Abstract
Mitochondria are essential organelles that are responsible for cellular energy production and cell death in response to various stimuli. Although C-terminal binding protein (CtBP) functions as a metabolic sensor in transcriptional corepressor complex, it is unclear whether CtBP controls gene transcription in response to metabolic stress. In this study, we found that CtBP represses Bcl-2-associated X protein (Bax) transcription in glucose-rich media by binding to the E-box region of the Bax promoter. Glucose withdrawal leads to the dissociation of CtBP from the Bax promoter and significant changes of the histone codes in the Bax promoter. CtBP knockout increases Bax transcription, ablates mitochondrial morphology and reduces mitochondrial activities. Ectopic expression of CtBP or knockdown of Bax in ctbp-knockout cells recovers mitochondrial morphology and function, suggesting that CtBP functions as a metabolic sensor that maintains mitochondrial activities. Our findings provide insights into how the intracellular energy level is reflected into gene transcription involved in mitochondrial morphology and function.
Similar content being viewed by others
Main
C-terminal binding protein (CtBP) was originally identified as an E1A-binding protein1 and as a transcriptional corepressor that associates with DNA-specific transcriptional repressors in development and tumorigenesis.2, 3 CtBP is also known to repress gene transcription in a histone deacetylase (HDAC)-dependent4, 5 and -independent manner.6, 7, 8 CtBP belongs to a family of 2-hydroxyacid dehydrogenases; CtBP senses metabolic changes by differentially binding to NADH and controls the transcription of downstream genes through monomer–dimer transition.6, 9, 10 Thus, CtBP serves as a metabolic sensor in transcriptional corepressor complex.11, 12, 13
Glucose is the main source of cellular carbohydrate and is a metabolic fuel for activating metabolic pathways in all mammalian cells. Thus, glucose homeostasis is the most tightly regulated function in the body. Meanwhile, metabolic stress, such as imbalance of glucose homeostasis, triggers a variety of cellular responses including cellular growth and gene expression. Most cells efficiently produce high-energy molecules in the mitochondria through the TCA cycle and oxidative respiration. Moreover, mitochondria mediate cell death by regulating the balance between pro-apoptotic and antiapoptotic proteins. Therefore, metabolic stresses such as glucose deprivation are likely to affect mitochondrial function. However, the detailed mechanisms of these effects remain unclear.
According to the primary function of glucose in living cells, we first investigated the effect of glucose deprivation on cellular activities. In this study, we found that mitochondria-dependent apoptosis occurred in response to glucose deprivation and this apoptosis was mediated by the activation of Bcl-2-associated X protein (Bax) gene expression. Similar to the earlier findings that CtBP serves as a metabolic sensor and a transcriptional corepressor,11, 12, 13 we also found that CtBP associated with and repressed Bax promoter and was dissociated from the promoter in response to glucose deprivation. In addition, mitochondrial morphological changes and decreased mitochondrial activities were observed in ctbp-knockout cells, and the knockdown of Bax in ctbp-knockout cells recovered the mitochondrial morphology and function. Our results showed that the transcriptional corepressor CtBP1 maintains mitochondrial activities by regulating Bax transcription dependent on the cellular glucose level.
Results
Glucose deprivation induces apoptotic cell death
As glucose is the main source of energy production in living cells, we first investigated whether the imbalance of glucose homeostasis triggers apoptotic cell death. Prolonged incubation of cells in low-glucose (1 mM) media, as well as in glucose-free (0 mM) media, induced apoptotic cell death (Figure 1a and b). Low ATP level and reduced mitochondrial membrane potential were observed under low-glucose or glucose-free conditions, indicating that apoptosis induced by low glucose is mitochondria dependent (Supplementary Figure 1). Bcl-2 family members play a critical role in the regulation of the mitochondrial pathway that leads to apoptotic cell death by directly targeting mitochondria.14 This led us to examine the pro-apoptotic regulators that affect mitochondria under low-glucose conditions. To determine which pro-apoptotic regulator affects mitochondrial activities in low-glucose conditions, we screened mitochondria-targeted pro-apoptotic proteins induced by low glucose. We found that glucose deprivation significantly induced Bax expression at the transcriptional level in a time- (Supplementary Figure 2) and dose-dependent manner (Figure 1c and d). In addition, the glucose transporter-specific inhibitor, cytochalasin B, significantly increased the mRNA level of Bax (Figure 1e).
