Abstract
Background:
Anoctamin (ANO)/transmembrane member 16 (TMEM16) proteins mediate diverse physiological and pathophysiological functions including cancer cell proliferation. The present study aimed to identify the role of ANOs in pancreatic cancer.
Methods:
In an initial screen of ANOs, ANO9/TMEM16J was overexpressed in pancreatic cancer cells, and its role in the pathogenesis of pancreatic cancer was evaluated using an integrated in vitro and in vivo approach. To determine clinical relevance of the experimental findings, the prognostic value of ANO9 was evaluated in patients with pancreatic cancer.
Results:
The ANO9 mRNA and protein levels were increased in pancreatic cancer-derived cells. Exogenous expression of ANO9 in PANC-1 cells significantly increased cell proliferation in cell cultures and in mice. In contrast, knockdown of ANO9 in AsPC-1, BxPC-3, and Capan-2 cells strongly inhibited cell proliferation. Mechanistic analysis suggested that physical association of ANO9 with epidermal growth factor receptor (EGFR) underlies ANO9-induced cell proliferation. Knockdown of ANO9 augmented the effects of the EGFR inhibitor and the cytotoxic agent on pancreatic cancer cell proliferation. In addition, high ANO9 expression is a poor prognostic factor in patients with pancreatic cancer.
Conclusions:
The ANO9/TMEM16J appears to be a clinically useful prognostic marker for pancreatic cancer and a potential therapeutic target.
Similar content being viewed by others
Main
Pancreatic cancer, a leading cause of cancer death, is difficult to treat because clinical presentation is often late, and the disease is resistant to conventional chemotherapy (Stocken et al, 2008; Ryan et al, 2014). Approximately 57% of patients with pancreatic cancer have distant metastasis at diagnosis, and the 5-year survival rate is only 2% (Siegel et al, 2015). Studies of pancreatic cancer have focussed on the KRAS oncogene, because >90% of pancreatic ductal adenocarcinomas (PDACs) carry mutations in this gene. Mutations in TP53, CDKN2A, SMAD4, MLL3, TGFBR2, ARID1A, SF3B1, and genes involved in chromatin modification and DNA damage repair systems also contribute to disease development and progression (Biankin et al, 2012; Zorde Khvalevsky et al, 2013; Ryan et al, 2014). However, an effective drug target for pancreatic cancer has not yet been developed.
The transmembrane member 16 (TMEM16) family of proteins, also known as anoctamins (ANOs), mediate diverse physiological functions such as ion transport, phospholipid movement across the membrane, and regulation of other membrane proteins (Caputo et al, 2008; Pedemonte and Galietta, 2014; Picollo et al, 2015). Since ANO1/TMEM16A was identified as a Ca2+-activated Cl− channel (CaCC) in 2008 (Caputo et al, 2008; Schroeder et al, 2008; Yang et al, 2008), the biological roles and clinical implications of other ANO/TMEM16 proteins have been described. For example, ANO2/TMEM16B and ANO6/TMEM16F were also found to possess CaCC activity (Pedemonte and Galietta, 2014; Kim et al, 2015), and ANO3/TMEM16C facilitates sodium-activated potassium currents in primary sensory neurons of rats and regulates pain processing (Huang et al, 2013). Recessive mutations of ANO5/TMEM16E cause proximal limb girdle muscular dystrophy and distal non-dysferlin Miyoshi muscular dystrophy (Bolduc et al, 2010). The ANO6 possesses phospholipid scramblase activity that is required for lipid scrambling in platelets during blood coagulation (Pedemonte and Galietta, 2014). In addition, ANO10/TMEM16K is associated with autosomal recessive cerebellar ataxia (Vermeer et al, 2010).
Some ANOs have been shown to be involved in the development of cancers. For example, overexpression of ANO1 has been observed in gastrointestinal stromal tumours, breast cancer, and head and neck cancer (Duvvuri et al, 2012; Liu et al, 2012; Britschgi et al, 2013; Simon et al, 2013; Sauter et al, 2015) and is correlated with poor prognosis in breast cancer and head and neck cancer (Duvvuri et al, 2012; Britschgi et al, 2013). The ANO6 is associated with the metastatic potential of breast cancer (Dutertre et al, 2010), and ANO7/TMEM16G is upregulated in prostate cancer (Bera et al, 2004). In addition, ANO9/TMEM16J, also known as tumour protein p53-inducible protein 5 (TP53I5), was found to be expressed in human colorectal, lung, and breast cancers in an in silico screening of p53-associated genes (Katoh and Katoh, 2004). However, its biological role and clinical relevance in cancer are not known.
In a preliminary study, we examined the expression of all human ANO members (ANO1–ANO10) in normal pancreatic cells and pancreatic cancer cells to identify protein(s) associated with pancreatic cancer. Our results showed that ANO9 is overexpressed in pancreatic cancer cells, and high ANO9 expression is a poor prognostic factor in patients with pancreatic cancer. In addition, we examined the role of ANO9 in the pathogenesis of pancreatic cancer using an integrated in vitro, in vivo, and human clinical approach.
