Abstract
Hepatitis B virus (HBV) causes chronic hepatitis in hundreds of millions of people worldwide, which can eventually lead to hepatocellular carcinoma (HCC). The molecular mechanisms underlying HBV persistence are not well understood. TRAIL, the TNF-related apoptosis-inducing ligand, has recently been implicated in hepatocyte death during HBV infection. We report here that the HBV core protein (HBc) is a potent inhibitor of TRAIL-induced apoptosis. Overexpressing HBc significantly decreased TRAIL-induced apoptosis of human hepatoma cells, whereas knocking-down HBc expression in hepatoma cells transfected with HBV genome enhanced it. When present in the same cell, HBc blocked the pro-apoptotic effect of the HBV X protein (HBx). The resistance of HBc-expressing cells to TRAIL-induced apoptosis was associated with a significant reduction in death receptor 5 (DR5) expression. Upon transfection, HBc significantly repressed the promoter activity of the human DR5 gene. Importantly, HBc gene transfer inhibited hepatocyte death in a mouse model of HBV-induced hepatitis; and in patients with chronic hepatitis, DR5 expression in the liver was significantly reduced. These results indicate that HBc may prevent hepatocytes from TRAIL-induced apoptosis by blocking DR5 expression, which in turn contributes to the development of chronic hepatitis and HCC. They also call into question the potential side effects of HBc-based vaccines.
Similar content being viewed by others
Main
Hepatitis B virus (HBV) infection remains to be a major health problem worldwide despite the availability of an effective vaccine. More than 350 million people are chronically infected with HBV who are at a high risk of developing hepatitis, cirrhosis and hepatocellular carcinoma.1 To date, the molecular mechanisms of chronic HBV infection have not been elucidated in detail. Recent studies suggest that resistance of HBV-infected hepatocytes to apoptosis may contribute to the development of chronic hepatitis.2 Apoptosis of hepatocytes during HBV infection is mediated by several molecular pathways, which involve at least three members of the TNF superfamily, that is, TNF, Fas ligand (FasL) and TRAIL. Unlike TNF and FasL, TRAIL preferentially induces apoptosis of tumor cells and virus-infected cells but not normal cells.3, 4, 5 Blocking the TRAIL pathway using soluble death receptor 5 (DR5) significantly ameliorates liver inflammation in a mouse model of hepatitis B.6 Interestingly, two HBV proteins, HBV X (HBx)5 and truncated middle hepatitis B surface protein (MHBs(t)),7 were recently found to sensitize hepatocytes to TRAIL-induced apoptosis.
The TRAIL apoptotic pathway is strictly controlled at both the receptor and intracellular signaling levels. Five receptors for TRAIL have been identified, including two death receptors (DR4 and DR5, also known as TRAIL-R1 and TRAIL-R2, respectively) and two decoy receptors (decoy receptor 1(DcR1, TRAIL-R3, and TRID) and decoy receptor 2 (DcR2, TRAIL-R4, and TRUNDD)).4, 8, 9 In mice, only one membrane TRAIL receptor has been identified, which shares 79% sequence homology with human DR5.10 And there are two murine decoy receptors that contain no intracellular death domain.11 In addition to decoy receptors, a variety of intracellular regulators of the apoptotic pathway can modulate the cell's sensitivity to TRAIL.12 We recently found that HBx enhances TRAIL-induced apoptosis through Bax upregulation, whereas MHBs(t) does this through ERK2 activation.5, 7
HBV core protein (HBc), a 21–22 kDa peptide, self-assembles to form the subviral 30–32 nm nucleocapsid particles, which pack the viral polymerase and pregenomic RNA during RNA replication. HBc is highly immunogenic, eliciting strong immune responses during HBV infection. For this reason, HBc has been the top choice for the HBV vaccine development. However, HBc may also mediate hepatocarcinogenesis through several mechanisms.13, 14 The goal of this study is to determine for the first time whether HBc is involved in modulating the TRAIL apoptotic pathway. We report here that HBc blocks TRAIL-induced hepatocyte apoptosis through inhibiting DR5 expression.
Results
HBc transfection decreases the sensitivity of hepatoma cells to TRAIL-induced apoptosis
To investigate the effect of HBc on TRAIL-induced apoptosis, we transfected a widely used human hepatoma cell line BEL7402 with pcDNA3 or pcDNA3 plasmid that carries the HBc gene. Transfected cells were selected in G418-containing medium and tested for HBc expression. As shown in Figure 1a, HBc protein was detected only in BEL7402 cells that were transfected with pcDNA3-HBc. By CCK-8 and TUNEL, a dose-dependent cytotoxicity of TRAIL was detected. BEL7402 cells transfected with HBc exhibited a much lower rate of cell death than control BEL7402 cells and BEL7402 cells transfected with pcDNA3 (Figure 1b and c). Specifically, following incubation with 10 ng/ml TRAIL for 24 h, 22.9±1.3% of control BEL7402 cells underwent apoptosis; this was decreased to 13.1±1.0% for BEL7402 cells transfected with HBc (P<0.01). Similarly, transfection of SMMC7721 cells with HBc but not control pcDNA3 inhibited TRAIL-induced apoptosis (Figure 1d) (P<0.01). These results indicate that HBc is able to decrease the sensitivity of hepatoma cells to TRAIL-induced apoptosis.
HBc transfection decreases the sensitivity of hepatoma cells to TRAIL-induced apoptosis. (a) The hepatoma cell line BEL7402, BEL7402 transfected with pcDNA3 (BEL7402-pcDNA3), or pcDNA3-HBc (BEL7402-HBc) were examined for HBc expression by western blot. (b) BEL7402, BEL7402-pcDNA3 or BEL7402-HBc cells were treated with different concentrations of TRAIL. The cytotoxicity was examined using the CCK-8 kit 24 h later. Three independent experiments using triplicates were carried out. The differences between BEL7402 cells transfected with HBc and other groups are statistically significant (P<0.01). (c) BEL7402, BEL7402-pcDNA3 or BEL7402-HBc cells were treated with PBS or 10 ng/ml TRAIL. Twenty-four hours later, cells were labeled by TUNEL and examined by flow cytometry as described in Materials and Methods. The data shown are representative of five experiments. The differences between BEL7402 cells transfected with HBc and other groups are statistically significant (P<0.01). (d) The hepatoma cell lines SMMC7721 transfected with pcDNA3 or pcDNA3-HBc were treated with PBS or 100 ng/ml TRAIL. Twenty-four hours later, cells were labeled by TUNEL and examined by flow cytometry. The data shown are representative of three experiments. The differences between SMMC7721 cells transfected with HBc and other groups are statistically significant (P<0.01)
To test this theory using a loss-of-function approach, antisense oligonucleotide specific to HBc gene, PS-asODNs/HBc, was used to block HBc expression in BEL7402 cells transfected with pcDNA3-HBV1.1 (Figure 2a and b). We found that HBc knockdown significantly upregulated the sensitivity of BEL7402 cells transfected with pcDNA3-HBV1.1 to TRAIL-induced apoptosis. Thus, 54.7±4.8% of BEL7402-HBV1.1 cells pretreated with PS-rODNs underwent TRAIL-induced apoptosis; this was increased to 68.4±5.6% for BEL7402-HBV1.1 cells pretreated with PS-asODNs/HBc (P<0.01) (Figure 2c).