Mitochondria-dependent apoptotic cell death in HEK 293 cells. (a) Glucose deprivation induces apoptotic cell death. HEK 293 cells were cultured in medium containing the indicated concentration of D-glucose for 48 h and cell death was measured by flow cytometry. (b) A graph of the percentage of apoptotic cell death obtained in (a). (c) Bax protein was increased in low-glucose conditions. HEK 293 cells were cultured in medium containing the indicated concentration of D-glucose for 12 h. Each sample was subjected to SDS-PAGE and immunoblotted with anti-CtBP1, anti-Bax (Santa Cruz Biotechnology) and anti-β-actin antibodies (Sigma). (d and e) The quantitative analysis of Bax mRNA expression. HEK 293 cells were stimulated with different glucose concentrations (d) or the glucose transporter-specific inhibitor, cytochalasin B, (e) for 12 h. Relative mRNA levels were analyzed by quantitative real-time PCR
Ctbp-knockout cells are more sensitive to apoptosis by glucose deprivation
As a metabolic sensor, CtBP can regulate cellular activities in response to the change of NAD+/NADH.9, 10, 15 Also, CtBP acts as a negative regulator of apoptosis.16, 17, 18, 19 However, the precise mechanism by which CtBP regulates apoptosis at the transcriptional level remains unclear. To investigate whether CtBP is involved in apoptosis induced by low glucose, we used ctbp mouse embryonic fibroblast (MEF) cells.20 We incubated both ctbp+/− and ctbp−/− cells in glucose-free media. We then stained the cells with trypan blue (TB) and counted living cells (TB-negative cells) at the indicated time points (Figure 2a). After 48 h of incubation, ctbp−/− cells appeared more sensitive to apoptosis than ctbp+/− cells. In addition, we confirmed that ctbp−/− cells died more rapidly than ctbp+/− cells by labeling the cells with annexin-V and propidium iodide after the 48h of incubation in low-glucose conditions (Figure 2b and c).
CtBP has an anti-apoptotic function in low-glucose-induced apoptosis. (a) Detection of cell death by cell counting during glucose deprivation. ctbp MEF cells (3.5 × 105) were grown on 100-mm culture dishes for the indicated time in glucose-free medium. Cells were counted after staining with trypan blue solution. Cell number: n=log 2 Ct/C0 (Ct: cell number at time point, C0: initial cell number). (b) ctbp−/− MEF cells were more sensitive to apoptosis than ctbp+/− MEF cells upon glucose deprivation. ctbp MEF cells were cultured in the indicated glucose concentration for 48 h. Cells were harvested and incubated with both annexin-V and propidium iodide. Apoptotic cell death was assessed by flow cytometry. (c) A graph of results obtained in (b). Error bars in (c) denote S.D. (n=3)
To further define the effect of CtBP on apoptosis, we established stable MEF cells that express wild-type CtBP1 (ctbp−/− (WT)) (Supplementary Figure 3) and investigated the apoptosis of ctbp−/− (WT) cells in low-glucose conditions. The restoration of CtBP1 in ctbp−/− cells significantly retarded apoptosis after 48 h of incubation under glucose-free conditions (Figure 2b and c), indicating that CtBP plays a role in preventing cell death during glucose deprivation.
CtBP is a transcriptional corepressor of Bax gene promoter
We found that low glucose triggered the expression of the Bax gene and that CtBP has an anti-apoptotic function in low-glucose conditions (Figures 1c and 2). Moreover, ctbp-knockout cells were reported to be hypersensitive to apoptosis, and several pro-apoptotic genes are highly induced in ctbp-knockout cells.16 Therefore, we postulated that CtBP1 represses Bax transcription by its association with Bax promoter under normal glucose conditions. We first confirmed that Bax is highly expressed in ctbp−/− cells (Figure 3a). When ctbp MEF cells were transfected with a Bax promoter-driven luciferase reporter gene, the luciferase activity was highly detected in ctbp−/− cells (Figure 3b). In addition, the ectopic expression of CtBP1 in ctbp−/− cells and HeLa cells reduced the luciferase activity, confirming that CtBP represses Bax at the transcriptional level (Figure 3c). As there are several reported transcription factor-binding sites in the Bax promoter (Figure 3d),21, 22, 23 we examined the CtBP1-binding region in the human Bax promoter by chromatin immunoprecipitation (ChIP) assays. ChIP analysis revealed that CtBP1 preferentially binds to the E-box region conserved in the proximal promoters of both human and mouse Bax gene (Figure 3e, Supplementary Figure 4), consistent with the earlier findings that CtBP1 associates with several E-box-binding transcription factors to repress its target gene.3
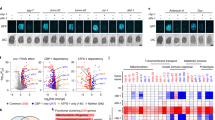