Materials and methods
Cell culture, plasmids, and antibodies
The PANC-1 and HEK293T cells were maintained in high-glucose Dulbecco’s modified Eagle’s medium (DMEM) supplemented with 10% fetal bovine serum and 1% penicillin and streptomycin (Invitrogen, Carlsbad, CA, USA). The AsPC-1, BxPC-3, and Capan-2 cells were maintained in Roswell Park Memorial Institute (RPMI) medium supplemented with 10% fetal bovine serum and 1% penicillin and streptomycin. Immortalised hTERT-HPNE cells were obtained from American Type Culture Collection (Manassas, VA, USA) and maintained in 75% DMEM and 25% Medium M3 Base (Incell Corp, San Antonio, TX, USA; Cat. No. M300F-500) supplemented with 5% fetal bovine serum, 10 ng ml−1 human recombinant epidermal growth factor, and 750 ng ml−1 puromycin. Cells were grown at 37 °C in a 5% CO2 incubator. Human ANO9 cDNA was purchased from GE Dharmacon (Lafayette, CO, USA; Clone ID: 40122929). The coding regions of ANO9 were subcloned into the pCMV-Myc-N vector with an N-terminal Myc-tag using EcoRI and XhoI restriction enzymes. The coding regions of ANO9 were also subcloned into the doxycycline-inducible (Tet-On) pTRIPZ lentiviral vector using AgeI and EcoRI restriction enzymes. Short hairpin RNA (shRNA) targeting ANO9 (base sequence: 5′-GAUACAGGGACUACCGCAA-3′) was also inserted into pTRIPZ. To produce replication-incompetent lentiviral particles, pTRIPZ vectors containing ANO9 cDNA or shRNA against ANO9 were transfected into HEK293T cells together with the psPAX2 packaging plasmid (Addgene, Cambridge, MA, USA; 12260) and pMD2.G envelope (Addgene 12259); supernatants were collected 24 h after transfection. Epidermal growth factor receptor (EGFR) cDNA was purchased from Sino Biological Inc. (Beijing, China; Clone ID: H10001-NY), and coding regions of EGFR were subcloned into a pCMV vector with a human influenza haemagglutinin (HA) tag. The mammalian-expressible plasmids for ANO5 were purchased from GeneCopoeia (Rockville, MD, USA; Clone ID: EX-T8509-M02). The mammalian expression plasmids for ANO1 and ANO6 have been previously described (Kim et al, 2015; Jun et al, 2016). The coding regions of ANO1, ANO5, and ANO6 were subcloned into the pCMV-Myc-N vector with an N-terminal Myc tag. For immunoblotting, antibodies targeting ANO9 were generated by immunising rabbits with synthetic peptides (RLREKMVHGRQRLGGVGAGSRPPM; near the carboxy terminus of ANO9) using the custom antibody production services of Young In Frontier Inc. (Seoul, Korea). The following antibodies were purchased from commercial sources: antibodies against ANO9 for immunohistochemistry (LifeSpan BioSciences, Seattle, WA, USA), ANO1, ANO6, β-actin, aldolase A (Santa Cruz Biotechnology, Santa Cruz, CA, USA), EGFR, phospho-EGFR, extracellular signal-regulated kinase1/2 (ERK1/2), phospho-ERK1/2, AKT1, phospho-AKT1, HA, Flag, and Myc (Cell Signaling Technology, Beverly, MA, USA).
Chemicals, transfection, and transduction
Doxycycline, hexadimethrine bromide (Polybrene), dimethyl sulphoxide, erlotinib, and gemcitabine were purchased from Sigma-Aldrich (St Louis, MO, USA). CellTiter 96 AQueous One Solution Cell Proliferation assay was purchased from Promega (Madison, WI, USA). Plasmids were transiently transfected into PANC-1 or HEK293T cells using TransIT-X2 reagent (Mirus Bio Corporation, Madison, WI, USA). For lentiviral transduction, PANC-1, AsPC-1, BxPC-3, and Capan-2 cells were treated with Polybrene for 4 h, and then viral supernatants were added to the cell culture medium. To select cells that stably express the transgene, cells were treated with puromycin (5 μg ml−1).
Quantitative PCR analysis
Quantitative PCR (qPCR) was performed using the Applied Biosystems StepOne system (Applied Biosystems, Foster City, CA, USA). According to the comparative threshold cycle (Ct) method (Livak and Schmittgen, 2001), expression of the target gene was normalised to the expression of the housekeeping gene glyceraldehyde 3-phosphate dehydrogenase (GAPDH), yielding the delta Ct (dCt) value. Subtracting the dCt value under control conditions from the dCt value under experimental conditions yielded the delta delta Ct (ddCt) value. Detailed methods are available in the Supplementary Methods.
Immunoblotting and surface biotinylation
Surface biotinylation and immunoblotting were performed as described previously (Lee et al, 2015). For immunoblotting, HEK293T, hTERT-HPNE, PANC-1, AsPC-1, BxPC-3, and Capan-2 cells were grown and treated according to each experimental condition. The cells were washed three times with ice-cold PBS and then harvested with lysis buffer consisting of 50 mM Tris-HCl (pH 7.4), 150 mM NaCl, 10% (v/v) glycerol, 1% (v/v) Triton X-100, 5 mM EDTA, 5 mM EGTA, and complete protease inhibitor cocktail (Roche Applied Science, Mannheim, Germany). The cells were centrifuged, and the cell lysates were diluted in a 5 × sodium dodecyl sulphate (SDS) sample buffer and separated by SDS–PAGE. The separated proteins were transferred to a nitrocellulose membrane that was incubated with the appropriate primary and secondary antibodies. Protein bands were detected by enhanced chemiluminescence (Amersham Biosciences, Piscataway, NJ, USA), and band densities were analysed using Multi Gauge ver. 3.0 software (Fujifilm, Tokyo, Japan).
For biotinylation of plasma membrane proteins, cells were treated with buffer containing Sulfo-NHS-SS-Biotin (Pierce, Rockford, IL, USA). After 30 min, the cells were washed with quenching buffer to remove excess biotin and washed twice with PBS. After harvesting the cells and adding NeutrAvidin solution (UltraLink Immobilized NeutrAvidin Beads 10%; Pierce), the cells were incubated overnight at 4 °C. Avidin-bound complexes were pelleted (16 000 g) and washed three times. The biotinylated proteins were eluted in a 2 × sample buffer, separated by SDS–PAGE, and immunoblotted with primary and secondary antibodies.
Immunoprecipitation and immunocytochemistry
For the immunoprecipitation assay, PANC-1 cells were co-transfected with the Myc- ANO1, Myc-ANO5, Myc-ANO6, Myc-ANO9, and HA-EGFR plasmids. The cell lysates were mixed with anti-Myc antibodies and incubated overnight at 4 °C in lysis buffer. The immune complexes were collected by binding to protein G/A-Sepharose and washed three times with lysis buffer. The immunoprecipitates and lysates were then separated by SDS–PAGE and immunoblotted with anti-HA antibodies.
To determine the cellular localisation of ANO9, immunocytochemistry was performed in PANC-1 cells transfected with the Myc-ANO9 plasmid using anti-Myc antibodies. The localisation of endogenous ANO9 was evaluated in hTERT-HPNE, PANC-1, AsPC-1, BxPC-3, and Capan-2 cells. The cells cultured on coverslips were fixed in 10% formalin for 10 min and permeabilised with 0.1% Triton X-100 for 10 min at room temperature. Nonspecific binding sites were blocked by incubation for 1 h at room temperature with blocking medium consisting of 0.1 ml PBS containing 5% horse serum, 1% bovine serum albumin, and 0.1% gelatin. The cells were then stained with the appropriate primary antibodies and followed by fluorophore-conjugated secondary antibodies. Images were obtained with a Zeiss LSM780 confocal microscope (Carl Zeiss, Berlin, Germany).
