Abstract
Apical periodontitis, dominated by dense inflammatory infiltrates and increased osteoclast activities, can lead to alveolar bone destruction and tooth loss. It is believed that miRNA participates in regulating various biological processes, osteoclastogenesis included. This study aims to investigate the differential expression of miRNAs in rat apical periodontitis and explore their functional target genes. Microarray analysis was used to identify differentially expressed miRNAs in apical periodontitis. Bioinformatics technique was applied for predicting the target genes of differentially expressed miRNAs and their biological functions. The result provided us with an insight into the potential biological effects of the differentially expressed miRNAs and showed particular enrichment of target genes involved in the MAPK signaling pathways. These findings may highlight the intricate and specific roles of miRNA in inflammation and osteoclastogenesis, both of which are key aspects of apical periodontitis, thus contributing to the future investigation into the etiology, underlying mechanism and treatment of apical periodontitis.
Similar content being viewed by others
Introduction
Maxillofacial bones, like other skeletal tissues in vertebral bodies, are principal structural connective tissues constituting the contour of the organism and protecting key organs. Different from the axial skeleton and appendicular skeleton, the main biological significance of maxillofacial bones, especially maxillar and mandible, is supporting or conducting of the masticatory forces, as well as maintenance of the maxillofacial aesthetics and oral cavity. Despite of the differences in function, they all share the similar mechanism in bone biology.
It is well known that bone morphology and function in postnatal vertebrates is maintained through the balancing activities of the bone-forming osteoblasts (which are part of a group of cells broadly referred to as osteoblast lineage cells) and the bone-resorbing osteoclasts, a process namely bone remodeling. The balance between the two activities is under the spatio-temporal control of several signaling pathways (1–5). Any interruption of the balance between osteoblasts and osteoclasts can compromise bone remodeling. Chronic inflammatory diseases in oromaxillofacial region, such as periodontitis and apical periodontitis, are characterized by abundant osteoclast proliferation and enhanced resorptive activity relative to formation, thus lead to bone loss and tissue destruction in different sites (6–8).
Apical periodontitis, an infectious oral disease represents aggressive immune and inflammatory activation, can be viewed as the result of a dynamic encounter between microbes and host defense. It has already been proved that apical peiodontitis is dominated by dense inflammatory infiltrates and increased osteoclast activities (9,10). Lymphocytes, macrophages and neutrophils have been identified in focus and the number or activity of osteoclasts elevated (11,12). This disturbs the bone remodeling by enhancing bone resorption relative to formation (13–16). Apical periodontitis can lead to alveolar bone defect and tooth loss. Left untreated, the disease can cause systemic disorders such as rheumatoid arthritis, cardiovascular diseases, and diabetes (17,18). The understanding of the pathogenesis and treatment is of importance to the population. However, the mechanisms of host immuno-inflammatory responses and osteoclastogenesis in apical periodontitis remain unclear. Recently, accumulating evidence demonstrates that miRNAs play a critical role in these processes (19).
miRNAs are small non-coding RNAs of approximately 18–22 nucleotides (nt) that regulate gene function post-transcriptionally (20,21). miRNAs are transcribed from endogenous miRNA genes and generate primary (pri-) miRNAs. Pri-miRNAs are processed into single hairpins or precursor miRNAs (pre-miRNAs) by the RNAase III enzyme Drosha in the nucleus. Pre-miRNAs are then shuttled into the cytoplasm by Exportin-5 and further processed by the RNAase enzyme Dicer to generate mature miRNAs. miRNAs function in the form of ribonucleoproteins called miRISCs (miRNA-inducing silencing complexes) (21), which comprise Argonaute and GW-182 family proteins. miRISCs use the miRNAs as guides for the sequence-specific silencing of messenger RNAs that contain complementary sequence through inducing the degradation of the mRNAs or repressing their translation (22–24). miRNAs are able to regulate the expression of multiple targets by binding to the 3'-UTR of genes. Thus, miRNAs mediate rapid fine-tuning of gene expression by targeting existed mRNAs. There are several studies have been carried out in order to understand how miRNAs influence osteoclatogenesis (25–30). The global loss of miRNA activity in osteoclast precursor cells causes a blockage in osteoclastogenesis that absence of mature miRNAs by conditional knock-out model in mononuclear preosteoclasts and in mature multinucleated osteoclasts results in increased bone mass due to decreased number and activity of osteoclasts (25,31,32).
There are two principal facts of microRNAs that should be kept in mind that one miRNA can target hundreds of mRNAs and act selectively in different tissues, whereas one mRNA can have multiple mass binding sites for different miRNAs (33,34). These two fundamental properties predict that miRNAs function as powerful molecular managers that operate by controlling several gene regulatory networks simultaneously (20). Meanwhile, it raises the concern that analysis of microRNA in bone homeostasis should be an informatic integration. Bioinformatic analytic tools provide such possibility. To this end, the roles of miRNAs in the differentiation and recruitment of osteoclasts derived from hematopoietic stem cells remains to be established.