CtBP controls Bax transcription upon glucose deprivation. (a) Bax was highly expressed in ctbp−/− MEF cells. Meanwhile, the expression of Bax was reduced in ctbp−/− (WT) MEF cells. Stably expressed myc-CtBP1 was detected with either anti-CtBP1 antibody or anti-myc antibody. (b) Bax promoter-driven luciferase activity was increased in ctbp−/− MEF cells compared with that in ctbp+/− MEF cells. (c) Ectopically expressed CtBP1 decreased Bax reporter gene expression in both ctbp−/− MEF and HeLa cells. (d) Diagram of human Bax promoter. P1 contains NF-κB-binding site, P2 contains both NF-κB- and p53-binding sites, P3 contains the interspace between the p53-binding site and the E-box region, and P4 contains four E-box regions. (e) ChIP analysis showed that CtBP1 preferentially bound to the E-box region of the Bax promoter. (f–h) Dynamic changes of chromatin-associated proteins in human Bax promoter upon glucose deprivation. ChIP analysis was performed using the indicated antibodies. Error bars in (b) and (c) denote S.D. (n=3)
As glucose deprivation affects Bax transcription and CtBP1 represses Bax transcription in normal glucose conditions, we investigated whether glucose deprivation changes the repressive function of CtBP1 and the histone codes in the Bax promoter for activating Bax transcription. When glucose was depleted from the media, CtBP1 dissociated from the E-box region in the Bax promoter (Figure 3f). At the same time, transcriptionally active histone codes (acetylated H3-K9 and trimethylated and dimethylated H3-K4) were significantly increased around the E-box region, whereas transcriptionally repressive histone codes (trimethylated and dimethylated H3-K9) were decreased (Figure 3g). In addition, these changes in the histone codes were mediated by PCAF, JARID1C,24 HDAC4 and JMJD2A,25 respectively (Figure 3h), indicating that CtBP1 associates with histone-modifying enzymes and is involved in the repression of Bax transcription in glucose-rich conditions. Also, the dissociation of CtBP1 from the Bax promoter de-represses Bax transcription during glucose deprivation.
CtBP maintains the mitochondrial morphology and function
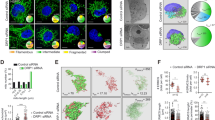
Mitochondrial fusion and fission are required for mitochondrial dynamics.26 As Bax is a mitochondrial protein involved in mitochondrial dynamics27, 28 and is increased in ctbp-knockout cells (Figure 3a), we hypothesized that ctbp knockout would affect mitochondrial morphology. To address this, we cocultured MEF cells of ctbp+/− and ctbp−/− in the same plate and compared their mitochondrial morphology by staining with Red-Mitotracker (Figure 4a) and quantitative analysis (Supplementary Figure 5). Interestingly, mitochondrial elongation was observed in ctbp-knockout cells, whereas mitochondria in ctbp+/− cells appeared spotted as normal. In addition, we observed that the stable expression of CtBP1 in ctbp−/− cells apparently recovered mitochondrial morphology to normal (Supplementary Figure 6a). This result indicates that the loss of CtBP leads to mitochondrial morphological changes. Moreover, recent studies have shown that mitochondrial morphological changes, i.e., mitochondrial fission and cristae reorganization, occur during apoptosis.29, 30 We thus investigated mitochondrial morphology during glucose deprivation. When HeLa cells were cultured in glucose-free media, mitochondria appeared elongated at a relatively early (12 h) time point, but ultimately became fragmented with long-term culture (48 h) (Supplementary Figure 7). As those of HeLa cells, mitochondria in MEF cells were also observed as elongated tubules during 16 h of incubation in glucose-free media (Figure 4b, Supplementary Figure 6b). Next, we further investigated the mitochondrial ultrastructure by transmission electron microscopy (TEM). TEM analysis showed that mitochondria in ctbp−/− cells were swollen and their mitochondrial cristae were nearly hollow, even in glucose-rich conditions, compared with the mitochondria of ctbp+/− cells, which were filled with densely packed cristae (Figure 4c and d). Meanwhile, the stable expression of CtBP1 in ctbp−/− cells recovered densely packed cristae in mitochondria (Supplementary Figure 8). Furthermore, when glucose was removed from the media, the mitochondria of most cells swelled and their cristae became hollow (Figure 4c and d, Supplementary Figure 8). We next examined the effect of ctbp knockout on mitochondrial function. Interestingly, ctbp knockout decreased the cellular ATP level, oxygen consumption rate and mitochondrial membrane potential even in glucose-rich media. Meanwhile, the stable expression of CtBP1 in ctbp−/− MEF cells restored these values to the levels observed in ctbp+/− cells (Figure 4e–g). These findings suggest that CtBP is very important for maintaining normal mitochondrial morphology and function.