Cell viability and colony formation
For cell viability assay, PANC-1 cells were transduced with lentiviral particles for doxycycline-inducible ANO9 overexpression, and AsPC-1, BxPC-3, and Capan-2 cells were transduced with lentiviral particles for doxycycline-inducible shRNA targeting ANO9 (shANO9). Cell proliferation was evaluated using a 3-(4,5-dimethylthiazol-2-yl)-5-(3-carboxymethoxyphenyl)-2-(4-sulfophenyl)-2H-tetrazolium (MTS) assay (CellTiter 96 AQueous One Solution, Promega). To evaluate colony formation, cells were plated in 6-well plates (300 cells per plate PANC-1; 3000 cells per plate AsPC-1, BxPC-3, and Capan-2), and colonies were counted 2 weeks later using crystal violet stain.
Xenograft model
This study was approved by the Committee on Animal Research at Yonsei Medical Center (protocol number 2015-0187), and all animal procedures were performed according to the Yonsei Medical Center Animal Research Guidelines that adhered to the standards articulated in the Animal Research: Reporting of In Vivo Experiments (ARRIVE) (https://www.nc3rs.org.uk/arrive-guidelines). The PANC-1 xenograft tumours were developed in 6-week-old female nude mice by implanting 1 × 107 PANC-1 cells transduced with pTRIPZ-ANO9/TMEM16J subcutaneously on the right flank. Tumour growth was recorded once a week in three dimensions using a digital caliper. Tumour volume (mm3) was calculated as (length × width × height)/2. To induce ANO9 expression in the PANC-1 xenograft, doxycycline was added to the drinking water (1 g doxycycline and 50 g sucrose in 1 litre water). The ANO9 protein expression in the tumour tissues was confirmed by immunostaining.
Patients
This study was approved by the institutional review board of Gangnam Severance Hospital, Yonsei University, Seoul, Korea (IRB No. 3-2015-0157). A total of 78 patients with pancreatic cancer who visited Gangnam Severance Hospital from August 1998 to May 2012 were enrolled in this study. Inclusion criteria were as follows: (1) age >18 years, (2) histologically confirmed PDAC, (3) curative resection of pancreatic cancer, and (4) available electronic medical records including treatment information. Exclusion criteria were: (1) unresectable disease treated with systemic chemotherapy, (2) other histologic types such as cystic neoplasms, neuroendocrine tumours, or lymphomas, and (3) synchronous malignancies. The following baseline data were recorded at the time of diagnosis; age, sex, tumour differentiation (grade), tumour stage (T stage), node stage (N stage), lymphovascular invasion, and operation types.
Tissue immunohistochemistry
Paraffin blocks containing pancreatic cancer tissues were selected by reviewing all haematoxylin and eosin-stained slides of surgical specimens. Tissue cores (1.5 mm diameter) were extracted from each donor block and precisely arrayed in a formalin-fixed and paraffin-embedded tissue microarray (TMA) block. Sections of the TMA block were mounted on poly-L-lysine-coated glass slides, deparaffinised with xylene, and rehydrated. Antigen retrieval was performed by autoclaving the slides in 10 mM citric acid buffer. The slides were incubated for 1 h at room temperature with primary antibodies against ANO9 and then incubated with biotinylated secondary antibodies for 30 min and peroxidase-labelled streptavidin for 30 min. The slides were developed with 3,3′-diaminobenzidine and counterstained with haematoxylin. The staining intensity of ANO9 was scored as negative (0), mild (1+), moderate (2+), or strong (3+) by pathologists who did not have information on the study samples. Positive staining in <5% of tumour cells was considered negative.
Electrophysiology
To measure the Cl− channel activity of ANO9, whole-cell patch clamp recordings were performed in HEK293T cells using ANO1 as a control, as reported previously (Jung et al, 2013; Lee et al, 2015; Jun et al, 2016). Detailed methods are available in the Supplementary Methods.
Statistical analysis
Results of multiple experiments are presented as mean±s.e.m., and P<0.05 was considered significant. Results of cell culture experiments were compared using Student’s t-test or analysis of variance followed by Dunnett’s multiple comparison test, as appropriate, using GraphPad Prism 6 (GraphPad Software, Inc., La Jolla, CA, USA). Patient characteristics according to ANO9 expression level were compared using the χ2 or Fisher’s exact test. Disease-free survival (DFS) was defined as the time from surgery to the first documented recurrence or death from any cause. Overall survival (OS) was calculated from the date of surgery to death from any cause. Time-to-event end point was analysed using the Kaplan–Meier method, and DFS and OS of subgroups were compared using the log-rank test. Univariate and multivariate analyses were performed to identify independent prognostic factors for survival using the Cox proportional hazard regression model. Hazard ratios (HRs) and 95% confidence intervals (CIs) were determined using SPSS Statistics 18.0 software (IBM Corporation, Armonk, NY, USA). We further evaluated the prognostic relevance of ANO9 on OS using a public cancer gene expression database (PROGgene V2, http://www.compbio.iupui.edu/proggene) (Goswami and Nakshatri, 2014). Data sets that included >100 patients were analysed by Cox proportional hazard model. Meta-analyses were performed using Comprehensive Meta-Analysis software (version 3.3.070, Englewood, NJ, USA).
Results
ANO9/TMEM16J is overexpressed in pancreatic cancer cells
To determine whether a specific ANO protein is associated with pancreatic cancer, we evaluated expression levels of ANO/TMEM16 family genes in three PDAC cell lines (AsPC-1, BxPC-3, and PANC-1) and an immortalised normal pancreas cell line (hTERT-HPNE) by qPCR. Of the 10 ANOs examined (ANO1–ANO10), all Ct values except for ANO5 were <37 and the ddCt values of the three cancer cell lines were analysed (Figure 1A). The expression of ANO1 and ANO9 appeared to be upregulated in AsPC-1 and BxPC-3 cells compared with the hTERT-HPNE control. We focussed on the pathogenic role of ANO9 in pancreatic cancer, because the role of ANO1 in cancer pathogenesis has been examined in studies of several other cancers (Duvvuri et al, 2012; Liu et al, 2012; Britschgi et al, 2013; Simon et al, 2013).