Here, we report the identification of miRNAs that are differentially expressed during apical periodontitis. By comparing the bone lesion regions in the rat molars, we have identified miRNAs that may play a role in the osteoclastogenesis during this process.
Materials and Methods
Ethics Statement
Experiments were performed according to the Regulations for the Administration of Affairs Concerning Experimental Animals (Ministry of Science and Technology, China, revised in June 2004). The study is approved by Ethical Committees of West China School of Stomatology, Sichuan University and State Key Laboratory of Oral Diseases. The rats were allowed access to feed and water ad libitum under normal conditions and humanely sacrificed as necessary.
Animals and Sample Collection
The Sprague-Dawley rats used in this experiment were obtained from the Dashuo Laboratory Animal Company of Sichuan, China. The induction of periapical lesions was performed as described previously (35,36). Briefly, four-week-old Sprague-Dawley rats were anaesthetized with 10% chloral hydrate (0.3 mL per 100 g animal weight) by intraperitoneal injection, and mounted on a jawretraction board. The pulps of bilateral mandibular first molars were exposed to the oral environment with a No. 1/4 dental round bur under a surgical microscope. Rats were allowed to recover and were maintained under standard conditions until they were sacrificed at 28 days after pulp exposure. Rats without pulp exposures were established as controls. 28 days after the surgery, animals were sacrificed and bone blocks containing the mandibular first molars' roots were dissected, gently cleaned of soft tissue and preserved in RNAlater (Life Technology Corporation, Foster city, CA, USA) at −80°C until processing. Bone blocks were ground in pre-cooled sterile mortars and pestles in liquid nitrogen. Total RNA including small RNAs were extracted using the miRNeasy Mini kit (Qiagen, Valencia, CA, USA) according to manufacturer's instructions. The same RNA samples were used for the experiments in both microarray and qRT-PCR assays.
microRNA Microarray Analysis
In this study, bone blocks of apical periodontitis and of negative controls with four biological repeats for each were tested. The RNA was quantitated by using the NanoDrop (Thermo Scientific, Wilmington, DE, USA) and Agilent 2100 Bioanalyzer (Agilent Technologies Inc, Wilmington, DE, USA) to determine the quantity and purity of the samples. Agilent microarray hybridization was carried out by the ShanghaiBio Corporation. miRNA microarray profiling was performed as previously described. Data analysis was performed by using Gene-Spring software 11.0 (Agilent Technologies, Santa Clara, CA, US). A miRNA was designated as highly expressed if expression in apical periodontitis was more than 1.5-fold compared to that in negative controls. miRNAs considered to be significantly differentially expressed were then included for further analysis. The correlation analysis was used to inquire the correlation between any two of all eight samples.
Validation of microarray Results by qRT-PCR
The purified total RNA including small RNAs were used as templates. Reverse-transcription was performed with the TaqMan MicroRNA Reverse Transcription Kit using small RNA-specific RT primer. The reactions were incubated at 16 °C for 30 min, 42 °C for 30 min and 85 °C for 5 min, chilled on ice for 5 min, and the cDNA was stored at −20 °C. The qRT-PCR was performed with the TaqMan Small RNA Assay following the manufacturer's instructions in 20 μL reaction mixtures. U6 was used as endogenous control to normalize Ct values obtained for each gene. miRNA expression was compared by ΔΔCt. Data were compared by one-way ANOVA followed by the post-hocTukey's test.
Target Prediction and Functional Analysis
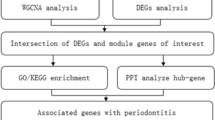
Three types of miRNA target prediction databases, including TargetScan (http://www.targetscan.org/) and miRanda algorithm base (http://www.microrna.org/microrna/home.do), were used to predict the target genes of differentially expressed miRNAs. Unique profiles were defined and significant profiles selected (P<0.01). The intersection of these two datasets was assayed according to the prediction results. To determine the potential biological functions and pathways, GO terms and Pathway annotation of the miRNA targets were found using the DAVID database (http://david.abcc.ncifcrf.gov/content.jsp) and the SBC Analysis System annotation tool (http://sas.ebioservice.com/), respectively.
Statistical Analysis
All experiments were performed independently at least three times in triplicate. Relative miRNA levels were calculated in comparison to internal U6 standards. Numerical data are presented as mean±SD. The difference between means was analyzed with one-way ANOVA. Differences were considered significant when P<0.05. All statistical analyses were done with the software SPSS19.0 (SPSS Inc. Chicago, USA).