HBc knockdown increases the sensitivity of HBV1.1-transfected BEL7402 cells to TRAIL-induced apoptosis. (a) and (b) BEL7402-HBV1.1 cells were treated with 10 μmol/l antisense oligonucleotide against HBc (PS-asODNs/HBc), or random control oligonucleotide (PS-rODNs). Forty-eight hours later, HBc expression was examined by western blot. Graphics show mean values and S.D. corresponding to three independent experiments. The differences between BEL7402-HBV1.1 cells treated with PS-asODNs/HBc and other groups are statistically significant (P<0.01). (c) BEL7402-HBV1.1, pretreated with PS-asODNs/HBc or PS-rODNs for 24 h, was exposed to 10 ng/ml TRAIL for another 24 h. Then TUNEL assay was used to examine the apoptosis rate. The data shown are representative of three experiments. The differences between BEL7402-HBV1.1 cells treated with PS-asODNs/HBc and PS-rODNs are statistically significant (P<0.01)
Taken together, these results establish for the first time that, unlike other HBV proteins that promote apoptosis, HBc acts as an inhibitor of TRAIL-induced apoptosis of hepatocytes.
HBc blocks the proapoptotic effect of HBx
We and others5, 15 reported that HBx significantly enhances TRAIL-induced apoptosis of hepatocytes. To determine the consequence of expressing two HBV proteins with opposing activities, we transiently transfected BEL7402-HBx cells, which stably expressed HBx,5 with pcDNA3-HBc or control plasmid pcDNA3. By western blot, we found that HBx was expressed at similar levels in BEL7402-HBx cells with or without HBc transfection (Figure 3a). However, transfection of BEL7402-HBx cells with HBc, but not control pcDNA3, significantly inhibited TRAIL-induced apoptosis (Figure 3b). Specifically, 40.1±4.4% of control BEL7402-HBx cells underwent apoptosis following TRAIL treatment; this was reduced to 26.9±4.3% for BEL7402-HBx cells transfected with HBc, which is similar to that of control BEL7402 cells not transfected with HBx (P<0.01). These results indicate that HBc is able to block the effect of HBx on TRAIL-induced apoptosis.
HBc blocks the pro-apoptotic effect of HBx. BEL7402-HBx were transfected with HBc or pcDNA3. Twenty-four hours later, (a) western blot assay was used to detect HBx expression. (b) BEL7402-pcDNA3, BEL7402-HBx, BEL7402-HBx transfected with HBc, or pcDNA3 were treated with 10 ng/ml TRAIL for another 24 h. TUNEL assay was used to examine the apoptotic rate. The data shown are representative of three experiments. The differences between BEL7402-HBx cells transfected with pcDNA3-HBc and pcDNA3 are statistically significant (P<0.01)
HBc blocks hepatocyte apoptosis during HBV-induced hepatitis
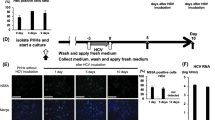
Our previous studies demonstrated that TRAIL plays an important role in hepatic cell death during HBV-induced hepatitis.6 To determine whether HBc is involved in modulating TRAIL-induced hepatocyte apoptosis, we performed in vivo HBc gene transfer in an acute model of hepatitis B. As shown in Figure 4a, hydrodynamic administration of pcDNA3-HBV1.1 (to induce hepatitis) together with HBc plasmids led to significant expressions of HBc in hepatocytes. Histochemical analysis of the liver sections revealed less inflammation in the HBc group as compared with control hepatitis groups (Figure 4b). HBc expression also reduced blood levels of ALT (Figure 4c). This was accompanied by a significant reduction of apoptotic cell numbers in the liver (Figure 4d). Specifically, 12.4±2.4% of hepatocytes in control hepatitis B groups underwent apoptosis. This was reduced to 7.0±1.3% for pcDNA3-HBV1.1 together with HBc groups (Figure 4e) (P<0.01). These data indicate that HBc may inhibit the apoptosis of hepatocytes during HBV-induced hepatitis and control the extent of liver injury.
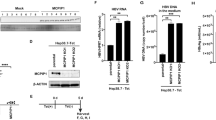
HBc impairs the apoptosis of hepatocytes in HBV-induced hepatitis. Four groups of BALB/c mice, five mice per group, were hydrodynamically injected with 120 μg of control plasmid pcDNA3 (pcDNA3 group), or 60 μg of pcDNA3-HBc plus 60 μg of pcDNA3 (HBc group), or 60 μg of pcDNA3-HBV1.1 plus 60 μg of pcDNA3 (HBV1.1 group, acute hepatitis B model), or 60 μg of pcDNA3-HBV1.1 plus 60 μg of pcDNA3-HBc (HBV1.1+HBc group). Forty-eight hours later, liver tissues were sectioned and examined for HBc expression by immunohistochemistry (a), or for pathology by hematoxylin and eosin staining (b) and serum ALT levels (c). HBc overexpression significantly ameliorated the liver injury in this model of hepatitis B (P<0.01). The apoptosis rate of hepatocytes was examined by TUNEL assay. (d) Liver sections were stained by TUNEL. Original magnifications, × 200. (e) Single cell suspensions of freshly isolated hepatocytes were stained by TUNEL and examined by flow cytometry. The data shown are representative of three experiments. The differences between HBc overexpression group (HBV1.1+HBc) and acute hepatitis B model group are statistically significant (P<0.01)
HBc downregulates DR5 expression both in vitro and in vivo
To explore the mechanisms through which HBc ameliorates TRAIL-induced hepatic cell death, we examined a panel of apoptosis mediators/regulators in BEL7402 cells that do or do not express HBc. Unexpectedly, we found that HBc transfection significantly decreased the expression of DR5 at both mRNA (Figure 5a,b and d) and protein levels (Figure 5c, e and f and Supplementary Figure 1), with no detectable effect on the expression of DR4, DcR1 or DcR2. Similar results were obtained when SMMC7721 cells were used for HBc transfection (Figure 5g and our unpublished data) (P<0.01). In addition, we also tested the DR5 expression in BEL7402-HBx cells with or without HBc transfection. As shown in Figure 5h and Supplementary Figure 2, cell surface levels of DR5 expression were significantly decreased in BEL7402-HBx cells after HBc transfection (P<0.01).