The ANO9/TMEM16J is expressed in pancreatic cancer cell lines. (A) Gene expression of ANO/TMEM16 family members in the pancreatic cancer cell lines PANC-1, AsPC-1, and BxPC-3 and the immortalised normal pancreas cell line hTERT-HPNE was determined by qPCR. The −delta delta Ct (−ddCt) values of the three cancer cell lines are depicted. The first delta (dCt) represents the difference in mRNA levels of the target gene and the housekeeping gene GAPDH, and the second delta (ddCt) indicates the difference from dCt of hTERT-HPNE. The mRNA level difference can be estimated using the 2−ddCt method. (B) Expression of ANO9 in PANC-1, AsPC-1, BxPC-3, and Capan-2 cells was re-evaluated by qPCR. (C) Protein levels of ANO9 in hTERT-HPNE, PANC-1, AsPC-1, BxPC-3, and Capan-2 cells were determined by immunoblotting. Results of multiple experiments are summarised in the bottom panel. The specificity of ANO9 antibodies was examined using protein samples from HEK293T cells transfected with pCMV-Myc-ANO9 (upper left panel, see Supplementary Figure 1). (D) Results of the surface biotinylation assay in PANC-1 cells transfected with pCMV-Myc-ANO9 indicates that ANO9 is expressed on the plasma membrane. ANO9 was blotted with anti-Myc antibodies. (E) Immunofluorescence of ANO9 in PANC-1 cells expressing Myc-ANO9 shows that ANO9 is expressed on the plasma membrane (arrowhead). Immunostaining was performed with anti-Myc antibodies. (F) Expression and localisation of endogenous ANO9 were evaluated in HPNE, PANC-1, AsPC-1, BxPC-3, and Capan-2 cells by immunofluorescence with anti-ANO9 antibodies. Ct, threshold cycle. Data are presented as mean±s.e.m. **P<0.01.
To confirm the overexpression of ANO9 in pancreatic cancer cells, we evaluated ANO9 expression in PANC-1, AsPC-1, BxPC-3, and Capan-2 cells by qPCR and immunoblotting. Our results showed a 27–29-fold increase in ANO9 mRNA in AsPC-1, BxPC-3, and Capan-2 cells compared with that of hTERT-HPNE (Figure 1B). The ANO9 mRNA levels were lower in PANC-1 cells than in the other pancreatic cancer cell lines. We then evaluated ANO9 protein levels in hTERT-HPNE, PANC-1, AsPC-1, BxPC-3, and Capan-2 cells by immunoblot analysis (Figure 1C). The specificity of ANO9 antibodies generated in this study was verified using protein samples from HEK293T cells with exogenous ANO1, ANO5, ANO6, and ANO9 expression (Supplementary Figure 1A). We found that ANO9 protein was highly overexpressed in AsPC-1, BxPC-3, and Capan-2 cells (Figure 1C), whereas PANC-1 has little endogenous ANO9. Analysis of ANO9 expression in normal and cancer tissues using a public database (Human Protein Atlas; http://www.proteinatlas.org) (Uhlen et al, 2015) showed that 4 of 12 pancreatic cancer tissues examined expressed ANO9, whereas none of the normal pancreatic tissues expressed ANO9 (Supplementary Table 1). These results support a possible pathogenic role of ANO9 in pancreatic cancer.
The ANO/TMEM16 proteins are plasma membrane proteins having 10 transmembrane domains (Brunner et al, 2014). Therefore, we examined the intracellular localisation of ANO9 in PANC-1 cells transfected with pCMV-Myc-ANO9 using a cell surface biotinylation assay (Figure 1D) and immunocytochemistry (Figure 1E). In addition, we evaluated the localisation of endogenous ANO9 in the pancreatic cell lines (Figure 1F). The results indicated that a significant amount of ANO9 protein is expressed at the cell surface in both exogenously and endogenously ANO9-expressing cells.
ANO9 overexpression induces cell proliferation and ERK/EGFR activation in PANC-1 cells
The PANC-1 cells, which showed a relatively low ANO9 expression (Figure 1C), were chosen for gain-of-function experiments. After transduction with lentiviral particles generated from pTRIPZ-ANO9, a colony of PANC-1 cells that stably expressed doxycycline-inducible ANO9 was selected by puromycin (5 μg ml−1). Colony formation ability according to ANO9 expression level was then evaluated in the ANO9-expressing PANC-1 cells and untransduced PANC-1 cells (controls). Our results showed that colony formation was significantly increased by doxycycline treatment (1 μg ml−1) in the ANO9-expressing PANC-1 cells but not in untransduced PANC-1 cells (Figure 2A). Similar results were obtained in the MTS-based cell viability assay using PANC-1 cells transfected with pCMV-Myc-ANO9 (Supplementary Figure 2).
Exogenous ANO9 promotes proliferation of PANC-1 cells via ERK and EGFR activation. (A) Colony formation assay. After transduction with lentiviral particles containing doxycycline-inducible ANO9 (pTRIPZ-ANO9), PANC-1 cells that stably expressed ANO9 were selected by puromycin (5 μg ml−1). Control (untransduced) or transduced PANC-1 cells were plated on 6-well plates (300 cells per plate), and the number of colonies was determined after 2 weeks. Results of multiple experiments are summarised in the bottom panel. Doxycycline treatment increased colony formation of PANC-1 cells transduced with the ANO9-expressing lentivirus but not that of untransduced cells (controls). (B) Protein levels of ANO9 and cell signalling proteins were determined in PANC-1 cells transduced with lentiviral particles containing doxycycline-inducible ANO9 after treatment with doxycycline for 1 week. Results of multiple experiments are summarised in the right panel. Overexpression of ANO9 increased protein levels of phosphorylated ERK1/2 (pERK1/2), total EGFR, and phosphorylated EGFR (pEGFR). (C) Immunoprecipitation assays were performed with anti-Myc antibodies in PANC-1 cells transfected with plasmids expressing HA-EGFR, Myc-ANO1, Myc-ANO5, Myc-ANO6, and Myc-ANO9. The results of four experiments are summarised in the right panel. EGFR showed the strongest association with ANO9 and only minimal association with ANO6. (D) Gene expression of ANO9 and EGFR was evaluated in PANC-1 cells by qPCR. ANO9 overexpression did not affect mRNA levels of EGFR. Data are presented as mean±s.e.m. NS=not significant; *P<0.05; **P<0.01.
To explore the molecular mechanism underlying ANO9-induced cell proliferation, we evaluated the effects of ANO9 overexpression on AKT/protein kinase B, ERK, and EGFR signalling that are activated in many cancer cells and associated with cell proliferation mediated by other ANO proteins (Bill et al, 2015). We found that ANO9 overexpression in PANC-1 cells did not affect protein levels of AKT1 or phopho-AKT1 but increased phospho-ERK1/2 levels without affecting total ERK1/2 levels (Figure 2B), implying that ANO9-mediated cell proliferation is associated with activation of the mitogen-activated protein kinase (MAPK) pathway. In addition, both total EGFR and phospho-EGFR levels were increased in PANC-1 cells overexpressing ANO9 (Figure 2B). The ANO1 is thought to directly associate with and activate EGFR that promotes tumourigenesis of head and neck cancers (Bill et al, 2015). Therefore, we examined whether ANO9 physically associated with EGFR in an immunoprecipitation assay and compared its EGFR-binding ability with that of other ANOs. Our results showed that both ANO1 and ANO9 form protein complexes with EGFR in PANC-1 cells, but the binding of ANO9 to EGFR was 132% higher than that of ANO1 (Figure 2C). Results of qPCR revealed that ANO9 overexpression did not affect mRNA levels of EGFR (Figure 2D). Collectively, these results indicate that protein complex formation between ANO9 and EGFR, rather than transcriptional regulation, is involved in ANO9-mediated upregulation and activation of EGFR.