CtBP affects mitochondrial morphology. (a and b) Confocal microscopy showed that ctbp knockout triggers mitochondrial elongation (a) and that glucose deprivation (16 h) elongates mitochondria (b). Both ctbp+/− and ctbp−/− MEF cells were cocultured and distinguished by staining with anti-CtBP1 antibody followed by FITC-conjugated secondary antibody. Mitochondria were stained with Red-Mitotracker. Images are shown at × 1000 magnification. (c and d) Ultrastructure of mitochondrial cristae in ctbp MEF cells. TEM analysis showed that ctbp knockout causes swollen mitochondria and hollow cristae. Images are shown at × 25 000 magnification (N: nucleus; M: mitochondria). (e) Measurement of cellular ATP generation by mitochondria in ctbp MEF cells. The level of ATP in lysates of 3 × 105 cells was detected using the ATP assay kit (Roche). *P=0.000764; **P=0.005406. (f) Oxygen consumption rate (OCR) in ctbp MEF cells. Cells (1 × 106) were collected and resuspended in culture medium. The oxygen consumption rates of cells were monitored using a Clark-type oxygen electrode (Instech). *P=0.000139; **P=0.000369. (g) Mitochondrial membrane potential was decreased in ctbp-knockout cells. Mitochondria were stained with DiOC6(3) and fluorescence intensity was measured by flow cytometry. *P=0.000142; **P=0.000053. Error bars in (e), (f) and (g) denote S.D. ((e): n=6; (f): n=8; (g): n=3)
Role of Bax in mitochondrial morphology and function
Swollen mitochondria with hollow cristae in ctbp-knockout cells appeared very similar to the mitochondria under glucose-free conditions. Moreover, as CtBP controlled Bax transcription in response to glucose deprivation, CtBP may regulate mitochondrial morphology and function by controlling Bax gene expression. We thus investigated the location of Bax protein in ctbp MEF cells. Glucose deprivation in ctbp+/− cells triggered the translocation of Bax from cytosol to mitochondria (Figure 5a). Intriguingly, a significant amount of Bax in ctbp−/− cells was located in mitochondria even in glucose-rich media (Figure 5a). Next, to determine whether the mitochondrial morphological and structural changes are due to the translocation of Bax by loss of CtBP and/or glucose deprivation, we knocked down the Bax mRNA level by infection with lentivirally expressed shBax in ctbp−/− cells (Figure 5b). As expected, the knockdown of Bax restored mitochondrial morphology based on confocal microscopy findings (Figure 5c–e). In addition, the hollow cristae in swollen mitochondria reverted to the densely packed cristae observed in normal mitochondria (Figure 5f). Moreover, the knockdown of Bax reduced mitochondrial length to normal (Figure 5g).
The role of Bax in mitochondrial morphology and function. (a) Translocation of Bax protein during glucose deprivation. After 16 h of glucose deprivation, ctbp MEF cells were fractionated by syringe filtration. Each fraction was resolved by SDS-PAGE and probed with the indicated antibodies. Anti-Hsp60 and anti-tubulin antibodies (Santa Cruz Biotechnology) were used for fractionation controls of mitochondria and cytosol, respectively. (b) Knockdown of Bax reduced both the mRNA and the protein levels of Bax. ctbp−/− MEF cells were infected with either lentivirally expressed shBax or scrambled shRNA as a control. (c and d) The recovery of mitochondrial elongation by knockdown of Bax in ctbp−/− MEF cells. Mitochondrial elongation by the induction of Bax mRNA in ctbp−/− cells was restored by knockdown of Bax. (e) The number of mitochondria counted in (c) and (d) (n=100). (f) The ultrastructure of mitochondrial cristae in ctbp−/− cells infected with shRNA. Mitochondrial cristae in ctbp−/− MEF cells, which stably express shRNA for knockdown of Bax, were observed by TEM. Swollen mitochondrial hollow cristae structure was restored to normal by knockdown of Bax. (g) Measurement of mitochondrial length. Mitochondrial length in either scrambled shRNA- or shBax-infected ctbp−/− cells was measured using TEM. (h–j) Recovery of cellular ATP generation, oxygen consumption rate and mitochondrial membrane potential by knockdown of Bax in ctbp−/− cells. *P=0.00262; *P=0.02416; *P=0.036706 in (h), (i) and (j), respectively. Error bars in (h), (i) and (j) denote S.D. ((h): n=4; (i): n=4; (j): n=3). (k) Reduction of apoptosis in Bax-knockdown ctbp−/− MEF cells in low-glucose conditions. ctbp−/− cells infected with either control shRNA or shBax were cultured in medium containing the indicated concentration of D-glucose for 48 h. Apoptosis was measured by flow cytometry after staining cells with both annexin-V and propidium iodide
To delineate the role of Bax in mitochondrial function in ctbp−/− cells, we measured the ATP level, oxygen consumption rate and mitochondrial membrane potential in Bax-knockdown ctbp−/− cells (Figure 5h–j). Bax knockdown in ctbp−/− cells increased cellular ATP level, oxygen consumption rate and mitochondrial membrane potential, indicating that the expression level of Bax controls metabolic processes such as ATP production and oxygen consumption in mitochondria. Finally, we also observed that knockdown of Bax significantly reduced apoptotic cell death in ctbp−/− cells under low-glucose conditions (Figure 5k). Bax reportedly colocalizes with both mitochondrial fusion and fission sites and Bax induces mitochondrial fusion in non-apoptotic cells.26, 27, 28 These reports are consistent with our findings that Bax overexpression in ctbp−/− MEF cells and Bax induction in ctbp+/− cells during glucose deprivation lead to the change of mitochondrial morphology and function.