HBc transfection decreases DR5 expression in hepatoma cells. (a) Human hepatoma cell BEL7402, BEL7402-pcDNA3 or BEL7402-HBc were examined for TRAIL receptor expression by RT-PCR. The data shown are representative of three experiments. (b)The receptor /β-actin ratios were determined by densitometry. (c) TRAIL receptor levels were examined by flow cytometry. The differences in DR5 protein expression between BEL7402-pcDNA3 and BEL7402-HBc cells are statistically significant (P<0.01). Graphics shown are representative of five experiments. DR5 expression in BEL7402, BEL7402-pcDNA3 or BEL7402-HBc was detected by real-time PCR (d) and western blot (e and f). (g) SMMC7721, SMMC7721-pcDNA3 or SMMC7721-HBc were examined for DR5 protein expression by flow cytometry. Graphics show mean values and S.D. corresponding to three independent experiments. The differences in DR5 expression between SMMC7721 cells that do or do not express HBc are statistically significant (P<0.01). (h) BEL7402-HBx, transfected with HBc or pcDNA3, were examined for DR5 protein expression by flow cytometry. The differences in DR5 expression between BEL7402-HBx cells that do or do not express HBc are statistically significant (P<0.01). (i) BEL7402-pcDNA3 and BEL7402-HBc cells were firstly treated with 3 μg DR5 blocking antibody HS201 or mouse IgG as an isotype control and then exposed to 10 ng/ml TRAIL. The apoptosis rate was determined by TUNEL assay. The data shown are representative of three experiments. DR5 blocking antibody significantly inhibited TRAIL-induced apoptosis in both BEL7402-pcDNA3 and BEL7402-HBc cells (P<0.01)
To determine whether DR5 downregulation at the cell surface is sufficient to inhibit TRAIL-induced apoptosis, we used an antagonistic antibody, HS201, to block DR5-mediated effects. As shown in Figure 5i, HS201 inhibited TRAIL-induced apoptosis both in BEL7402-pcDNA3 and BEL7402-HBc cells. But after DR5 blockade, there was no significant difference in TRAIL-induced apoptosis between BEL7402-pcDNA3 cells and BEL7402-HBc cells. This data demonstrates that DR5 deregulation is likely responsible for the effect of HBc on TRAIL-induced apoptosis.
On the other hand, HBc transfection did not significantly alter the levels of FLIP, Bax (Supplementary Figure 3), Bid, and procaspases 3, 8 and 9 (Figure 6a). Upon treatment with TRAIL, procaspases 3, 8,9 and Bid are degraded in BEL7402 cells leading to a reduction in the levels of these proteins as compared with controls (Figure 6a and b). Degradation of procaspases 3, 8, 9 and Bid was significantly decreased in cells transfected with HBc (Figure 6a and b). This is consistent with the theory that there is a crosstalk between the mitochondrial and non-mitochondrial pathways of apoptosis. Thus, downregulated DR5 expression may contribute to the decrease in the activation of procaspases 3, 8, 9 and Bid.16
HBc transfection blocks TRAIL-induced degradation of procaspases 3, 8, 9 and Bid. (a) BEL7402, BEL7402 transfected with pcDNA3 or HBc were treated with (+) or without (−) 10 ng/ml of TRAIL for 24 h. The levels of procaspase 8, procaspase 9, procaspase 3 and Bid were examined by western blot. β-actin was used as a loading control. The procaspase 8, procaspase 9, procaspase 3 and Bid/β-actin ratios were determined by densitometry. (b) The relative level of activation is calculated according to the formula: (the value of the mock group/β-actin of the mock group – the value of TRAIL-treated group/β-actin of TRAIL-treated group) × 100%. Mean values obtained in three independent experiments are shown. The differences in relative level of procaspases and Bid activation between BEL7402 cells that do or do not express HBc are statistically significant (P<0.01)
To determine whether HBc also regulates DR5 expression in normal mice or mice with hepatitis B virus transfection, we examined the DR5 protein levels in hepatocytes after hydrodynamic injection with pcDNA3-1.1HBV and/or pcDNA3-HBc for 48 h. As shown in Figure 7a, 14.7±1.79% and 13.5±0.9% of hepatocytes were DR5-positive in pcDNA3- and HBV1.1-treated groups, respectively; these were reduced to 8.8±1.6% and 9.1±1.1% when HBc was co-transfected(P<0.01). Similar changes in DR5 expression were observed by immunohistochemistry and western blot (Figure 7b and c). In addition, the degradation of procaspases 3, 8, 9 and Bid was significantly reduced in HBc co-transfected group as compared with the control group (Figure 7d) (P<0.01). These results are consistent with the reduced apoptotic rate of hepatocytes in HBc-expressing groups (Figure 4e). Thus, the inhibitory effects of HBc on TRAIL-induced apoptosis may be due to decreased DR5 expression.