Knockdown of ANO9 inhibits pancreatic cancer cell proliferation
Next, we performed loss-of-function experiments in the ANO9-overexpressing AsPC-1, BxPC-3, and Capan-2 pancreatic cancer cells (Figure 3). After transduction with lentiviral particles generated from pTRIPZ-ANO9-shRNA, a colony of each cell line that stably expressed doxycycline-inducible ANO9 shRNA was selected by puromycin (5 μg ml−1). Knockdown of ANO9 by doxycycline treatment (1 μg ml−1) was confirmed by immunoblotting (Figure 3B and Supplementary Figure 1B). Colony formation was strongly reduced by doxycycline in cells expressing shRNA against ANO9 but not in untransduced control cells (Figure 3A, C, and D), indicating that ANO9 depletion inhibits cell proliferation in pancreatic cancer cells.
The ANO9 knockdown inhibits proliferation of AsPC-1, BxPC-3, and Capan-2 cells. (A) Colony formation assay in AsPC-1 cells. After transduction with lentiviral particles containing doxycycline-inducible shRNA against ANO9 (shANO9), AsPC-1 cells that stably expressed shANO9 were selected by puromycin (5 μg ml−1). Untransduced AsPC-1 cells (controls) or transduced AsPC-1 cells were plated on 6-well plates (3000 cells per plate), and the number of colonies was counted after 2 weeks. Results of multiple experiments are summarised in the bottom panel. Doxycycline (1 μg ml−1) reduced colony formation in AsPC-1 cells transduced with the shANO9-expressing lentivirus but not in untransduced cells. (B) Protein levels of ANO9 and cell signalling proteins were determined in AsPC-1 cells transduced with lentiviral particles containing doxycycline-inducible shANO9 by immunoblotting after treatment with doxycycline for 1 week. Doxycycline reduced ANO9 expression in AsPC-1 cells by 82±3%. Results of multiple experiments are summarised in the right panel. Cells expressing shANO9 showed lower levels of phosphorylated ERK1/2 (pERK1/2), total EGFR, and phosphorylated EGFR (pEGFR). (C and D) Colony formation of BxPC-3 (C) and Capan-2 (D) cells transduced with lentiviral particles containing doxycycline-inducible shANO9 was assessed as described in (A). Knockdown of ANO9 inhibited colony formation in BxPC-3 and Capan-2 cells. Data are presented as mean±s.e.m. NS=not significant; *P<0.05; **P<0.01.
To investigate the effects of ANO9 knockdown, the expression and activation of AKT, ERK1/2, and EGFR were assessed in AsPC-1 cells (Figure 3B). The results were exactly opposite to that of ANO9 overexpression in PANC-1 cells (Figure 2B), showing that ANO9 depletion inhibited phosphorylation of ERK1/2 without affecting protein level and decreased both total protein level and phosphorylation of EGFR (Figure 3B). These results support the roles of ERK and EGFR in ANO9-induced tumourigenesis.
ANO9 promotes tumour growth in mice
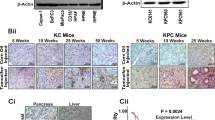
We then examined the effects of ANO9 on pancreatic tumour growth using nude mice implanted with PANC-1 cells that stably express doxycycline-inducible ANO9. Our results showed that tumour growth was strongly increased in mice treated with doxycycline (Figure 4A and B). Immunohistochemistry confirmed ANO9 overexpression in the xenograft tumours of mice treated with doxycycline (Figure 4C and D).
The ANO9 promotes tumour growth in mice. (A and B) The PANC-1 xenograft tumours were developed in 6-week-old female nude mice by implanting 1 × 107 PANC-1 cells expressing doxycycline-induced ANO9 subcutaneously on the right flank. Tumour growth of doxycycline-treated mice and untreated mice was recorded once a week. Representative examples are shown in (A), and results of multiple experiments are depicted in (B). ANO9 overexpression markedly increased tumour growth in the PANC-1 xenograft model. (C and D) Immunostaining of ANO9 in the xenograft tumour tissues confirmed doxycycline-induced ANO9 overexpression. Data are presented as mean±s.e.m. *P<0.05.
ANO9 is associated with poor survival in patients with pancreatic cancer
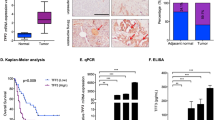
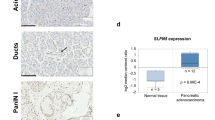
To investigate the effects of ANO9 on pancreatic cancer in humans, we evaluated survival in 78 patients with pancreatic cancer according to ANO9 expression level. The ANO9 protein level in tumour tissues was compared with that of normal pancreatic tissues (Figure 5A–C). Low ANO9 expression (0 or 1+) was observed in 55 (70.5%) patients, and high ANO9 expression (2+ or 3+) was observed in 23 (29.5%) patients. Baseline characteristics of patients according to ANO9 expression are shown in Supplementary Table 2. Median patient age was 64 years (range, 37–83), and 39 patients (50%) were men. The two groups did not differ significantly according to any of the demographic parameters.
Survival analysis of patients with pancreatic cancer according to ANO9 expression level. (A–C) Representative ANO9 immunohistochemistry results of normal pancreas tissue (A), tumour with low ANO9 expression (B, +0), and tumour with high ANO9 expression (C, +3). (D and E) Kaplan–Meier survival curves for disease-free survival (DFS, D) and overall survival (OS, E) according to ANO9 expression level in patients with pancreatic cancer. (F) Meta-analysis of OS using data from three independent cohorts of patients with pancreatic cancer (GSE21501, GSE71729, and TCGA-PAAD) in PROGgene V2 (http://www.compbio.iupui.edu/proggene). Data sets of >100 subjects containing ANO9 expression data were analysed. Increased expression of ANO9 is significantly associated with poor survival (P=0.012).