Discussion
Glucose is the primary carbohydrate source in living cells and is converted to many other metabolites that are necessary for cellular activities. Metabolic stress from changes in glucose influx influences a variety of cellular events. Mitochondria play an essential role in maintaining cellular activities by regulating the balance of energy. Therefore, cellular glucose concentration is closely related to mitochondrial function. In this study, we found that mitochondria-dependent apoptosis occurs under low-glucose conditions and this apoptosis is mediated by Bax gene transcription. In low-glucose conditions, Bax gene transcription is induced by the release of CtBP from its promoter region. On the basis of the localization of Bax in mitochondria,27 it is possible that the changes in mitochondrial morphology and function are mediated by an increased Bax gene transcription. Therefore, as a metabolic sensor and transcriptional corepressor, CtBP maintains mitochondrial activities by regulating Bax transcription. Recently, CtBP was identified as a transcriptional corepressor of PRDM16 for the selective repression of white fat genes.31 However, little is known about the mechanism by which CtBP controls the cellular activities by the change of energy metabolism at the transcriptional level. In this study, we found that CtBP controls Bax transcription in response to glucose deprivation, and Bax induction by the functional loss of CtBP leads to intermediate mitochondrial elongation, hollow-cristae formation in swollen mitochondria and a more rapid mitochondrial fragmentation upon cell death. Opa1 is also able to control apoptotic cristae remodeling.32, 33 Furthermore, Bax associates with Opa1 in the junction between the inner and outer mitochondrial membranes, whereas knockdown of Bax blocks the release of Opa1 from mitochondria.34 It is therefore possible that the decrease of mitochondrial activities by knockout of ctbp occurs by a collaboration between Bax and Opa1.
Notably, the loss of CtBP affects mitochondrial activities through structural changes and facilitates apoptosis in response to glucose deprivation. Given the structural and functional activities of CtBP, our findings provide new insights into both transcriptional regulation by metabolic stress and its effects on cellular activities.
Materials and Methods
Cell culture, transfection and stable cell establishment
HEK 293 and HeLa cells were grown in Dulbecco's modified Eagle medium (DMEM) containing 10% (v/v) fetal bovine serum and penicillin–streptomycin. CtBP MEF cells were grown in DMEM containing 10% FBS, penicillin–streptomycin and 2 mM L-glutamine. Transient transfection was performed using Lipofectamine reagent (Invitrogen). To generate ctbp−/− (WT) cells, ctbp−/− cells were transfected with pcDNA4-CtBP1-myc plasmid and selected in medium containing 500 μg/ml of Zeocin (Invitrogen). For cell culture in different glucose concentrations, cells were cultured in glucose-free medium containing 10% dialyzed FBS (Invitrogen) and the indicated D(+)-glucose (Sigma) concentrations.
RNA purification and quantitative real-time PCR
RNA was purified using Trizol reagent (Invitrogen). Quantitative RT-PCR was performed as described by the manufacturer (Finnzymes). Briefly, 2 × qPCR master mix was diluted to 1 × reaction solution with pre-designed primers (Supplementary Table 1), cDNA and ultra-purified water. The accumulation of specific products in a reaction was continuously monitored by iCycler Real-Time PCR (Bio-Rad).
Lentivirus-based shRNA
A total of 6 × 106 293 FT cells were transfected with either mouse Bax shRNA (5′-CCGGCTACAGGGTTTCATCCAGGATCTCGAGATCCTGGATGAAACCCTGTAGTTTTT-3′) (Open Biosystems) or control shRNA (Sigma) containing lentivirus packaging plasmids using Lipofectamine 2000 reagent (Invitrogen). After 72 h, virus-containing supernatants were harvested and centrifuged at 1500 r.p.m. for 10 min to pellet cell debris. To test lentivirus-containing mBax shRNA, when ctbp−/− cells were 30% confluent, they were treated with 1 ml of lentiviral stock. After 48 h, cells were selected for 2 weeks in 2 μg/ml puromycin.