Results
Identification of Differentially Expressed miRNAs in Rat Apical Periodontitis
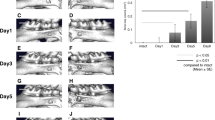
Total RNA including small RNAs extracted from bone blocks of apical periodontitis and of negative controls, was sent to ShanghaiBio Corporation for microarray hybridization (Figure 1A and 1B). Based on the microarray analysis, a total of 49 miRNAs were detected to be differentially expressed in bone lesions of apical periodontitis. 21 miRNAs were down-regulated while 28 were up-regulated in apical periodontitis compared with control animals (P<0.05) (Figure 2A). Correlation analysis, on the basis of miRNA expression profiles, showed variance between the 8 samples (Figure 2B). Although apical periodontitis groups and control groups represent similarity within groups, the miRNA expression profile of apical periodontitis sample 1, sample 2 and sample 4 are distinct from samples from control groups, as well as apical periodontitis sample 3 (Figure 2B). In addition, 21 miRNAs were recognized as significantly differentially expressed with 7 down-regulated and 14 up-regulated (P<0.01) (Table 1).
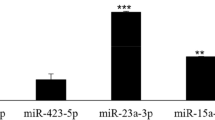
The expression profiles of miRNAs in rat apical periodontitis were analyzed using miRNA microarray. A: A heatmap generated from microarray data reflecting miRNA expression values. 49 miRNAs were differentially expressed at least 1.5-fold (P<0.05) between apical periodontitis group and control group. Bar graph showed fold changes. B: Correlation analysis showed the miRNA expression profile of apical periodontitis sample 1, sample 2 and sample 4 are distinct from samples from control groups, as well as apical periodontitis sample 3. C: Validation of microarray data by qRT-PCR. The fold change of 3 down-regulated miRNAs (rno-miR-17-5p, rno-miR-130b, rno-miR-142) and 2 up-regulated miRNAs (rno-miR-130a, rno-miR-132) were verified. * Statistical significance between apical periodontitis group and control group, P<0.05.
Confirmation of Differentially Expressed miRNAs by realtime qPCR
From the differentially expressed miRNAs identified from apical periodontitis and control groups, we selected 5 miRNAs for confirmation by qPCR. The miRNAs were chosen based on high differential expression and included both those with increases and decreases in expression fold-differences. All the miRNAs assayed by qPCR were confirmed to be differentially expressed (Figure 2C), which validated the microarray approach. Thus, the array results for all differentially expressed miRNAs are expected to be accurate. The predicted targets of these chosen microRNAs were retrieved from two target prediction databases mentioned before. Only the predicted targets found in both prediction databases were retained (Table 2).
Target Prediction and Functional Annotation
In order to analyze the potential functions of these miRNA-target genes, GO (Gene Ontology) term and pathway analyses were applied. For each differentially expressed, array-identified miRNA (P<0.01), the predicted targets were retrieved from two target prediction databases.
The GO enrichment analysis is a powerful tool by which these genes are correlated with biological processes, cellular components and molecular functions. GO terms with P<0.001 are considered statistically significant. FDR (false discovery rate) is also taken into consideration to define P value threshold at which the FDR would be less than 25%. Under “Biological Processes” category of GO classification (Figure 3), positive regulation of nucleobase, nucleoside, nucleotide and nucleic acid metabolic process, positive regulation of nitrogen compound metabolic process and positive regulation of developmental process are the top three over-represented terms (Figure 3 and 4). The GO terms of positive regulation of macromolecule biosynthetic process transmembrane receptor protein tyrosine kinase signaling pathway and phosphate metabolic process are statistically significant (Figure 3 and 4). These categories have been described as associated with macrophage differentiation (37,38), which is closely related to osteoclast precursor (39). The molecular function subcategories are most notably associated with various binding functions such as enzyme binding, kinase binding and protein kinase binding indicating (Figure 5A). Within the GO category of cellular components, the terms, Golgi apparatus, cell fraction and membrane fraction are prominent (Figure 5B).
Gene Ontology analysis of target genes of 21 differential miRNAs (P<0.01). GO terms with P<0.001 and FDR≤25% were taken into consideration. GO terms overrepresented and associated with apical periodontitis under biological process category. The numbers of the pie's every section represented the counts of putative target genes mapping to corresponding GO terms.