Kaplan–Meier plots for DFS and OS are presented in Figure 5D and E. Univariate analysis was carried out to determine whether demographic parameters or ANO9 expression level affect DFS and OS. Factors that were significant at P<0.3 were tested using multivariate analysis to identify independent factors associated with DFS and OS. Results of univariate analysis showed that DFS was lower in patients with high ANO9 expression levels in tumour tissues than in patients with low ANO9 expression (median survival 13.3 vs 7.5 months, P=0.04). This finding was confirmed by multivariate analysis (P=0.03, Supplementary Table 3 and Figure 5D). Similarly, patients with high ANO9 expression levels showed lower OS in multivariate analysis (median survival time 30.4 vs 20.2 months, P=0.01; Supplementary Table 3 and Figure 5E).
We also investigated the effect of ANO9 on survival using the PROGgene V2 prognostics database (Goswami and Nakshatri, 2014). Data sets of >100 subjects with pancreatic cancer and containing ANO9 expression data were included in the survival analysis. A meta-analysis of OS using data from three independent cohorts (GSE21501, GSE71729, and TCGA-PAAD) indicated that increased expression of ANO9 is significantly associated with poor survival in pancreatic cancer (overall P=0.012; Figure 5F).
ANO9 knockdown augments the anticancer effects of gemcitabine and erlotinib
To explore the possibility of ANO9 inhibition for clinical application, we examined the combinatorial effects of ANO9 knockdown and treatment with anticancer agents. The effects of gemcitabine and erlotinib in cells with or without doxycycline-inducible ANO9 shRNA were assessed by the colony formation assay or MTS-based cell viability assay. In control cells lacking ANO9 shRNA, treatment with doxycycline alone did not affect cell proliferation (Figure 3). However, doxycycline reduced colony formation and cell viability in AsPC-1 cells transduced with doxycycline-inducible ANO9 shRNA (Figure 6). Gemcitabine (Figure 6A–C) and erlotinib (Figure 6D and E) further reduced colony formation and cell viability in ANO9-depleted cells. ANO9 knockdown appeared to lower the IC50 value of erlotinib from 4.51 to 1.50 μ M in the colony formation assay (P=0.006, Figure 6E). The ANO9 knockdown showed similar effects on cell proliferation in BxPC-3 cells (Supplementary Figure 3) and Capan-2 (Supplementary Figure 4) cells. Collectively, these results support the potential of ANO9 as a target for drug development in pancreatic cancers, in particular those with high ANO9 expression.
Knockdown of ANO9 augments the anticancer effects of gemcitabine and erlotinib in AsPC-1 cells. (A–C) The antiproliferative effects of the cytotoxic agent gemcitabine were examined in AsPC-1 cells expressing doxycycline-inducible shRNA against ANO9 (shANO9) using the colony formation assay (A and B) and the MTS-based cell viability assay (C). (D–F) The anticancer effects of the EGFR inhibitor erlotinib were examined in AsPC-1 cells expressing doxycycline-inducible shANO9 using the colony formation assay (C and D) and the MTS-based cell viability assay (F). ANO9 knockdown additively inhibited cell proliferation when combined with gemcitabine or erlotinib. Data are presented as mean±s.e.m.
Discussion
In the present study, we demonstrate that ANO9 may serve as a marker for poor prognosis in pancreatic cancer and a potential target for drug development via an integrated in vitro, in vivo, and clinical investigation. Our results showed that ANO9 is overexpressed in most pancreatic cancer cell lines tested and in ∼30% of tissue samples obtained from patients with pancreatic cancer. Moreover, high ANO9 expression was associated with increased proliferation of pancreatic cancer cells and poor survival in patients with pancreatic cancer.
Experiments in PDAC cell lines indicate that ANO9 promotes proliferation in pancreatic cancer cells. Gain-of-function experiments in PANC-1 cells showed that exogenous ANO9 increased colony formation (Figure 2) and in vivo tumour growth (Figure 4), and loss-of-function experiments in AsPC-1, BxPC-3, and Capan-2 cells showed that ANO9 depletion inhibited cell proliferation (Figure 3). Experiments investigating the underlying mechanism of these ANO9 effects showed that activation of MAPK and EGFR was associated with ANO9-mediated cell proliferation (Figures 2B and 3B).
Activating KRAS mutations are found in >90% of pancreatic cancers (Kanda et al, 2012). Although KRAS mutations are strongly associated with the initiation of pancreatic adenocarcinoma, additional factors such as EGFR signalling are necessary for KRAS-driven pancreatic tumourigenesis (Ardito et al, 2012; Navas et al, 2012). Of the ANO/TMEM16 proteins, ANO1 is the most well studied in terms of its functional and clinical roles. Overexpression of ANO1 is observed in several cancers and is associated with lower survival in patients with these cancers (Duvvuri et al, 2012; Ruiz et al, 2012; Britschgi et al, 2013; Rodrigo et al, 2015). Results of previous studies also provide evidence for the involvement of EGFR activation in ANO1-related cancer cell growth and proliferation (Duvvuri et al, 2012; Britschgi et al, 2013; Bill et al, 2015). In the present study, results of the immunoprecipitation assay suggested that physical association with EGFR is responsible for ANO9-induced activation of EGFR (Figure 2C). Interestingly, the viability of PANC-1 cells was increased by exogenous expression of ANO1 and ANO9 that associate with EGFR, but not by ANO6 that does not associate with EFGR (Figure 2C), implying that not all ANOs induce cell proliferation via EGFR (Supplementary Figure 2).
Notably, the binding ability of ANO9 to EGFR was almost double that of ANO1 (Figure 2C), indicating that ANO9 has a higher potential to induce EGFR-mediated cell proliferation. How ANO9 can upregulate and activate EGFR remains a question. Most ANO proteins are located at the cell surface and some (e.g., ANO1, ANO2, ANO6) have CaCC activity (Kim et al, 2015; Pedemonte and Galietta, 2014). Although ANO9 is expressed at the plasma membrane (Figure 1E–F), it did not show CaCC activity (Supplementary Figure 5), indicating that ANO9-mediated EGFR activation is not associated with CaCC activity.
To better understand the clinical implications of ANO9 expression, we analysed the survival rate of 78 patients who underwent curative resection for primary pancreatic cancer; 70 of these patients (90%) had TNM stage 2 disease. Immunohistochemistry results showed that ANO9 expression levels were high (2+ or 3+) in 29.5% of the pancreatic cancer tissue samples. Patient characteristics and tumour grade did not differ significantly according to the ANO9 expression level (Supplementary Table 2). However, in univariate and multivariate survival analyses, DFS and OS were significantly shorter in patients with high ANO9 expression, indicating that ANO9 is a poor prognostic factor for pancreatic cancer (average hazard ratio, 1.74–2.64; Figure 5 and Supplementary Table 3). These results were supported by a meta-analysis of pancreatic cancer data sets (Figure 5F). In contrast, a previous study suggested that decreased expression of ANO9 was correlated with poor prognostic outcomes in colorectal cancer (Li et al, 2015). Therefore, we also analysed the relationship between ANO9 expression and survival of patients with colorectal cancer in public data sets. Interestingly, the results indicated that ANO9 was not significantly correlated with OS in colorectal cancer (Supplementary Figure 6). The reasons for the differential effects of ANO9 between pancreatic cancer and colorectal cancer are unclear. However, ANO9 is not normally expressed in the pancreas, whereas it is expressed in normal colorectal tissues (Supplementary Table 1), suggesting that aberrant expression of ANO9 in pancreatic cancer tissues may cause detrimental outcomes, whereas the native expression of ANO9 in colorectal cancers does not have negative effects.