Chromatin immunoprecipitation assay
HEK 293 cells were fixed with 1% (v/v) formaldehyde. Cells were lysed with lysis buffer (5 mM PIPES (pH 8.0), 85 mM KCl and 0.5% (v/v) NP-40). After centrifugation at 5000 r.p.m., nuclear pellets were resuspended in sonication buffer (50 mM Tris-Cl (pH 8.0), 10 mM EDTA and 1% SDS). To shear DNA, lysates were sonicated and centrifuged for 10 min. Lysates were transferred to fresh tubes and diluted 10-fold in IP buffer (16.7 mM Tris-Cl (pH 8.0), 167 mM NaCl, 0.01% (w/v) SDS, 1% (v/v) Triton X-100 and 1.2 mM EDTA). This ‘chromatin solution’ (25–100 μg) was incubated with anti-CtBP1, anti-dimethyl H3K4, anti-trimethyl H3K4, anti-dimethyl H3K9, anti-trimethyl H3K9 (Upstate), anti-acetyl H3K9, anti-Histone H3, anti-HDAC4, anti-JARID1C, anti-JMJD2A (Abcam), anti-PCAF or anti-rabbit normal IgG (Santa Cruz Biotechnology) overnight. The next day, protein A/G PLUS agarose was added and incubated for 2 h; the resulting immune complexes were centrifuged and washed with washing buffer (20 mM Tris-Cl (pH 8.0), 150 mM NaCl, 0.5% (w/v) SDS, 1% Triton X-100 and 2 mM EDTA) three times and with washing buffer containing 300 mM NaCl two times. Immune complexes were then eluted by adding 250 μl of elution buffer (1% (w/v) SDS and 0.1 M NaHCO3) two times. Then 20 μl of 5 M NaCl was added and crosslinks between DNA and proteins were reversed at 65°C by overnight incubation. The next day, free DNA was precipitated with ethanol. Purified DNA was quantified by PCR using pre-designed primer sets (Supplementary Table 2).
Fluorescence microscopy
HeLa and ctbp MEF cells were grown on coverslips for 16 h. Cells were stained with 50 nM MitoTracker Red CMXRos (Molecular Probes, M-7512) for 30 min at 37°C. After three washes with PBS, cells were fixed with 4% (w/v) paraformaldehyde and blocked with 2% (w/v) BSA in PBS. Cells were then labeled with anti-CtBP1 (Santa Cruz Biotechnology) or anti-myc (Covance) antibody and with FITC-conjugated goat anti-mouse antibody (Jackson Immunoresearch). Labeled cells were visualized with LSM 510 META (Carl Zeiss).
FACS analysis
To analyze apoptotic cell death, HEK 293 and ctbp MEF cells were harvested and washed three times with PBS. Cells were then incubated with FITC annexin-V (BD Pharmingen) and propidium iodide (Sigma) for 15 min and analyzed by flow cytometry (Beckman Coulter). To measure mitochondrial membrane potential, cells were washed with PBS and incubated with 80 nM DiOC6(3) (Molecular Probes) for 15 min at 37°C. DiOC6(3) fluorescence was measured immediately by flow cytometry.
Measurement of cellular ATP and oxygen consumption rate
Cellular concentrations of ATP were measured using the ATP Bioluminescence Assay Kit CLS II (Roche). Briefly, after cells were subcultured for 16 h, 3 × 105 cells were collected and resuspended in 500 μl of ATP assay dilution buffer (100 mM Tris (pH 7.75) and 4 mM EDTA). Samples were boiled for 2 min and centrifuged for 5 min at 1000 × g. Supernatants were collected and 50 μl samples were analyzed in the luminometer (Perkin Elmer). For the measurement of oxygen consumption rate, cells were trypsinized and centrifuged at 1500 r.p.m. for 2 min at 4°C. Cells were washed with PBS and then 1 × 106 cells were collected and resuspended in 200 μl of culture medium. The oxygen consumption rate of the cells was monitored continuously for 15 min using a Clark-type oxygen electrode (Instech Laboratories).
Transmission electron microscope
Cells were washed with PBS and fixed overnight in a mixture of 2.5% (w/v) glutaraldehyde (Sigma) and 2% (w/v) paraformaldehyde in 0.1 M phosphate buffer (pH 7.2). Cells were post-fixed for 90 min in 2% (w/v) osmium tetroxide in 0.1 M phosphate buffer, pH 7.2, at room temperature. The samples were then washed briefly with deuterated H2O, dehydrated throughout a graded 50, 60, 70, 80, 90, 95 and 100% ethanol (χ2) series, infiltrated with propylene oxide (Acros Organics) and EPON epoxy resin mix, and finally embedded with epoxy resin. Samples were then loaded into capsules and polymerized at 38°C for 12 h and 60°C for 48 h. Sections for light microscopy were cut at 1.0 nm and stained with 1% (w/v) toluidine blue for 45 s on a hot plate at 80°C. Thin sections were cut using an ultramicrotome (RMC MT-XL) and collected on a copper grid. Appropriate areas for thin sectioning were cut at 65 nm and stained with saturated 4% (w/v) uranyl acetate and 4% (w/v) lead citrate before examination with a transmission electron microscope (JEM-1400, Japan) at 80 kV.