Regarding pathway analysis, the gene sets of Biocarta and KEGG pathways were used. There are 42 and 54 terms statistically significant in Biocarta and KEGG pathway, respectively (P<0.001) (Table 3 and 4). Some of these pathways were consistent with biological processes already revealed by GO analysis. MAPK Signaling Pathway, EGF Signaling Pathway, HIV-I Nefnegative effector of Fas and TNF in Biocarta and Pathways in cancer, Metabolic pathways, MAPK signaling pathway in KEGG were highly enriched (Figure 6A and 6B). The functional identity of the target genes, as revealed by different bioinformatic approaches confirmed that miRNAs may have important implications for osteoclatogenesis through their effects on signaling pathways. Erk1Erk2 Mapk Signaling pathway, p38 MAPK Signaling Pathway and Angiotensin II mediated activation of JNK Pathway via Pyk2 dependent signaling are statically significant as well (Figure 6A). Furthermore, in our prediction, T cell receptor signaling pathway, Toll-like receptor signaling pathway and B cell receptor signaling pathway are as well statistically significant (Figure 6B).
Discussion
Apical periodontitis is characterized by inflammation and elevated osteoclastic activity. Alveolar bone loss surrounding the tooth root is result of disease in some intractable cases and the prognosis could be negative even with periodically endodontic follow-up (40). Recent studies indicated that miRNAs are involved in chronic inflammatory diseases, such as arthritis and periodontitis (41–44). With apparently different etiology and clinical features, the involvement of microRNAs in apical periodontitis and the underlying mechanism remained unexplored. In this present study, microarray analysis is applied to screen and compare differentially expressed miRNAs in rat apical periodontitis. Our results might contribute to the identification of regulatory factors involved in apical periodontitis and inflammation.
Regardless of etiology, diseases with bone loss always represents enhanced bone resorption relative to formation, hence the progress in understanding the pathogenesis and successful treatment of this family of diseases requires an understanding of osteoclast biology. Bone resorption is a multistep process initiated by the proliferation of immature osteoclast precursors, the commitment of these cells to the osteoclast phenotype, and finally, degradation of the organic and inorganic phases of bone by the mature resorptive cells (45). Each stage of osteoclast differentiation requires regulatory factors, that also serve as markers of osteoclast function, and bone resorbing activity (46–48), and could be regulated by microRNAs through the specific targeting with the protein-coding gene mRNAs.
According to microarray analysis, a total of 49 (P<0.05) and 21 (P<0.01) miRNAs were differentially expressed in apical periodontitis. In order to validate the microarray result, we selected 5 differentially expressed miRNAs for qRT-PCR assay. These microRNAs are predicted to be targeting on osteoclastogenesis associated genes. The Correlation Analysis was performed for the similarities and differences in miRNA expression profiles between groups. Although significant variance was observed between the apical periodontitis groups and the control groups, it also showed a difference between sample 3 and the other 3 samples in apical periodontitis group. The variability of the different samples indicated that the miRNA expression profiles may vary between individuals.
In conventional gene expression data analysis, enrichment analysis of differentially expressed genes has been successfully utilized to explore the underlying signaling pathways and genes. Target genes of the 21 differentially expressed miRNAs (P<0.01) were mapped to gene function databases, and the collected 374 putative target genes were performed in the GO (Gene Ontology) and pathway analysis.
GO terms of positive regulation of nitrogen compound metabolic process and positive regulation of developmental process are overrepresented. Terms of positive regulation of macromolecule biosynthetic process, transmembrane receptor protein tyrosine kinase signaling pathway and phosphate metabolic process associated with osteoclastogenesis were statistically significant changed as well (Figure 2B). Inducible nitric oxide synthase (iNOS) is well known for the contribution to osteoclast activity and alveolar bone loss. Nitric oxide (NO) mediates adaptive bone formation, protects osteocytes against apoptosis and mediates osteoclastic activity (49–51). GO results enlightened the role of miRNA in regulating nitrogen compound related genes and furthermore, in regulating bone destruction and remolding. Macromolecule synthesis is the main housekeeping function of immune cells (52). Macromolecule molecules participate in inducing expression of the proinflammatory cytokines (53). These cytokines, such as TNF-α, IL-1β, IL-6, IL-17 etc, have been demonstrated to promote osteoclast differentiation and function either directly or indirectly, by acting on cells of the osteoclastlineage or by acting on other cell types to regulate expression of the key osteoclastogenic factor receptor activator of nuclear factor kappaB ligand (RANKL) and/or its inhibitor osteoprotegerin (OPG) (54,55).
Among the panels of target genes fall into GO subcategories of transmembrane receptor protein tyrosine kinase signaling pathway and phosphate metabolic process, the most notable is CSF1R. CSF1R (also termed c-Fms) is recognizable for CSF1, which is more widely known as M-CSF. The biological effects of M-CSF are mediated by CSF1R (56) and supports survival and proliferation of myeloid progenitors and promotes generation of osteoclast precursors that express RANK receptor, and thus are competent to differentiate into osteoclasts in response to RANKL recognization (57).
