressed from the same pcDNA3 vector and plasmid amounts. Although the TA(1–10) and ΔN(3′–10) p63 proteins have the same steady-state levels as the respective p63γ, the HPRT fusion proteins, adding a 173 amino acid tail, are unstable compared with their respective p63γ counterparts. H1299 cells were transfected with 200 μg plasmid each. After 24 h, the same percent of cell lysates (20 μg per lane) was immunoblotted with 4A4. (b, c) Transcriptional activity of Brdm2-derived p63 products, compared with native p63γ. Luciferase reporter assays with the p53/p63-responsive human Mdm2 promoter BP100Luc. TA(1–10) and ΔN(3′–10) behave indistinguishably from genuine p63γ. Hprt-fusion proteins are inactive as transactivator, whereas ΔN-Hprt functions partially as transrepressor of TAp63γ and p53. (b, c bottom) Transactivation by the indicated p63 proteins. Respective protein amounts shown by 4A4 immunoblot in (b, c top). (d) Transrepression of co-expressed p53 or TAp63γ by the indicated p63 proteins, shown on top. Plasmid amounts were adjusted to obtain equal protein expression of the various p63 proteins
To further validate these results, we carried out immunoblot analysis on tissue lysates obtained from the snout and mouth region of WT and Brdm2/Brdm2 E15 littermates and compared them with corresponding lysates of McKeon p63 KO E15 embryos. The results show a unique and reproducible band in Brdm2/Brdm2 embryos that aligns with transfected p63Ex10-truncated ΔN(3′–10) control protein and is absent in WT and p63 KO embryos (Supplementary Figure 2). Taken together, we propose that the strong specific p63 immunostaining present in mutant skin and squamous epithelium (Figures 1 and 2) is largely because of p63Ex10-truncated ΔN(3′–10) and to a lesser extent to TA(1–10) proteins.
Next we assessed whether the Brdm2 proteins have p63γ-like transcriptional activity in vivo. Earlier, it was shown that the absence of p63 results in a strong reduction of CK14 and PERP, identifying these genes as important bona fide epidermal target genes of p63γ.25, 26, 27 CK14 is a marker of the basal layer of stratified epithelia, whereas PERP, a component of desmosomes which are specialized intraepithelial cell-cell adhesion junctions, is expressed in all cell layers of the epidermis.27 Importantly, indeed, both CK14 and PERP proteins were expressed in Brdm2/Brdm2 E15 skin at WT levels, thus confirming the in vivo functionality of Brdm2 proteins (Figure 6). In addition, our protein expression, RT-PCR and functional in vitro and in vivo data prove that the Brdm2 allele is not a null allele, as had been assumed all along in a series of earlier studies using the Brdm2 mouse model (see References2, 6, 8, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 28), but generates ΔN and TA variants of truncated p63 proteins that structurally and functionally resemble the native p63γ isoforms. Moreover, our data provide further biological and functional insight into the role of the long carboxy-terminal domain of p63 in skin and limb development.
Expression of CK14 and PERP, important bona fide epidermal target genes of p63γ, in E15 Brdm2/Brdm2 embryos. Skin of WT embryos at day E18 (a–c) and skin of Brdm2/Brdm2 embryos at day E15 (d–i) express CK14, a marker of the basal layer of stratified epithelia and PERP, a component of desmosomes. Immunoperoxidase staining
Discussion
Brdm2 mice are p63α/β isoform-specific knockout mice, gaining unexpected new importance
The skin and limb phenotype of both strains of targeted p63 mice that were generated 9 years ago to ablate the entire p63 gene is identical at birth.2, 3 Our results shed new and unexpected light on one of the two strains, the heavily used Brdm2 mice. Instead of being a global p63 KO model, we show that as a consequence of the ‘late’ insertion of the targeting vector only after exon 10, they are in fact p63α/β isoform-specific knockouts that express p63γ-like proteins at quasi WT levels. Thus, the dramatic phenotype of the mutant mice is entirely attributable to the absence of the carboxy-terminal tail present in p63α/β. With this reinterpretation, Brdm2 mice take on unexpected new importance. They positively map the p63 region required for skin and limb development to exons 11–14, which encode the functionally important SAM and post-SAM domains. Both domains are specific for p63α but not γ isoforms and absent in p53. In strong support of the importance of the SAM and post-SAM domains in ectodermal and limb development are the human ectodermal dysplasia syndromes of ankyloblepharon-ectodermal dysplasia-clefting, that is, the Hay–Wells and related syndromes. These congenital abnormalities of limbs, skin, hair, teeth, nails and palate are because of various heterozygous point mutations within the SAM and post-SAM domains of human p63.20, 21 Moreover, although ΔNp63α seems to be the predominant isoform expressed in mature stratified epithelia,1 it was unknown whether α isoforms are essential or if other isoforms can substitute for α in epithelial and limb development and homeostasis. Our results show that the carboxy-terminal domains of p63 are indeed required for the full morphogenetic program of skin and limb development, and likely also for maintenance of mature skin. The p63γ isoform thus allows to set up embryonic epidermal fate, whereas the α/β isoform is necessary for skin homeostasis. p63 γ-like isoforms cannot substitute for p63 α/β in these tissues (nor can p53 which is present in Brdm2 mice).