Abbreviations
- CtBP:
-
C-terminal binding protein
- Bax:
-
Bcl-2-associated X protein
- ChIP:
-
chromatin immunoprecipitation
- MEF:
-
mouse embryonic fibroblast
References
Schaeper U, Boyd JM, Verna S, Uhlmann E, Subramanian T, Chinnadurai G . Molecular cloning and characterization of a cellular phosphoprotein that interacts with a conserved C-terminal domain of adenovirus E1A involved in negative modulation of oncogenic transformation. Proc Natl Acad Sci USA 1995; 92: 10467–10471.
Chinnadurai G . Transcriptional regulation by C-terminal proteins. Int J Biochem Cell Biol 2007; 39: 1593–1607.
Berman LM, Blayers JP . C-terminal binding proteins: emerging roles in survival and tumorigenesis. Apoptosis 2006; 11: 879–888.
Dressel U, Bailey PJ, Wang SC, Downes M, Evans RM, Muscat GE . A dynamic role for HDAC7 in MEF2-mediated differentiation. J Biol Chem 2001; 276: 17007–17013.
Subramanian T, Chinnadurai G . Association of class I histone deacetylase with transcriptional corepressor CtBP. FEBS Lett 2003; 540: 255–258.
Kim JH, Cho EJ, Kim ST, Youn HD . CtBP represses p300-mediated transcriptional activation by direct association with its bromodomain. Nat Struct Mol Biol 2005; 12: 423–428.
Meloni AR, Lai CH, Yao TP, Nevins JR . A mechanism of COOH-terminal binding protein-mediated repression. Mol Cancer Res 2005; 3: 575–583.
Senyuk V, Sinha KK, Nucifora G . Corepressor CtBP1 interacts with and specifically inhibits CBP activity. Arch Biochem Biophys 2005; 441: 168–173.
Fjeld CC, Birdsong WT, Goodman RH . Differential binding of NAD+ and NADH allows the transcriptional corepressor carboxy-terminal binding protein to serve as a metabolic sensor. Proc Natl Acad Sci USA 2003; 100: 9202–9207.
Garriga-Canut M, Schoenike B, Qazi R, Bergendahl K, Daley TJ, Pfender RM et al. 2-deoxy-D-glucose reduces epilepsy progression by NRSF-CtBP-dependent metabolic regulation of chromatin structure. Nat Neurosci 2006; 9: 1382–1387.
Zhang Q, Piston DW, Goodman RH . Regulation of corepressor function by nuclear NADH. Science 2002; 295: 1895–1897.
Kumar V, Carlson JE, Ohgi KA, Edwards TA, Rose DW, Escalante CR et al. Transcription corepressor CtBP is an NAD+-regulated dehydrogenase. Mol Cell 2002; 10: 857–869.
Shi Y, Sawada J, Sui G, Affar el B, Whetstine JR, Lan F et al. Coordinated histone modifications mediated by a CtBP co-repressor complex. Nature 2003; 422: 735–738.
Green DR, Reed JC . Mitochondria and apoptosis. Science 1998; 281: 1309–1312.
Zhang Q, Wang SY, Nottke AC, Rocheleau JV, Piston DW, Goodman RH . Redox sensor CtBP mediates hypoxia-induced tumor cell migration. Proc Natl Acad Sci USA 2006; 103: 9029–9033.
Grooteclaes M, Deveraux Q, Hildebrand J, Zhang Q, Goodman RH, Frisch SM . C-terminal-binding protein corepresses epithelial and proapoptotic gene expression programs. Proc Natl Acad Sci USA 2003; 100: 4568–4573.
Zhang Q, Yoshimatsu Y, Hildebrand J, Frisch SM, Goodman RH . Homeodomain interacting protein kinase 2 promotes apoptosis by downregulating the transcriptional corepressor CtBP. Cell 2003; 115: 177–186.
Whang SH, Iordanov M, Zhang Q . c-Jun NH2-terminal kinase promotes apoptosis by down-regulating the transcriptional co-repressor CtBP. J Biol Chem 2006; 281: 34810–34815.
Paliwal S, Pande S, Kovi RC, Sharpless NE, Bardeesy N, Grossman SR . Targeting of C-terminal binding protein (CtBP) by ARF results in p53-independent apoptosis. Mol Cell Biol 2006; 26: 2360–2372.