HBc reduces DR5 expression and the degradation of procaspases 3, 8, 9 in vivo. Four groups of BALB/c mice, five mice per group, received hydrodynamic injection as described. (a) Forty-eight hours later, single cell suspension of fresh isolated hepatocytes was examined for DR5 expression by flow cytometry. The data shown are representative of three experiments. The differences between pcDNA3 group and HBc group, HBV1.1 group and HBV1.1 plus HBc group are statistically significant (P<0.01). (b) Liver sections were examined for DR5 expression by immunohistochemical staining. (c and d) Liver tissues were examined for DR5 by western blot. (e and f) Western blot was used to detect the procaspase 8, procaspase 9, procaspase 3 and Bid. β-actin was used as a loading control. Mean values obtained in three independent experiments are shown. The differences between pcDNA3 group and HBc group, HBV1.1 group and HBV1.1 plus HBc group are statistically significant (P<0.01)
HBc downregulates the DR5 promoter activity
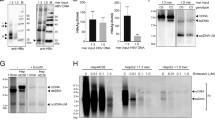
Our finding that HBc decreased the DR5 mRNA expression indicates that HBc may regulate DR5 gene expression at the transcriptional level. To test this theory, we used a dual luciferase reporter system to analyze the DR5 promoter activity in BEL7402 cell lines that do or do not express HBc. As shown in Figure 8a, BEL7402-pcDNA3 cells exhibited a strong DR5 promoter activity; this was significantly decreased in BEL7402-HBc cells (P<0.01). We then performed this experiment using BEL7402 cells transiently transfected with increasing amounts of pcDNA3 or pcDNA3-HBc. The result showed that higher amount of HBc plasmids had a more significant inhibition of the DR5 promoter (Supplementary Figure 4). Importantly, the transfection efficiencies in different groups were comparable (Supplementary Figure 5). Consistent with a report by Kwon et al.13 HBc transfection downregulated the activity of the p53 promoter (Figure 8a), although having no effect on the activity of the hTERT promoter (Figure 8a). As expected, HBx did not alter the expression of DR5 or the activity of its promoter (Figure 8b). These results demonstrate that HBc selectively regulates DR5 expression at the promoter level.
HBc decreases the DR5 promoter activity. (a) BEL7402-pcDNA3 BEL7402-HBc were transfected with 0.2 μg pDR5PF, 0.4 μg pDR5PF, p53 promoter pGL3-p53, or hTERT-306. (b) BEL7402 cells were co-transfected with pDR5PF and HBx. Control plasmid pGL3-basic was used to ensure that all groups received a total of 1 μg plasmids. Forty-eight hours later, the cells were lysed and luciferase activity was measured. Data represent the mean±S.D. of three independent experiments. The differences in relative activity of DR5 promoter and p53 promoter between BEL7402 cells that do or do not express HBc are statistically significant (P<0.01)
Reduced hepatic DR5 levels in patients with chronic hepatitis
Results described above support the view that HBc prevents TRAIL-induced hepatic cell death in HBV-induced hepatitis. As reduced apoptosis of HBV-infected hepatocytes may contribute to the development of chronic hepatitis,2 we asked whether HBc-mediated inhibition of TRAIL-induced apoptosis might play a role in human chronic hepatitis B. As a first step to address this question, we examined the relationship between DR5 and HBc expression in chronic hepatitis B patients. We found that 81% of chronic hepatitis B patients were HBc-positive in serum. As reported, DR4 and DR5 were both expressed in normal liver tissues.17 DR5 but not DR4 expression was significantly reduced as judged by both intensity and number of cells expressing it, in chronic hepatitis B patients (Figure 9a–c). Moreover, soluble DR5 in patients' sera was also decreased: 20.37±6.88 pg/ml as compared with 31.42±6.79 pg/ml in normal serum (P<0.001). These clinical data indicate that low expression of DR5 in chronic hepatitis B patients may play a role in the development of the disease, which may be related with the HBc expression in these patients.
Reduced DR5 expression in patients with chronic hepatitis B. Human liver sections from normal control and patients with chronic hepatitis B were examined for DR4 (a) and DR5 (b) expression by immunohistochemistry (Original magnifications, × 200). At least 50 independent fields per liver (at × 200 magnifications) were counted, with 100±5 hepatocytes per field. (c) The differences in DR5 expression between the normal control and chronic hepatitis B patients are statistically significant (P<0.01)
Discussion
TRAIL is an attractive drug for cancer therapy owing to its selective cytotoxicity to tumor cells.18, 19 However, recent studies of TRAIL function under physiological and pathological conditions, especially inflammatory conditions, have raised concerns about the TRAIL-based therapy. Although normal hepatocytes appear to be resistant to TRAIL,20 our previous studies revealed that HBV infection and HBx protein expression sensitized hepatocytes to TRAIL-induced apoptosis.6 Here we showed that HBc, another viral protein encoded by HBV, played an opposite role in TRAIL-induced hepatocyte apoptosis. The finding is important because it shows for the first time that different HBV proteins can have distinct effects on TRAIL-induced apoptosis.
The expression levels of death receptors may play a critical role in determining the strength and/or duration of death receptor-mediated apoptotic signaling in response to death ligands. It is well-known that TRAIL triggers apoptotic cascades through specifically binding to the death receptors, DR4 and DR5, which contain cytoplasmic death domains and are in turn responsible for recruiting adaptor molecules involved in caspase activation.21 Under physiologic conditions (37°C), DR5 is known to have a higher affinity for TRAIL than DR4.22 Furthermore, by using phage display of death receptor selective TRAIL variants, it was demonstrated that DR5 may play a more prominent role than DR4 in mediating apoptotic signals derived from TRAIL in cells expressing both death receptors.23 Recently, various agents including IFN-γ24, 25 have been reported to downregulate DR5 expression. Also, decreased DR5 expression was found in human tumors such as hepatoma26 and ovarian cancer.27 This indicates that downregulated DR5 may play a role in the pathogenesis of malignant tumors. In this study, we discovered that HBc significantly decreases DR5 mRNA and protein expression both in hepatoma cells and in murine hepatocytes. This finding was also confirmed in our cDNA microarray analysis (data not shown). Importantly, in patients with chronic hepatitis B, decreased DR5 expression was associated with the persistent presence of HBc in serum and in liver tissues. Moreover, we found that DR5-blocking antibody neutralizes the effect of HBc on TRAIL-induced apoptosis, further demonstrating that DR5 deregulation is most crucial for the reduced TRAIL-induced apoptosis. In addition, we also examined whether the Akt pathway was involved in the effect of HBc. As shown in the Supplementary Figure 6, the inhibitor of Akt did not alter the effect of HBc transfection on TRAIL-induced apoptosis.