Furthermore, our findings make it necessary to re-interpret other earlier results using Brdm2 mice. Any phenotype that was observed with this mouse strain is because of a selective loss of long p63 isoforms. Thus, earlier studies on the development of urogenital,11, 17, 18, 29, 30, 31 gonadal19, 22, 23 and sympathetic neuronal systems14 may have addressed the biological consequences of tipping the balance between full-length and carboxy-terminally truncated p63, rather than the effect of losing all p63 isoforms.
Finally, our results reconcile at least in part some unresolved controversies in the p63 field. Different conclusions may now be linked to the use of the two different KO models2, 3 that, as shown here are not interchangeable, but represent a complete KO3 in contrast to a p63α/β isoform-specific KO.2 One example is the controversial function of p63 in skin, assigning it an essential role in proliferation of skin stem cells3, 7 versus a role in terminal differentiation.2, 16 In another example, Brdm2 heterozygotes were not prone to or even protected from spontaneous or genotoxically induced tumors compared with their WT counterparts,8, 9 which was interpreted as evidence that the p63 gene functions as an oncogene by virtue of repressing the tumor suppressive mechanisms of cellular senescence.9, 32 This result contrasted with evidence for a p63 tumor suppressor function obtained in spontaneous tumorigenesis studies with the complete p63KO strain.10 In light of our findings here, some conclusions about p63's protective role in tumorigenesis might have to be re-examined. The exclusively expressed p63γ-like proteins in Brdm2 mice have transactivation and transrepression activities comparable with p63γ and could conceivably act similar to p53-like tumor suppressors, providing cancer protection. Future studies will need to define the exact functions of the p63α/β carboxy-terminal domains in development and tumorigenesis. Brdm2 mice will be a valuable tool in this endeavour.
Materials and Methods
Mice
Genotyping of p63 Brdm2/Brdm2 mice (maintained as C57Bl6/129 50 : 50; also available as JAX Mice # 003568) across the p63 genomic/vector border confirmed the reported insertion of the targeting vector between exon 10 and exon 10′.2 The gap-repair mechanism leads to a duplication of exons 5–10. A forward primer in intron 10 (gtgttggcaaggattctgagacc, nt 187617–187639, Genbank AF533892) and a reverse primer located in the targeting vector pTV12E60 (cloning vector pIRES1neo, Genbank U89673.1) (ggaagacaatagcaggcatgct, nt 2672–3128) yields a 410 bp product which maps as indicated by the red line in Figure 1a. p63 knockout mice (Yang A et al.3) were a gift from F McKeon and G Melino.