Hildebrand JD, Soriano P . Overlapping and unique roles for C-terminal binding protein 1 (CtBP1) and CtBP2 during mouse development. Mol Cell Biol 2002; 22: 5296–5307.
Miyashita T, Reed JC . Tumor suppressor p53 is a direct transcriptional activator of the human bax gene. Cell 1995; 80: 293–299.
Schmidt T, Körner K, Karsunky H, Korsmeyer S, Müller R, Möröy T . The activity of the murine Bax promoter is regulated by Sp1/3 and E-box binding proteins but not by p53. Cell Death Differ 1999; 6: 873–882.
Bentires-Alj M, Dejardin E, Viatour P, Van Lint C, Froesch B, Reed JC et al. Inhibition of the NF-κB transcription factor increases Bax expression in cancer cell lines. Oncogene 2001; 20: 2805–2813.
Iwase S, Lan F, Bayliss P, de la Torre-Ubieta L, Huarte M, Qi HH et al. The X-linked mental retardation gene SMCX/JARID1C defines a family of histone H3 lysine 4 demethylases. Cell 2007; 128: 1077–1088.
Whetstine JR, Nottke A, Lan F, Huarte M, Smolikov S, Chen Z et al. Reversal of histone lysine trimethylation by the JMJD2A family of histone demethylases. Cell 2006; 125: 467–481.
Cerveny KL, Tamura Y, Zhang Z, Jensen RE, Sesaki H . Regulation of mitochondrial fusion and division. Trends Cell Biol 2007; 17: 563–569.
Karbowski M, Lee YJ, Gaume B, Jeong SY, Frank S, Nechushtan A et al. Spatial and temporal association of Bax with mitochondrial fission sites, Drp1, and Mfn2 during apoptosis. J Cell Biol 2002; 159: 931–938.
Karbowski M, Norris KL, Cleland MM, Jeong SY, Youle RJ . Role of Bax and Bak in mitochondrial morphogenesis. Nature 2006; 443: 658–662.
Frank S, Gaume B, Bergmann-Leitner ES, Leitner WW, Robert EG, Catez F et al. The role of dynamin-related protein 1, a mediator of mitochondrial fission, in apoptosis. Dev Cell 2001; 1: 515–525.
Scorrano L, Ashiya M, Buttle K, Weiler S, Oakes SA, Mannella CA et al. A distinct pathway remodels mitochondria cristae and mobilizes cytochrome c during apoptosis. Dev Cell 2002; 2: 55–67.
Kajimura S, Seale P, Tomaru T, Erdjument-Bromage H, Cooper MP, Ruas JL et al. Regulation of the brown and white fat gene programs through a PRDM16/CtBP transcriptional complex. Genes Dev 2008; 15: 1397–1409.
Arnoult D, Grodet A, Lee YJ, Estaquier J, Blackstone C . Release of OPA1 during apoptosis participates in the rapid and complete release of cytochrome c and subsequent mitochondrial fragmentation. J Biol Chem 2005; 280: 35742–35750.
Cipolat S, Rudka T, Hartmann D, Costa V, Semeels L, Craessaerts K et al. Mitochondrial rhomboid PARL regulates cytochrome c release during apoptosis via OPA1-dependent cristae remodeling. Cell 2006; 126: 163–175.
Zhang D, Lu C, Whiteman M, Chance B, Armstrong JS . The mitochondrial permeability transition regulates cytochrome c release for apoptosis during endoplasmic reticulum stress by remodeling the cristae junction. J Biol Chem 2008; 283: 3476–3486.
Acknowledgements
This study was supported by KOSEF grants from the National Research Laboratory (ROA-2007-000-20002-0) and the Center for Functional Analysis for Human Genome (3344-20060070) to HDY.
Author information
Authors and Affiliations
Corresponding author
Additional information
Edited by RJ Youle
Supplementary Information accompanies the paper on Cell Death and Differentiation website (http://www.nature.com/cdd)
Supplementary information
Rights and permissions
About this article
Cite this article
Kim, J., Youn, H. C-terminal binding protein maintains mitochondrial activities. Cell Death Differ 16, 584–592 (2009). https://doi.org/10.1038/cdd.2008.186
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/cdd.2008.186
Keywords
This article is cited by
-
CtBP Neuroprotective Role in Toxin-Based Parkinson’s Disease Models: From Expression Pattern to Dopaminergic Survival
Molecular Neurobiology (2023)
-
Redox-dependent Brca1 transcriptional regulation by an NADH-sensor CtBP1
Oncogene (2010)
-
Porcine muscle sensory attributes associate with major changes in gene networks involving CAPZB, ANKRD1, and CTBP2
Functional & Integrative Genomics (2009)