Several studies have suggested the possibility of HBc as a gene regulatory protein based on the presence of nucleic acid-binding motifs, nuclear localization signals, phosphorylation sites, and its localization in the nucleus.28, 29, 30, 31 It was reported earlier that HBc inhibited the expression of the β-interferon gene and HBx.14, 32 Recently, HBc has been shown to repress the expression of p53.13 Our microarray analysis also indicated that HBc represses P53 but not NF-kB expression in hepatocytes (data not shown). Transcription factors such as NF-κB33 and p53,34, 35 can regulate the DR5 promoter activity. Here, we show for the first time that HBc may decrease DR5 expression by attenuating the DR5 promoter activity. As reported earlier, the DR5 gene promoter has no typical TATA-box, but has two Sp1 sites responsible for the basal transcription activity of the DR5 gene.36 Whether the HBc-p53 pathway plays a significant role in regulating DR5 expression needs to be further studied.
We and others reported that HBV transgenic mice spontaneously develop liver cancer around 20 months of age.37 Our recent study indicates that HBc is expressed at a higher level at this age than at a younger age (2, 6 and 12 months) (data not shown). It is worth noting that HBc is being tested as a potential carrier for poorly immunogenic peptides and carbohydrate B cell epitopes because of its strong immunogenicity.38 This study may call this into question. This is another reason why the biological functions of HBc need to be further studied.
In summary, we discovered that unlike other HBV proteins that promote apoptosis, HBc ameliorates TRAIL-induced apoptosis of hepatocytes. This indicates that different HBV proteins have different, even opposite, effects on TRAIL-induced hepatocyte apoptosis. If the pro-apoptotic proteins, such as HBx, are dominant, HBV-infected hepatocytes may die as a consequence and fulminant hepatitis may develop. By contrast, if the anti-apoptotic viral proteins such as HBc prevail, the infected hepatocytes may not undergo apoptosis and chronic HBV infection may ensue. These findings may help clarify the roles of imbalanced apoptosis in the pathogenesis of the hepatitis B infection.
Materials and Methods
Cells and cell culture
The human hepatoma cell lines BEL7402 and SMMC7721 were purchased from the Shanghai Institute of Cell Biology at the Chinese Academy of Sciences. All cells were maintained in RPMI-1640 containing 10% fetal bovine serum and were cultured in 5% CO2 atmosphere at 37°C.
Mice
Six- to 8-week-old male BALB/c mice were obtained from the Laboratory Animal Center of Shandong University. All mice were housed in the Institute of Immunology Shandong University animal facilities under pathogen-free conditions. All procedures were pre-approved by the Institutional Animal Care and Use Committee.
Plasmids
pcDNA3-HBc was generated by inserting a copy of the HBc gene (amplified from pUC19/3HBV which was provided by Professor Akira Nishizono, Oita Medical University, Japan) into the HindIII and EcoRV site of the pcDNA3 (which contains a neomycin resistance gene under the control of the CMV promoter). pcDNA3-HBV1.1 was constructed by inserting 110% of HBV genome from the p3.6 II/HBV (provided by Professor Y Wang, Institute of Biochemistry and Cell Biology, Chinese Academy) into EcoRV site of pcDNA3. pcDNA3-HBx was generated by inserting a copy of the HBx gene into the HindIII and XbaI site of the pcDNA3. Qiagen plasmid mini/maxi kits were used to extract and purify the recombinant plasmids.
Liposome-mediated stable transfection
Human hepatoma cell line BEL7402 was transfected with pcDNA3-HBc, pcDNA3-HBx, pcDNA3-HBV1.1 or pcDNA3 using lipofectamineTM2000 (GIBCO Inc.) per manufacturer's instructions. Briefly, BEL7402 cells were seeded into 24-well plates with 1.5 × 105 cells per well and cultured for 20 h. The medium was removed and cells were exposed to the transfection complexes (1 μg of DNA and 2 μl of liposome mixed in 100 μl of serum-free Opti-MEM) for 6 h at 37°C. Cells were then cultured in a complete medium for 24 h. The transfected cells were selected in a medium containing 380 μg/ml of G418 (GIBCO Inc.) for ∼4 weeks. Then BEL7402-pcDNA3 cells, BEL7402-HBc BEL7402-HBx and BEL7402-HBV1.1 cells were obtained. Human hepatoma cell line SMMC7721 was also transfected with pcDNA3-HBc or pcDNA3 and was selected in a medium containing 1.5 mg/ml G418, respectively.
Cytotoxicity of TRAIL
BEL7402 cells, BEL7402-pcDNA3 cells and BEL7402-HBc cells were seeded into 96-well microtiter plates with 2 × 104 cells per well and cultured for 20 h. Different concentrations of TRAIL (0, 2, 5, 10, 20, 50 ng/ml) (PeproTech, USA) were added to the medium and cells were incubated for another 24 h. Cell Counting Kit 8 (CCK-8) (Donjindo, Japan) was used to detect the cytotoxicity of TRAIL. The absorbance (A) at the wavelength of 450 nm was measured using a microplate reader (Bio-RAD Model 680, USA). Cytotoxicity Rate (%)=(1-mean A value of the experimental group/mean A value of the control group) × 100%.
TUNEL assay
Cells were seeded into 24-well plates with 1.5 × 105 cells per well and treated with recombinant soluble TRAIL (PeproTech) for 24 h. Apoptotic cells were labeled using an in situ apoptosis detection kit (Immunotech) by TdT (terminal deoxynucleotidyl transferase) -mediated dUTP nick end-labeling (TUNEL). The degree of apoptosis was determined by flow cytometry (Beckman Coulter FC500, Germany). For liver tissues, the paraffin sections were stained using TUNEL kit and observed on fluorescence microscopy. Simultaneously, liver tissues grinded into single cell suspension were labeled by TUNEL and apoptotic rates were detected by flow cytometry.
Semi-quantitative and real-time PCR analysis
Total RNA was extracted from the cultured cells with Trizol reagent (Bio Basic Inc., Canada) in accordance with the manufacturer's protocol. Reverse transcription reaction was performed and the products of cDNA were used as a template for PCR. Semi-quantitative PCR for DR4, DR5, DcR1, DcR2 and β-actin were performed as described earlier.7 PCR products were analyzed on a 2% agarose gel by electrophoresis and ethidium bromide staining. Gels were scanned on a digital imaging system and the band intensity values, including those of β-actin, were transformed logarithmically. The TRAIL receptor mRNA levels were calculated as ratios of TRAIL receptor signals over that of the β-actin. In this way, each receptor mRNA value was normalized against the β-actin mRNA content. Results presented are representative of three individual experiments.