Gemcitabine and erlotinib are used clinically to treat pancreatic cancer (Ryan et al, 2014), and these drugs effectively inhibited pancreatic cancer cell proliferation. We found that ANO9 depletion provided an additive effect in inhibiting proliferation of ANO9-positive cancer cells (Figure 6 and Supplementary Figures 3 and 4). Notably, AsPC-1 and Capan-2 cells, which carry KRAS mutations (Navas et al, 2012), were sensitive to ANO9 inhibition, suggesting that ANO9 may be an effective target in patients with ANO9-positive pancreatic cancers regardless of KRAS mutation status. Gemcitabine has been shown to promote pancreatic cancer stemness that is closely associated with drug resistance and tumour relapse (Zhang et al, 2016). Combination therapy with an ANO9 inhibitor may help prevent drug resistance and tumour relapse associated with the prolonged use of gemcitabine. Because the EGFR tyrosine kinase inhibitor erlotinib is widely used to treat pancreatic cancer (Ryan et al, 2014), ANO9-induced EGFR upregulation may have clinical significance. Overexpression of EGFR is a predictive biomarker for response to anti-EGFR therapy in some cancers (Lievre et al, 2006; Pirker et al, 2012), suggesting that ANO9 may serve as a useful biomarker for response to EGFR inhibition therapy in pancreatic cancer.
In conclusion, we propose ANO9 as a prognostic marker for poor survival in pancreatic cancer and a potential biomarker for anti-EGFR therapy. Future clinical trials evaluating the prognostic and biomarker value of ANO9 are needed to confirm our conclusions. In addition, our results suggest that ANO9 inhibitors that block ANO9-mediated EGFR upregulation may enhance the therapeutic outcome of chemotherapy with gemcitabine or erlotinib.
Change history
05 December 2017
This paper was modified 12 months after initial publication to switch to Creative Commons licence terms, as noted at publication
References
Ardito CM, Gruner BM, Takeuchi KK, Lubeseder-Martellato C, Teichmann N, Mazur PK, Delgiorno KE, Carpenter ES, Halbrook CJ, Hall JC, Pal D, Briel T, Herner A, Trajkovic-Arsic M, Sipos B, Liou GY, Storz P, Murray NR, Threadgill DW, Sibilia M, Washington MK, Wilson CL, Schmid RM, Raines EW, Crawford HC, Siveke JT (2012) EGF receptor is required for KRAS-induced pancreatic tumorigenesis. Cancer Cell 22 (3): 304–317.
Bera TK, Das S, Maeda H, Beers R, Wolfgang CD, Kumar V, Hahn Y, Lee B, Pastan I (2004) NGEP, a gene encoding a membrane protein detected only in prostate cancer and normal prostate. Proc Natl Acad Sci USA 101 (9): 3059–3064.
Biankin AV, Waddell N, Kassahn KS, Gingras MC, Muthuswamy LB, Johns AL, Miller DK, Wilson PJ, Patch AM, Wu J, Chang DK, Cowley MJ, Gardiner BB, Song S, Harliwong I, Idrisoglu S, Nourse C, Nourbakhsh E, Manning S, Wani S, Gongora M, Pajic M, Scarlett CJ, Gill AJ, Pinho AV, Rooman I, Anderson M, Holmes O, Leonard C, Taylor D, Wood S, Xu Q, Nones K, Fink JL, Christ A, Bruxner T, Cloonan N, Kolle G, Newell F, Pinese M, Mead RS, Humphris JL, Kaplan W, Jones MD, Colvin EK, Nagrial AM, Humphrey ES, Chou A, Chin VT, Chantrill LA, Mawson A, Samra JS, Kench JG, Lovell JA, Daly RJ, Merrett ND, Toon C, Epari K, Nguyen NQ, Barbour A, Zeps N, Kakkar N, Zhao F, Wu YQ, Wang M, Muzny DM, Fisher WE, Brunicardi FC, Hodges SE, Reid JG, Drummond J, Chang K, Han Y, Lewis LR, Dinh H, Buhay CJ, Beck T, Timms L, Sam M, Begley K, Brown A, Pai D, Panchal A, Buchner N, De Borja R, Denroche RE, Yung CK, Serra S, Onetto N, Mukhopadhyay D, Tsao MS, Shaw PA, Petersen GM, Gallinger S, Hruban RH, Maitra A, Iacobuzio-Donahue CA, Schulick RD, Wolfgang CL, Morgan RA, Lawlor RT, Capelli P, Corbo V, Scardoni M, Tortora G, Tempero MA, Mann KM, Jenkins NA, Perez-Mancera PA, Adams DJ, Largaespada DA, Wessels LF, Rust AG, Stein LD, Tuveson DA, Copeland NG, Musgrove EA, Scarpa A, Eshleman JR, Hudson TJ, Sutherland RL, Wheeler DA, Pearson JV, McPherson JD, Gibbs RA, Grimmond SM (2012) Pancreatic cancer genomes reveal aberrations in axon guidance pathway genes. Nature 491 (7424): 399–405.
Bill A, Gutierrez A, Kulkarni S, Kemp C, Bonenfant D, Voshol H, Duvvuri U, Gaither LA (2015) ANO1/TMEM16A interacts with EGFR and correlates with sensitivity to EGFR-targeting therapy in head and neck cancer. Oncotarget 6 (11): 9173–9188.
Bolduc V, Marlow G, Boycott KM, Saleki K, Inoue H, Kroon J, Itakura M, Robitaille Y, Parent L, Baas F, Mizuta K, Kamata N, Richard I, Linssen WH, Mahjneh I, de Visser M, Bashir R, Brais B (2010) Recessive mutations in the putative calcium-activated chloride channel Anoctamin 5 cause proximal LGMD2L and distal MMD3 muscular dystrophies. Am J Hum Genet 86 (2): 213–221.