The GO analysis result is in line with our knowledge that apical periodontitits is the outcome of confront of stimuli and host reaction that is the activation of immune system and breakdown of the balance between osteogenesis and osteoclastogenesis inclined to enhanced osteoclast activity. To further analyze the downstream biological mechanism, pathway analysis was performed. The results of KEGG and Biocarta pathway analysis were in accordance with the GO analysis. Most of the genes targeted by significantly changed miRNAs were enriched in several signaling pathways, including MAPK signaling pathway.
It is well known that RANKL, in conjunction with M-CSF, play a pivotal role both in adaptive immunity and osteoclastogenesis, and thus provides firm molecular evidence to interconnect immune and skeletal systems. RANKL and RANK are members of the tumor necrosis factor (TNF) and TNF receptor superfamilies, respectively. Clinical studies indicated that inflammatory lesions of bone are characterized by abundant osteoclast proliferation and TNF is a potent osteoclastogenic agent which appears to mediate orthopedic implant loosening, a disorder accompanied by local secretion of TNF (58). Microbial lipopolysaccharide, which is likely to be an important pathogenetic factor in the alveolar bone loss seen in apical periodontitis, plays a critical role in stimulating the development of host immune reactions and causing severe tissue destruction (59). RANKL activates the receptor RANK in a trimeric symmetric complex with TRAF6. MAPKs (mainly including ERK, JNK and p38 MAPK) are located at the downstream of the TRAF6 signalling complexes, and they play an important role in RANKL-induced osteoclastogenesis by triggering a cascade reaction and up-regulating expressions of essential transcription factors c-Fos, MITF, PU.1 and NFATc1 (60,61).
From the expression level, target prediction, and functional analysis, the most promising miRNA appeared to be miR-130a. The potential target genes of miR-130a include TNFSF11 and CSF1. TNFSF11 is the gene that encodes the TNF family cytokine, RANKL (62). CSF1, or M-CSF, as one of the essential genes required for osteoclastognesis, is of crucial importance for the proliferation and survival of the osteoclast precursor cells. CSF1 participates in the later differentiation stage of osteoclastogenesis through activating Akt, c-Fos and ERK pathways that may interact with RANKL signals (2,63,64). Taking these results into account, it is reasonable to assume that the downregulation of miR-130a in apical periodontitis may increase the expression of TNFSF11 and CSF1, subsequently activating MAPK signaling pathway and thus promoting osteoclastic differentiation in alveolar progenitors. Future studies to validate the function and mechanism of miR-130a in osteoclastogenesis of alveolar bone marrow cells are warranted.
Due to the cherry-picking study of the microRNAs, so far there are only a few miRNAs known to contribute to osteoclast lineage commitment. Other than the global intervention of microRNA machinery, miR-155 (65), miR-146a (66), and miR-223 (28,67) were identified to be related with osteoclastogenesis. miRNAs were identified in periodontitis gingiva as well (68). Nevertheless, a lack of systemic analysis of microRNAs involvement in apical periodontitis is obvious.
In this study, we used microarray analysis to identify differentially expressed miRNAs during apical periodontitis. Bioinformatics technology provided us insight into the potential biological effects and showed particular enrichment of target genes involved in the MAPK signaling pathways. These findings may shed light on the regulatory mechanisms underlying apical periodontitis and may open new avenues for future research into the etiology, mechanism and treatment of apical periodontitis. The role of certain miRNAs in apical periodontitis is currently being further investigated.
References
Del Fattore A, Teti A, Rucci N . Osteoclast receptors and signaling. Arch Biochem Biophys. 2008;473:147–160.
Ross FP, Teitelbaum SL . alphavbeta3 and macrophage colony-stimulating factor: partners in osteoclast biology. Immunol Rev. 2005;208:88–105.
Miyazaki T, Tanaka S, Sanjay A, Baron R . The role of c-Src kinase in the regulation of osteoclast function. Mod Rheumatol. 2006;16:68–74.
Edwards CM, Mundy GR . Eph receptors and ephrin signaling pathways: a role in bone homeostasis. Int J Med Sci. 2008;5:263–272.
Sims NA, Gooi JH . Bone remodeling: Multiple cellular interactions required for coupling of bone formation and resorption. Semin Cell Dev Biol. 2008;19:444–451.
Duan L, Ren Y . Role of notch signaling in osteoimmunology—from the standpoint of osteoclast differentiation. Eur J Orthod. 2013;35:175–182.
Kim TH, Choi SJ, Lee YH, Song GG, Ji JD . Combined therapeutic application of mTOR inhibitor and vitamin D(3) for inflammatory bone destruction of rheumatoid arthritis. Med Hypotheses. 2012;79:757–760.