Real-time PCR was performed on a LightCycler (Roche Diagnostics, Mannheim, Germany) using the LightCycler FastStart DNA Master SYBR Green I kit. Primers for DR5 and Actin were as follows: DR5, Forward: 5-GGGCCACAGGGACACCTT-3; Reverse: 5-GCATCTCGCCCGGTTTT-3; β-actin Forward: AGTTGCGTTACACCCTTTC; Reverse: CCTTCACCGT TCCAGTTT. PCR conditions used were as follows: denaturation at 95 °C for 15 seconds, annealing at 60 °C for 15 seconds, and extension at 72 °C for 15 s. Results are presented as ratios of target genes/β-actin.
Detection of TRAIL receptor expression by flow cytometry
BEL7402 cells, BEL7402-pcDNA3 cells and BEL7402-HBc cells or freshly isolated mouse hepatocytes were collected and washed two times in PBS. They were then incubated with mouse anti-human DR4, DR5, DcR1, and DcR2 mAbs (Fourth Military Medical University, Xi'an, China) at 4°C for 30 min. After washing, cells were treated with FITC-conjugated goat anti-mouse IgG (Immunotech) for 30 min. The levels of the TRAIL receptors were determined as an average fluorescence intensity by flow cytometry.
Western blot
Cells were washed three times in PBS and incubated in the lysis buffer (50 mM Tris Cl (pH 8.0), 150 mM NaCl, 0.1% SDS, 1% Nonidet P-40, 0.02% sodium azide, 100 μg/ml PMSF, 1 μg/ml peptin and 1 μg/ml aprotinin) for 30 min on ice. Equal amount of protein from each lysate was subjected to 12% SDS-PAGE and transferred onto a nitrocellulose membrane. After blocking for 3 h in 5% nonfat dried milk containing 0.1% Tween 20, the membrane was incubated at 4°C overnight in the presence of Abs to human/mouse DR5, caspase-3, caspase-8, caspase-9, Bid, Flip, Bax,HA and β-actin (Santa Cruz Biotechnology, USA). The membrane was washed and further incubated for 1 h at room temperature with HRP-conjugated secondary Abs (Zhongshan Co., Beijing, China). Following washing, immunoreactive bands were detected using Western Lightning Chemilluminescence Reagent (Amersham Biosciences).
Blockade of HBc expression by antisense oligonucleotides
A 16-mer PS-ODNs (5′-CTCCAAATTCTTTATA-3′), corresponding to the nucleotides ∼1918–1933 and complementary to HBc poly-A region of HBV genome (adr subtype), called PS-asODNs/HBc, was synthesized. A random 16-mer PS-ODNs (5′-AGTCAGTCAGTCAGTC-3′), unrelated to the HBV sequence, called PS-rODNs, was designed and synthesized as a control. The powder was dissolved in RPMI-1640 complete culture medium. BEL7402-HBV1.1 cells, seeded into 24-well microtiter plates with 1 × 105 cells per well, were at first treated with 10 μmol/l PS-asODNs/HBc or PS-rODNs for 24 h. Then TRAIL (10 ng/ml) was added to the medium and cells were incubated for another 24 h. The apoptosis rate was determined by TUNEL assay.
Blockade of DR5-mediated apoptosis by antagonistic antibody HS201
BEL7402-pcDNA3 and BEL7402-HBc cells, were incubated in 24-well microtiter plates at 1.5 × 105 cells per well, with either 3 μg/ml DR5 blocking antibody (HS201, Apotech) or mouse IgG1 isotype control for 30 min on ice. TRAIL (10 ng/ml) was then added to the culture, and cells were incubated for another 24 h. The apoptosis rate was determined by the TUNEL assay.
Hydrodynamic injection
To express HBV antigens in vivo and to induce acute hepatitis, BALB/c mice received hydrodynamics-based tail vein injections of plasmids as described.39 Briefly, groups of mice were injected i.v. with (1) 120 μg of pcDNA3, (2) 60 μg of pcDNA3-HBc and 60 μg of pcDNA3, (3) 60 μg of pcDNA3-HBV1.1 and 60 μg of pcDNA3, or (4) 60 μg of pcDNA3-HBV1.1 and 60 μg of pcDNA3-HBc, within 5–8 s in a volume of saline equivalent to 8% of the mouse body weight. Mice injected with pcDNA3 or saline served as control. Forty-eight hours later, the levels of alanine aminotransferase in the serum were determined on a SpectraMax Plus spectrophotometer using the 3V Infinity Reagents (3V Biotech) per manufacturer's instructions. Liver tissues were sectioned, stained with HE and examined by light microscopy.
Immunohistochemical staining
Paraffin sections of mouse and human liver tissues were dewaxed with xylene, treated with 3% H2O2 for 10 min and washed three times with PBS. Thereafter, the sections were incubated with anti-HBc (Zsbio) or anti-DR5 antibody (CHEMICON) at 4°C overnight. After washing, a biotin-labeled secondary antibody (Maixin co., Fuzhou) was added to the sections followed by streptavidin-conjugated peroxidase. After incubation with the DAB substrate, the sections were counterstained with hematoxylin. The percentage of positive hepatocytes was determined by microscopy.
Dual luciferase reporter assay
Luciferase assays were performed with the Promega Luciferase Assay System (Promega, Madison, WI, USA), according to the manufacturer's specifications. pGL3-basic luciferase reporter vector and pDR5PF plasmid, containing DR5 promoter sequence (−2500/+3), were kindly provided by Dr. Sakai (Kyoto Prefectural University). pGL3-p53 plasmid was kindly provided by professor Kakar (University of Louisville). hTERT promoter hTERT-306, containing hTERT promoter sequence (−306/+135), was constructed by us. BEL7402-pcDNA3 and BEL7402-HBc cells in a 24-well plate, were transiently transfected with 0.2 μg or 0.4 μg pDR5PF using Lipofectamine 2000 (Invitrogen, USA). p53 promoter pGL3-p53, was used as a positive control, whereas hTERT-306 was used as a negative control. In addition, we also transfected pDR5PF into BEL7402 cells stably expressing HBx. Control plasmid pGL3-basic was used to ensure that the total amount of the plasmids used in each well was 1 μg. Twenty-five ng of pRL-TK was also included in each group to normalize transfection efficiency. The cells were harvested 48 h after the transfection and used for the luciferase assay (Promega, USA). The firefly luciferase activity was normalized against the Renilla reniformis luciferase activity of the co-transfected pRL-TK to control for variations in transfection efficiency.