Histological analysis and immunoperoxidase staining
Embryos were fixed in 10% buffered formalin, dehydrated, embedded in paraffin, and sectioned at 4 μm thickness. Sections were dewaxed, rehydrated, and stained in hematoxylin and eosin. Immunoperoxidase staining with p63-specific monoclonal antibody 4A4 was carried out using a standard protocol. Briefly, for antigen retrieval, deparaffinized slides were boiled in citric acid buffer (10 mM, pH 6.0) in a microwave 2 × 5 min, (alternatively for H-137 with Trilogy, Cell Marque, CA, USA), washed in PBS, quenched in 1% H2O2 for 15 min, washed and blocked with 10% serum for 20 min (Histostain SP Broad Spectrum HRP kit, ZYMED, USA). Sections were incubated with either 4A4 IgG (diluted 1 : 200 in 2% blocking serum, Santa Cruz Biotechnology, Inc. 2145, Delaware Avenue, Santa Cruz, CA, USA), H-137 (Santa Cruz, diluted 1 : 100), anti-cytokeratin 14 (CK14) (gift of Alea Mills) or anti-PERP specific antibody (gift of Laura Attardi) for 1–2 hrs at room temperature (RT) or at 4°C overnight. After washing three times in PBS, sections were incubated in biotinylated secondary antibody for 45 min at RT, washed and incubated with streptavidin-HRP before development with diaminobenzidine for 10 min, followed by dehydration and coverslipping.
Quantitative real-time reverse transcription PCR (qRT-PCR)
Total RNA was isolated from WT and Brdm2 E15 embryos or low passage MEFs using PureLink (Invitrogen, USA). RNA was reverse transcribed using the iScript cDNA Synthesis Kit (BioRad, CA, USA) according to manufacturer's instructions. IQ SYBR Green Supermix (BioRad) was used for real-time PCR applications. The primer sets are listed in Supplementary Table S1. PCR conditions were: 2 min at 95°C, followed by 40 cycles of 95°C for 15 s, 58–62°C for 30 s, and 72°C for 30–120 s. p63 transcript levels were normalized to GAPDH or loricrin (Figure 4b), calculated using the method.
Cloning of expression plasmids
cDNAs of all Brdm2 allele-derived p63 products were amplified by PCR, sequence confirmed and inserted into pCDNA3. Mouse TAp63γ, ΔNp63γ and p53 were also cloned into pcDNA3.
Immunoblots
Total cell lysates (20 μg) were separated on sodium dodecyl sulfate (SDS)-polyacrylamide gels and transferred to nitrocellulose, followed by incubation with 4A4 antibody (1 : 500 in PBS containing 5% milk powder and 0.05% Tween 20). Detection was carried out by a peroxidase-coupled secondary antibody (Jackson, USA) and ECL chemiluminescence (Pierce, USA).
Luciferase assays
H1299 cells (1 × 105 cells/well) were transfected with lipofectamine 2000 in duplicates with 100 ng of BP100luc and 50 ng of renilla reporters. To compensate for the instability of Hprt-fusion proteins, amounts of the indicated p63 expression plasmids were adjusted to achieve equal protein expression, as shown by 4A4 immunoblots (Figure 5b, d and f; 20 μg lysates each). For assessment of transactivation (Figure 5c and e), p63 plasmids were 25 ng of mouse TAp63γ and TA(1–10), and 50 ng TA-Hprt (Figure 5c), or 12.5 ng mouse ΔNp63γ and ΔN(3′–10), and 75 ng ΔN-Hprt (Figure 5e). For assessment of transrepression of TAp63γ (25 ng plasmid) or p53 (5 ng plasmid) (Figure 5g and h), ΔNp63 expression plasmids were transfected in excess (8-fold each of ΔNp63γ and ΔN(3′–10) and 48-fold of ΔN-Hprt plasmids) with respect to TAp63γ to achieve equal transrepressor protein levels (Figure 5f). Values were normalized for renilla.
Accession codes
Abbreviations
- SAM:
-
sterile α-motif
- Neo:
-
Neomycin
- KO:
-
knock out
- Hprt:
-
Hypoxanthin-Guanin-Phosphoribosyltransferase
- Puro:
-
Puromycin
- B:
-
Brdm2
- H&E:
-
Hematoxylin & Eosin
- UTR:
-
untranslated region
- WT:
-
wild-type
- MEFs:
-
mouse embryo fibroblasts
- PERP:
-
p53 apoptosis effector related to PMP-22
- SDS:
-
Sodium dodecyl sulfate
- IgG:
-
Immunoglobulin
- Ex:
-
Exon
- Luc:
-
luciferase
- RT:
-
room temperature
References
Yang A, Kaghad M, Wang Y, Gillett E, Fleming MD, Dotsch V et al. p63, a p53 homolog at 3q27–29, encodes multiple products with transactivating, death-inducing, and dominant-negative activities. Mol Cell 1998; 2: 305–316.