Sato K, Takayanagi H . Osteoclasts, rheumatoid arthritis, and osteoimmunology. Curr Opin Rheumatol. 2006;18:419–426.
Belibasakis GN, Rechenberg DK, Zehnder M . The receptor activator of NF-kappaB ligand-osteoprotegerin system in pulpal and periapical disease. Int Endod J. 2013;46:99–111.
da Silva RA, Ferreira PD, De Rossi A, Nelson-Filho P, Silva LA . Toll-like receptor 2 knockout mice showed increased periapical lesion size and osteoclast number. J Endod. 2012;38:803–813.
Marton IJ, Kiss C . Protective and destructive immune reactions in apical periodontitis. Oral Microbiol Immunol. 2000;15:139–150.
Tani-Ishii N, Wang CY, Stashenko P . Immunolocalization of bone-resorptive cytokines in rat pulp and periapical lesions following surgical pulp exposure. Oral Microbiol Immunol. 1995;10:213–219.
Artese L, Piattelli A, Quaranta M, Colasante A, Musani P . Immunoreactivity for interleukin 1-beta and tumor necrosis factor-alpha and ultrastructural features of monocytes/macrophages in periapical granulomas. J Endod. 1991;17:483–487.
Ataoglu T, Ungor M, Serpek B, Haliloglu S, Ataoglu H, Ari H . Interleukin-1beta and tumour necrosis factor-alpha levels in periapical exudates. Int Endod J. 2002;35:181–185.
Nair PN . Pathogenesis of apical periodontitis and the causes of endodontic failures. Crit Rev Oral Biol Med. 2004;15:348–381.
Reichert S, Machulla HK, Klapproth J, Zimmermann U, Reichert Y, Glaser C, Schaller HG, Schulz S . Interferon-gamma and interleukin-12 gene polymorphisms and their relation to aggressive and chronic periodontitis and key periodontal pathogens. J Periodontol. 2008;79:1434–1443.
Pizzo G, Guiglia R, Lo Russo L, Campisi G . Dentistry and internal medicine: from the focal infection theory to the periodontal medicine concept. Eur J Intern Med. 2010;21:496–502.
Zoellner H . Dental infection and vascular disease. Semin Thromb Hemost. 2011;37:181–192.
Lian JB, Stein GS, van Wijnen AJ, Stein JL, Hassan MQ, Gaur T, Zhang Y . MicroRNA control of bone formation and homeostasis. Nat Rev Endocrinol. 2012;8:212–227.
Bartel DP . MicroRNAs: genomics, biogenesis, mechanism, and function. Cell. 2004;116:281–297.
He L, Hannon GJ . MicroRNAs: small RNAs with a big role in gene regulation. Nat Rev Genet. 2004;5:522–531.
Martin R, Smibert P, Yalcin A, Tyler DM, Schafer U, Tuschl T, Lai EC . A Drosophila pasha mutant distinguishes the canonical microRNA and mirtron pathways. Mol Cell Biol. 2009;29:861–870.
Bartel DP . MicroRNAs: target recognition and regulatory functions. Cell. 2009;136:215–233.
Chiang HR, Schoenfeld LW, Ruby JG, Auyeung VC, Spies N, Baek D, Johnston WK, Russ C, Luo S, Babiarz JE, Blelloch R, Schroth GP, Nusbaum C, Bartel DP . Mammalian microRNAs: experimental evaluation of novel and previously annotated genes. Genes Dev. 2010;24:992–1009.
Sugatani T, Vacher J, Hruska KA . A microRNA expression signature of osteoclastogenesis. Blood. 2011;117:3648–3657.
Xia Z, Chen C, Chen P, Xie H, Luo X . MicroRNAs and their roles in osteoclast differentiation. Front Med. 2011;5:414–419.
Pauley KM, Cha S . miRNA-146a in rheumatoid arthritis: a new therapeutic strategy. Immunotherapy. 2011;3:829–831.
Shibuya H, Nakasa T, Adachi N, Nagata Y, Ishikawa M, Deie M, Suzuki O, Ochi M . Overexpression of microRNA-223 in rheumatoid arthritis synovium controls osteoclast differentiation. Mod Rheumatol. 2012 Aug 19. [Epub ahead of print]
Zhang J, Zhao H, Chen J, Xia B, Jin Y, Wei W, Shen J, Huang Y . Interferon-beta-induced miR-155 inhibits osteoclast differentiation by targeting SOCS1 and MITF. FEBS Lett. 2012;586:3255–3262.
Cheng P, Chen C, He HB, Hu R, Zhou HD, Xie H, Zhu W, Dai RC, Wu XP, Liao EY, Luo XH . miR-148a regulates osteoclastogenesis by targeting V-maf musculoaponeurotic fibrosarcoma oncogene homolog B. J Bone Miner Res. 2013;28:1180–1190.