Detection of DR4, DR5 and HBc expression in patients with chronic hepatitis B
All chronic hepatitis B patients were diagnosed according to the criteria of viral hepatitis issued at the Fifth National Conference on Infectious Diseases and Parasites. Patients co-infected with human immunodeficiency virus or hepatitis C virus were excluded. All procedures were approved by the Ethics Committee of the Shandong University, and appropriate consent was obtained from patients.
Sera of 110 patients with chronic hepatitis B and 30 normal healthy controls were collected at the Second Affiliated Hospital of Shandong University or the Infectious Disease Hospital of Jinan, Shandong. HBc expression and soluble DR5 levels in sera were detected using ELISA kits (Mdbio, China and Biosource, respectively).
Liver samples from 32 chronic hepatitis B patients were obtained by liver centesis. Eight normal liver tissue samples were obtained from patients with liver hemangioma or hepatorrhexis. DR4 and DR5 expression was determined by immunohistochemistry as described above.
Statistical analysis
The apoptosis rates and mRNA or protein expression ratios were analyzed by χ2 test. Enzyme and protein levels were analyzed by ANOVA.
Abbreviations
- DR5:
-
death receptor 5
- HBc:
-
HBV core protein
- HBV:
-
hepatitis B Virus
- HBx:
-
HBV X protein
- MHBs(t):
-
truncated middle hepatitis B surface protein
- TRAIL:
-
TNF-related apoptosis-inducing ligand
References
Beasley RP, Hwang LY, Lin CC, Chien CS . A prospective study of 22 707 men in Taiwan Hepatocellular carcinoma and hepatitis B virus. Lancet 1981; 2: 1129–1133.
Liu DX . A new hypothesis of pathogenetic mechanism of viral hepatitis B and C. Med Hypotheses 2001; 56: 405–408.
Wiley SR, Schooley K, Smolak PJ, Din WS, Huang CP, Nicholl JK S et al. Identification and characterization of a new member of the TNF family that induces apoptosis. Immunity 1995; 3: 673–682.
Pan G, Ni J, Wei YF, Yu G, Gentz R, Dixit VM . An antagonist decoy receptor and a death domain-containing receptor for TRAIL. Science 1997; 277: 815–818.
Liang X, Liu Y, Zhang Q, Gao L, Han L, Ma C et al. Hepatitis B virus sensitizes hepatocytes to TRAIL-induced apoptosis through Bax. J Immunol 2007; 178: 503–510.
Liu YG, Liu SX, Liang XH, Zhang Q, Gao LF, Han LH et al. Blockade of TRAIL pathway ameliorates HBV-induced hepatocyte apoptosis in an acute hepatitis model. Biochem Biophys Res Commun 2007; 352: 329–334.
Liang X, Du J, Liu Y, Cui M, Ma C, Han L et al. The hepatitis B virus protein MHBs(t) sensitizes hepatoma cells to TRAIL-induced apoptosis through ERK2. Apoptosis 2007; 12: 1827–1836.
Pan G, Ni J, Yu G, Wei YF, Dixit VM . TRUNDD, a new member of the TRAIL receptor family that antagonizes TRAIL signalling. FEBS Lett 1998; 424: 41–45.
Schneider P, Bodmer JL, Thome M, Hofmann K, Holler N, Tschopp J . Characterization of two receptors for TRAIL. FEBS Lett 1997; 416: 329–334.
Wu GS, Burns TF, Zhan Y, Alnemri ES, El-Deiry WS . Molecular cloning and functional analysis of the mouse homologue of the KILLER/DR5 tumor necrosis factor-related apoptosis-inducing ligand (TRAIL) death receptor. Cancer Res 1999; 59: 2770–2775.
Schneider P, Olson D, Tardivel A, Browning B, Lugovskoy A, Gong D et al. Identification of a new murine TNF receptor locus that contains two novel murine receptors for tumor necrosis factor-related apoptosis-inducing ligand (TRAIL). J Biol Chem 2002; 278: 5444–5454.
Sheridan JP, Marsters SA, Pitti RM, Gurney A, Skubatch M, Baldwin D et al. Control of TRAIL-induced apoptosis by a family of signaling and decoy receptors. Science 1997; 277: 818–821.
Kwon JA, Rho HM . Transcriptional repression of the human p53 gene by hepatitis B viral core protein (HBc) in human liver cells. Biol Chem 2003; 384: 203–212.
Kim JH, Kang S, Kim J, Ahn BY . Hepatitis B virus core protein stimulates the proteasome-mediated degradation of viral x protein. J Virol 2003; 77: 7166–7173.
Janssen HL, Higuchi H, Abdulkarim A, Gores GJ . Hepatitis B virus enhances tumor necrosis factor-related apoptosis-inducing ligand (TRAIL) cytotoxicity by increasing TRAIL-R1/death receptor 4 expression. J Hepatol 2003; 39: 414–420.
Deng Y, Lin Y, Wu X . TRAIL-induced apoptosis requires Bax-dependent mitochondrial release of Smac/DIABLO. Genes Dev 2002; 16: 33–45.
Kim TH, Seol DW, Esplen JE, Dorko K, Billiar TR, Strom SC . Apoptosis induced in normal human hepatocytes by tumor necrosis factor-related apoptosis-inducing ligand. Nat Med 2000; 6: 564–567.
Walczak H, Miller RE, Ariail K, Gliniak B, Griffith TS, Kubin M et al. Tumoricidal activity of tumor necrosis factor-related apoptosis-inducing ligand in vivo. Nat Med 1999; 5: 157–163.
Ashkenazi A, Pai RC, Fong S, Leung S, Lawrence DA, Marsters SA et al. Safety and antitumor activity of recombinant soluble Apo2 ligand. J Clin Invest 1999; 104: 155–162.
Hao C, Song JH, his B, Lewis J, Song DK, Petruk KC et al. TRAIL inhibits tumor growth but is nontoxic to human hepatocytes in chimeric mice. Cancer Res 2004; 64: 8502–8506.
Kischkel FC, Lawrence DA, Chuntharapai A, Schow P, Kim KJ, Ashkenazi A . Apo2 L/TRAIL-dependent recruitment of endogenous FADD and caspase-8 to death receptors 4 and 5. Immunity 2000; 12: 611–620.