Mills AA, Zheng B, Wang XJ, Vogel H, Roop DR, Bradley A . p63 is a p53 homologue required for limb and epidermal morphogenesis. Nature 1999; 398: 708–713.
Yang A, Schweitzer R, Sun D, Kaghad M, Walker N, Bronson RT et al. p63 is essential for regenerative proliferation in limb, craniofacial and epithelial development. Nature 1999; 398: 714–718.
Zheng B, Mills AA, Bradley A . A system for rapid generation of coat color-tagged knockouts and defined chromosomal rearrangements in mice. Nucleic Acids Res 1999; 27: 2354–2360.
McKeon F . p63 and the epithelial stem cell: more than status quo? Genes Dev 2004; 18: 465–469.
Koster MI, Kim S, Mills AA, DeMayo FJ, Roop DR . p63 is the molecular switch for initiation of an epithelial stratification program. Genes Dev 2004; 18: 126–131.
Senoo M, Pinto F, Crum CP, McKeon F . p63 Is essential for the proliferative potential of stem cells in stratified epithelia. Cell 2007; 129: 523–536.
Perez-Losada J, Wu D, DelRosario R, Balmain A, Mao JH . p63 and p73 do not contribute to p53-mediated lymphoma suppressor activity in vivo. Oncogene 2005; 24: 5521–5524.
Keyes WM, Vogel H, Koster MI, Guo X, Qi Y, Petherbridge KM et al. p63 heterozygous mutant mice are not prone to spontaneous or chemically induced tumors. Proc Natl Acad Sci USA 2006; 103: 8435–8440.
Flores ER, Sengupta S, Miller JB, Newman JJ, Bronson R, Crowley D et al. Tumor predisposition in mice mutant for p63 and p73: evidence for broader tumor suppressor functions for the p53 family. Cancer Cell 2005; 7: 363–373.
Cheng W, Jacobs WB, Zhang JJ, Moro A, Park JH, Kushida M et al. DeltaNp63 plays an anti-apoptotic role in ventral bladder development. Development 2006; 133: 4783–4792.
Davison TS, Vagner C, Kaghad M, Ayed A, Caput D, Arrowsmith CH . p73 and p63 are homotetramers capable of weak heterotypic interactions with each other but not with p53. J Biol Chem 1999; 274: 18709–18714.
Freedman DA, Epstein CB, Roth JC, Levine AJ . A genetic approach to mapping the p53 binding site in the MDM2 protein. Mol Med 1997; 3: 248–259.
Jacobs WB, Govoni G, Ho D, Atwal JK, Barnabe-Heider F, Keyes WM et al. p63 is an essential proapoptotic protein during neural development. Neuron 2005; 48: 743–756.
Keyes WM, Wu Y, Vogel H, Guo X, Lowe SW, Mills AA . p63 deficiency activates a program of cellular senescence and leads to accelerated aging. Genes Dev 2005; 19: 1986–1999.
Koster MI, Dai D, Marinari B, Sano Y, Costanzo A, Karin M et al. p63 induces key target genes required for epidermal morphogenesis. Proc Natl Acad Sci USA 2007; 104: 3255–3260.
Kurita T, Cunha GR, Robboy SJ, Mills AA, Medina RT . Differential expression of p63 isoforms in female reproductive organs. Mech Dev 2005; 122: 1043–1055.
Kurita T, Medina RT, Mills AA, Cunha GR . Role of p63 and basal cells in the prostate. Development 2004; 131: 4955–4964.
Livera G, Petre-Lazar B, Guerquin MJ, Trautmann E, Coffigny H, Habert R . p63 null mutation protects mouse oocytes from radio-induced apoptosis. Reproduction 2008; 135: 3–12.
McGrath JA, Duijf PH, Doetsch V, Irvine AD, de Waal R, Vanmolkot KR et al. Hay-Wells syndrome is caused by heterozygous missense mutations in the SAM domain of p63. Hum Mol Genet 2001; 10: 221–229.