Mizoguchi F, Izu Y, Hayata T, Hemmi H, Nakashima K, Nakamura T, Kato S, Miyasaka N, Ezura Y, Noda M . Osteoclast-specific Dicer gene deficiency suppresses osteoclastic bone resorption. J Cell Biochem. 2010;109:866–875.
Sugatani T, Hruska KA . Impaired micro-RNA pathways diminish osteoclast differentiation and function. J Biol Chem. 2009;284:4667–4678.
Jackson RJ, Standart N . How do microRNAs regulate gene expression? Sci STKE. 2007;2007:re1.
Hon LS, Zhang Z . The roles of binding site arrangement and combinatorial targeting in microRNA repression of gene expression. Genome Biol. 2007;8:R166.
Stashenko P, Teles R, D'Souza R . Periapical inflammatory responses and their modulation. Crit Rev Oral Biol Med. 1998;9:498–521.
Sasaki H, Balto K, Kawashima N, Eastcott J, Hoshino K, Akira S, Stashenko P . Gamma interferon (IFN-gamma) and IFN-gamma-inducing cytokines interleukin-12 (IL-12) and IL-18 do not augment infection-stimulated bone resorption in vivo . Clin Diagn Lab Immunol. 2004;11:106–110.
Way KJ, Dinh H, Keene MR, White KE, Clanchy FI, Lusby P, Roiniotis J, Cook AD, Cassady AI, Curtis DJ, Hamilton JA . The generation and properties of human macrophage populations from hemopoietic stem cells. J Leukoc Biol. 2009;85:766–778.
Martinez FO, Gordon S, Locati M, Mantovani A . Transcriptional profiling of the human monocyte-to-macrophage differentiation and polarization: new molecules and patterns of gene expression. J Immunol. 2006;177:7303–7311.
Teitelbaum SL . Osteoclasts: what do they do and how do they do it? Am J Pathol. 2007;170:427–435.
Wang J, Jiang Y, Chen W, Zhu C, Liang J . Bacterial flora and extraradicular biofilm associated with the apical segment of teeth with post-treatment apical periodontitis. J Endod. 2012 38:954–959.
Furer V, Greenberg JD, Attur M, Abramson SB, Pillinger MH . The role of microRNA in rheumatoid arthritis and other autoimmune diseases. Clin Immunol. 2010;136:1–15.
Nakasa T, Nagata Y, Yamasaki K, Ochi M . A mini-review: microRNA in arthritis. Physiol Genomics. 2011;43:566–570.
Lee YH, Na HS, Jeong SY, Jeong SH, Park HR, Chung J . Comparison of inflammatory microRNA expression in healthy and periodontitis tissues. Biocell. 2011;35:43–49.
Xie YF, Shu R, Jiang SY, Liu DL, Zhang XL . Comparison of microRNA profiles of human periodontal diseased and healthy gingival tissues. Int J Oral Sci. 2011;3:125–134.
Teitelbaum SL . Bone resorption by osteoclasts. Science. 2000;289:1504–1508.
Tsurukai T, Udagawa N, Matsuzaki K, Takahashi N, Suda T . Roles of macrophage-colony stimulating factor and osteoclast differentiation factor in osteoclastogenesis. J Bone Miner Metab. 2000;18:177–184.
Shiotani A, Takami M, Itoh K, Shibasaki Y, Sasaki T . Regulation of osteoclast differentiation and function by receptor activator of NFkB ligand and osteoprotegerin. Anat Rec. 2002;268:137–146.
Czupalla C, Mansukoski H, Pursche T, Krause E, Hoflack B . Comparative study of protein and mRNA expression during osteoclastogenesis. Proteomics. 2005;5:3868–3875.
Herrera BS, Martins-Porto R, Maia-Dantas A, Campi P, Spolidorio LC, Costa SK, van Dyke TE, Gyurko R, Muscara MN . iNOS-derived nitric oxide stimulates osteoclast activity and alveolar bone loss in ligature-induced periodontitis in rats. J Periodontol. 2011;82:1608–1615.
Kim YS, Kang SJ, Kim JW, Cho HR, Moon SB, Kim KY, Lee HS, Han CH, Ku SK, Lee YJ . Effects of Polycan, a beta-glucan, on experimental periodontitis and alveolar bone loss in Sprague-Dawley rats. J Periodontal Res. 2012;47:800–810.
Brandi ML, Hukkanen M, Umeda T, Moradi-Bidhendi N, Bianchi S, Gross SS, Polak JM, MacIntyre I . Bidirectional regulation of osteoclast function by nitric oxide synthase isoforms. Proc Natl Acad Sci U S A. 1995;92:2954–2958.