Truneh A, Sharma S, Silverman C, Khandekar S, Reddy MP, Deen KC et al. Temperature-sensitive differential affinity of TRAIL for its receptors DR5 is the highest affinity receptor. J Biol Chem 2000; 275: 23319–23325.
Kelley RF, Totpal K, Lindstrom SH, Mathieu M, Billeci K, Deforge L et al. Receptor-selective mutants of apoptosis-inducing ligand 2/tumor necrosis factor-related apoptosis-inducing ligand reveal a greater contribution of death receptor (DR) 5 than DR4 to apoptosis signaling. J Biol Chem 2005; 280: 2205–2212.
Bretz JD, Mezosi E, Giordano TJ, Gauger PG, Thompson NW, Baker JJ . Inflammatory cytokine regulation of TRAIL-mediated apoptosis in thyroid epithelial cells. Cell Death Differ 2002; 9: 274–286.
Tanaka F, Kawakami A, Tamai M, Nakamura H, Iwanaga N, Izumi Y et al. IFN-gamma/JAK/STAT pathway-induced inhibition of DR4 and DR5 expression on endothelial cells is cancelled by cycloheximide-sensitive mechanism: novel finding of cycloheximide- regulating death receptor expression. Int J Mol Med 2005; 15: 833–839.
Shin EC, Seong YR, Kim CH, Kim H, Ahn YS, Kim K et al. Human hepatocellular carcinoma cells resist to TRAIL-induced apoptosis, and the resistance is abolished by cisplatin. Exp Mol Med 2002; 34: 114–122.
Horak P, Pils D, Kaider A, Pinter A, Elandt K, Sax C et al. Perturbation of the tumor necrosis factor--related apoptosis-inducing ligand cascade in ovarian cancer: overexpression of FLIPL and deregulation of the functional receptors DR4 and DR5. Clin Cancer Res 2005; 11: 8585–8591.
Mondelli M, Tedder RS, Ferns B, Pontisso P, Realdi G, Alberti A . Differential distribution of hepatitis B core and E antigens in hepatocytes: analysis by monoclonal antibodies. Hepatology 1986; 6: 199–204.
Eckhardt SG, Milich DR, McLachlan A . Hepatitis B virus core antigen has two nuclear localization sequences in the arginine-rich carboxyl terminus. J Virol 1991; 65: 575–582.
Hatton T, Zhou S, Standring DN . RNA- and DNA-binding activities in hepatitis B virus capsid protein: a model for their roles in viral replication. J Virol 1992; 66: 5232–5241.
Liao W, Ou JH . Phosphorylation and nuclear localization of the hepatitis B virus core protein: significance of serine in the three repeated SPRRR motifs. J Virol 1995; 69: 1025–1029.
Whitten TM, Quets AT, Schloemer RH . Dentification of the hepatitis B virus factor that inhibits expression of the β interferon gene. J virol 1991; 65: 4699–4704.
Shetty S, Graham BA, Brown JG, Hu X, Vegh-Yarema N, Harding G et al. Transcription factor NF-kappaB differentially regulates death receptor 5 expression involving histone deacetylase 1. Mol Cell Biol 2005; 25: 5404–5416.
Takimoto R, El-Deiry WS . Wild-type p53 transactivates the KILLER/DR5 gene through an intronic sequence-specific DNA-binding site. Oncogene 2000; 19: 1735–1743.
Burns TF, Bernhard EJ, El-Deiry WS . Tissue specific expression of p53 target genes suggests a key role for KILLER/DR5 in p53-dependent apoptosis in vivo. Oncogene 2001; 20: 4601–4612.
Yoshida T, Maeda A, Tani N, Sakai T . Promoter structure and transcription initiation sites of the human death receptor 5/TRAIL-R2 gene. FEBS Lett 2001; 507: 381–385.
Barone M, Maiorano E, Ladisa R, Cuomo R, Pece A, Berloco P et al. Influence of ursodeoxycholate-enriched diet on liver tumor growth in HBV transgenic mice. Hepatology 2003; 37: 880–886.
Billaud JN, Peterson D, Lee BO, Maruyama T, Chen A, Sallberg M et al. Advantages to the use of rodent hepadnavirus core proteins as vaccine platforms. Vaccine 2007; 25: 1593–1606.
Liu F, Song Y, Liu D . Hydrodynamics-based transfection in animals by systemic administration of plasmid DNA. Gene Ther 1999; 6: 1258–1266.
Acknowledgements
We are grateful to Professor Sakai and Professor Yoshida from Department of Molecular Targeting Cancer Prevention, Kyoto Prefecture University, for providing the recombinant plasmid pDR5PF. We thank professor Kakar (University of Louisville), kindly providing the pGL3-p53 plasmid .We thank Professor Yuan Wang, from Institute of Biochemistry and Cell Biology, Chinese Academy of Sciences, for kindly offering the recombinant plasmid p3.6 II.This work was supported by grants from the Natural Science Foundation of China (Grant no.30700973, no.30772031, no.30128023 and no.30700357) and National Education Ministry of China (no. 20030422056).
Author information
Authors and Affiliations
Corresponding authors
Additional information
Edited by J Tschop
Supplementary Information accompanies the paper on Cell Death and Differentiation website (http://www.nature.com/cdd)
Rights and permissions
About this article
Cite this article
Du, J., Liang, X., Liu, Y. et al. Hepatitis B virus core protein inhibits TRAIL-induced apoptosis of hepatocytes by blocking DR5 expression. Cell Death Differ 16, 219–229 (2009). https://doi.org/10.1038/cdd.2008.144
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/cdd.2008.144
Keywords
This article is cited by
-
Hepatitis B virus core protein promotes the expression of neuraminidase 1 to facilitate hepatocarcinogenesis
Laboratory Investigation (2020)
-
Role of Core/Capsid Inhibitors in Functional Cure Strategies for Chronic Hepatitis B
Current Hepatology Reports (2020)
-
Tumor cell-intrinsic Tim-3 promotes liver cancer via NF-κB/IL-6/STAT3 axis
Oncogene (2018)
-
CUL4A facilitates hepatocarcinogenesis by promoting cell cycle progression and epithelial-mesenchymal transition
Scientific Reports (2015)
-
Molecular mechanisms of hepatic apoptosis
Cell Death & Disease (2014)