Moll UM, Slade N . p63 and p73: roles in development and tumor formation. Mol Cancer Res 2004; 2: 371–386.
Petre-Lazar B, Livera G, Moreno SG, Trautmann E, Duquenne C, Hanoux V et al. The role of p63 in germ cell apoptosis in the developing testis. J Cell Physiol 2007; 210: 87–98.
Petre-Lazar B, Moreno SG, Livera G, Duquenne C, Habert R, Coffigny H . p63 expression pattern in foetal and neonatal gonocytes after irradiation and role in the resulting apoptosis by using p63 knockout mice. Int J Radiat Biol 2006; 82: 771–780.
Reis-Filho JS, Milanezi F, Amendoeira I, Albergaria A, Schmitt FC . Distribution of p63, a novel myoepithelial marker, in fine-needle aspiration biopsies of the breast: an analysis of 82 samples. Cancer 2003; 99: 172–179.
Medawar A, Virolle T, Rostagno P, de la Forest-Divonne S, Gambaro K, Rouleau M et al. DeltaNp63 is essential for epidermal commitment of embryonic stem cells. PLoS ONE 2008; 3: e3441.
Boldrup L, Coates PJ, Gu X, Nylander K . DeltaNp63 isoforms regulate CD44 and keratins 4, 6, 14 and 19 in squamous cell carcinoma of head and neck. J Pathol 2007; 213: 384–391.
Ihrie RA, Marques MR, Nguyen BT, Horner JS, Papazoglu C, Bronson RT et al. Perp is a p63-regulated gene essential for epithelial integrity. Cell 2005; 120: 843–856.
Keyes WM, Mills AA . p63: a new link between senescence and aging. Cell Cycle 2006; 5: 260–265.
Kurita T, Cunha GR . Roles of p63 in differentiation of Mullerian duct epithelial cells. Ann NY Acad Sci 2001; 948: 9–12.
Kurita T, Mills AA, Cunha GR . Roles of p63 in the diethylstilbestrol-induced cervicovaginal adenosis. Development 2004; 131: 1639–1649.
Suzuki K, Haraguchi R, Ogata T, Barbieri O, Alegria O, Vieux-Rochas M et al. Abnormal urethra formation in mouse models of split-hand/split-foot malformation type 1 and type 4. Eur J Hum Genet 2008; 16: 36–44.
Guo X, Mills AA . p63, cellular senescence and tumor development. Cell Cycle 2007; 6: 305–311.
Acknowledgements
We thank A Mills for providing Brdm2 mice. This work was supported by the National Cancer Institute (UMM), the Carol Baldwin Breast Cancer Award (UMM), the Deutsche Krebshilfe (UMM), and the Deutsche Forschungsgemeinschaft (MD).
Author information
Authors and Affiliations
Corresponding author
Additional information
Edited by G Melino
Supplementary Information accompanies the paper on Cell Death and Differentiation website (http://www.nature.com/cdd)
Supplementary information
Rights and permissions
About this article
Cite this article
Wolff, S., Talos, F., Palacios, G. et al. The α/β carboxy-terminal domains of p63 are required for skin and limb development. New insights from the Brdm2 mouse which is not a complete p63 knockout but expresses p63 γ-like proteins. Cell Death Differ 16, 1108–1117 (2009). https://doi.org/10.1038/cdd.2009.25
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/cdd.2009.25
Keywords
This article is cited by
-
Late cornified envelope 1C (LCE1C), a transcriptional target of TAp63 phosphorylated at T46/T281, interacts with PRMT5
Scientific Reports (2018)
-
A Mutation in TP63 Causing a Mild Ectodermal Dysplasia Phenotype
Journal of Investigative Dermatology (2014)
-
p63 is a prosurvival factor in the adult mammary gland during post-lactational involution, affecting PI-MECs and ErbB2 tumorigenesis
Cell Death & Differentiation (2014)
-
ΔNp63 regulates select routes of reprogramming via multiple mechanisms
Cell Death & Differentiation (2013)
-
MicroRNA-203 contributes to skin re-epithelialization
Cell Death & Disease (2012)