Spies CM, Straub RH, Buttgereit F . Energy metabolism and rheumatic diseases: from cell to organism. Arthritis Res Ther. 2012;14:216.
Sanchez-Pernaute O, Filkova M, Gabucio A, Klein M, Maciejewska-Rodrigues H, Ospelt C, Brentano F, Michel BA, Gay RE, Herrero-Beaumont G, Gay S, Neidhart M, Juengel A . Citrullination enhances the proinflammatory response to fibrin in rheumatoid arthritis synovial fibroblasts. Ann Rheum Dis. 2012 Dec 12. [Epub ahead of print]
Walsh NC, Crotti TN, Goldring SR, Gravallese EM . Rheumatic diseases: the effects of inflammation on bone. Immunol Rev. 2005;208:228–251.
Braun T, Schett G . Pathways for bone loss in inflammatory disease. Curr Osteoporos Rep. 2012;10:101–108.
Arai F, Miyamoto T, Ohneda O, Inada T, Sudo T, Brasel K, Miyata T, Anderson DM, Suda T . Commitment and differentiation of osteoclast precursor cells by the sequential expression of c-Fms and receptor activator of nuclear factor kappaB (RANK) receptors. J Exp Med. 1999;190:1741–1754.
Hume DA, MacDonald KP . Therapeutic applications of macrophage colony-stimulating factor-1 (CSF-1) and antagonists of CSF-1 receptor (CSF-1R) signaling. Blood. 2012;119:1810–1820.
Merkel KD, Erdmann JM, McHugh KP, Abu-Amer Y, Ross FP, Teitelbaum SL . Tumor necrosis factor-alpha mediates orthopedic implant osteolysis. Am J Pathol. 1999;154:203–210.
Okahashi N, Inaba H, Nakagawa I, Yamamura T, Kuboniwa M, Nakayama K, Hamada S, Amano A . Porphyromonas gingivalis induces receptor activator of NF-kappaB ligand expression in osteoblasts through the activator protein 1 pathway. Infect Immun. 2004;72:1706–1714.
Yen ML, Hsu PN, Liao HJ, Lee BH, Tsai HF . TRAF-6 dependent signaling pathway is essential for TNF-related apoptosis-inducing ligand (TRAIL) induces osteoclast differentiation. PLoS One. 2012;7:e38048.
Asagiri M, Takayanagi H . The molecular understanding of osteoclast differentiation. Bone. 2007;40:251–264.
Xiong J, Onal M, Jilka RL, Weinstein RS, Manolagas SC, O'Brien CA . Matrix-embedded cells control osteoclast formation. Nat Med. 2011;17:1235–1241.
Wiktor-Jedrzejczak W, Bartocci A, Ferrante AW Jr., Ahmed-Ansari A, Sell KW, Pollard JW, Stanley ER . Total absence of colony-stimulating factor 1 in the macrophage-deficient osteopetrotic (op/op) mouse. Proc Natl Acad Sci U S A. 1990;87:4828–4832.
Yoshida H, Hayashi S, Kunisada T, Ogawa M, Nishikawa S, Okamura H, Sudo T, Shultz LD, Nishikawa S . The murine mutation osteopetrosis is in the coding region of the macrophage colony stimulating factor gene. Nature. 1990;345:442–444.
Bluml S, Bonelli M, Niederreiter B, Puchner A, Mayr G, Hayer S, Koenders MI, van den Berg WB, Smolen J, Redlich K . Essential role of microRNA-155 in the pathogenesis of autoimmune arthritis in mice. Arthritis Rheum. 2011;63:1281–1288.
Nakasa T, Shibuya H, Nagata Y, Niimoto T, Ochi M . The inhibitory effect of microRNA-146a expression on bone destruction in collagen-induced arthritis. Arthritis Rheum. 2011;63:1582–1590.
Fazi F, Rosa A, Fatica A, Gelmetti V, De Marchis ML, Nervi C, Bozzoni I . A minicircuitry comprised of microRNA-223 and transcription factors NFI-A and C/EBPalpha regulates human granulopoiesis. Cell. 2005;123:819–831.
Stoecklin-Wasmer C, Guarnieri P, Celenti R, Demmer RT, Keb-schull M, Papapanou PN . MicroRNAs and their target genes in gingival tissues. J Dent Res. 2012;91:934–940.
Acknowledgements
This study was supported by JCPT2011-9 and SKLODSCU 20130012 to XDZ, NSFC grant 81200760 and SKLODSCU 20130014 to LWZ.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Gao, B., Zheng, L. microRNA Expression in Rat Apical Periodontitis Bone Lesion. Bone Res 1, 170–185 (2013). https://doi.org/10.4248/BR201302006
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.4248/BR201302006