Britschgi A, Bill A, Brinkhaus H, Rothwell C, Clay I, Duss S, Rebhan M, Raman P, Guy CT, Wetzel K, George E, Popa MO, Lilley S, Choudhury H, Gosling M, Wang L, Fitzgerald S, Borawski J, Baffoe J, Labow M, Gaither LA, Bentires-Alj M (2013) Calcium-activated chloride channel ANO1 promotes breast cancer progression by activating EGFR and CAMK signaling. Proc Natl Acad Sci USA 110 (11): E1026–1034.
Brunner JD, Lim NK, Schenck S, Duerst A, Dutzler R (2014) X-ray structure of a calcium-activated TMEM16 lipid scramblase. Nature 516 (7530): 207–212.
Caputo A, Caci E, Ferrera L, Pedemonte N, Barsanti C, Sondo E, Pfeffer U, Ravazzolo R, Zegarra-Moran O, Galietta LJ (2008) TMEM16A, a membrane protein associated with calcium-dependent chloride channel activity. Science 322 (5901): 590–594.
Dutertre M, Lacroix-Triki M, Driouch K, de la Grange P, Gratadou L, Beck S, Millevoi S, Tazi J, Lidereau R, Vagner S, Auboeuf D (2010) Exon-based clustering of murine breast tumor transcriptomes reveals alternative exons whose expression is associated with metastasis. Cancer Res 70 (3): 896–905.
Duvvuri U, Shiwarski DJ, Xiao D, Bertrand C, Huang X, Edinger RS, Rock JR, Harfe BD, Henson BJ, Kunzelmann K, Schreiber R, Seethala RS, Egloff AM, Chen X, Lui VW, Grandis JR, Gollin SM (2012) TMEM16A induces MAPK and contributes directly to tumorigenesis and cancer progression. Cancer Res 72 (13): 3270–3281.
Goswami CP, Nakshatri H (2014) PROGgeneV2: enhancements on the existing database. BMC Cancer 14: 970.
Huang F, Wang X, Ostertag EM, Nuwal T, Huang B, Jan YN, Basbaum AI, Jan LY (2013) TMEM16C facilitates Na(+)-activated K+ currents in rat sensory neurons and regulates pain processing. Nat Neurosci 16 (9): 1284–1290.
Jun I, Cheng MH, Sim E, Jung J, Suh BL, Kim Y, Son H, Park K, Kim CH, Yoon JH, Whitcomb DC, Bahar I, Lee MG (2016) Pore dilatation increases the bicarbonate permeability of CFTR, ANO1 and glycine receptor anion channels. J Physiol 594 (11): 2929–2955.
Jung J, Nam JH, Park HW, Oh U, Yoon J, Lee MG (2013) Dynamic modulation of ANO1/TMEM16A HCO3(-) permeability by Ca2+/calmodulin. Proc Natl Acad Sci USA 110 (1): 360–365.
Kanda M, Matthaei H, Wu J, Hong SM, Yu J, Borges M, Hruban RH, Maitra A, Kinzler K, Vogelstein B, Goggins M (2012) Presence of somatic mutations in most early-stage pancreatic intraepithelial neoplasia. Gastroenterology 142 (4): 730–733 e739.
Katoh M, Katoh M (2004) Identification and characterization of human TP53I5 and mouse Tp53i5 genes in silico. Int J Oncol 25 (1): 225–230.
Kim HJ, Jun I, Yoon JS, Jung J, Kim YK, Kim WK, Kim BJ, Song J, Kim SJ, Nam JH, Lee MG (2015) Selective serotonin reuptake inhibitors facilitate ANO6 (TMEM16F) current activation and phosphatidylserine exposure. Pflugers Arch 467 (11): 2243–2256.
Lee CS, Jun I, Choi S, Lee JH, Lee MG, Lee SC, Kim EK (2015) A novel BEST1 mutation in autosomal recessive bestrophinopathy. Invest Ophthalmol Vis Sci 56 (13): 8141–8150.
Li C, Cai S, Wang X, Jiang Z (2015) Identification and characterization of ANO9 in stage II and III colorectal carcinoma. Oncotarget 6 (30): 29324–29334.
Lievre A, Bachet JB, Le Corre D, Boige V, Landi B, Emile JF, Cote JF, Tomasic G, Penna C, Ducreux M, Rougier P, Penault-Llorca F, Laurent-Puig P (2006) KRAS mutation status is predictive of response to cetuximab therapy in colorectal cancer. Cancer Res 66 (8): 3992–3995.
Liu W, Lu M, Liu B, Huang Y, Wang K (2012) Inhibition of Ca(2+)-activated Cl(-) channel ANO1/TMEM16A expression suppresses tumor growth and invasiveness in human prostate carcinoma. Cancer Lett 326 (1): 41–51.
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods 25 (4): 402–408.
Navas C, Hernandez-Porras I, Schuhmacher AJ, Sibilia M, Guerra C, Barbacid M (2012) EGF receptor signaling is essential for k-ras oncogene-driven pancreatic ductal adenocarcinoma. Cancer Cell 22 (3): 318–330.
Pedemonte N, Galietta LJ (2014) Structure and function of TMEM16 proteins (anoctamins). Physiol Rev 94 (2): 419–459.
Picollo A, Malvezzi M, Accardi A (2015) TMEM16 proteins: unknown structure and confusing functions. J Mol Biol 427 (1): 94–105.
Pirker R, Pereira JR, von Pawel J, Krzakowski M, Ramlau R, Park K, de Marinis F, Eberhardt WE, Paz-Ares L, Storkel S, Schumacher KM, von Heydebreck A, Celik I, O'Byrne KJ (2012) EGFR expression as a predictor of survival for first-line chemotherapy plus cetuximab in patients with advanced non-small-cell lung cancer: analysis of data from the phase 3 FLEX study. Lancet Oncol 13 (1): 33–42.
Rodrigo JP, Menendez ST, Hermida-Prado F, Alvarez-Teijeiro S, Villaronga MA, Alonso-Duran L, Vallina A, Martinez-Camblor P, Astudillo A, Suarez C, Maria Garcia-Pedrero J (2015) Clinical significance of Anoctamin-1 gene at 11q13 in the development and progression of head and neck squamous cell carcinomas. Sci Rep 5: 15698.
Ruiz C, Martins JR, Rudin F, Schneider S, Dietsche T, Fischer CA, Tornillo L, Terracciano LM, Schreiber R, Bubendorf L, Kunzelmann K (2012) Enhanced expression of ANO1 in head and neck squamous cell carcinoma causes cell migration and correlates with poor prognosis. PLoS One 7 (8): e43265.
Ryan DP, Hong TS, Bardeesy N (2014) Pancreatic adenocarcinoma. N Engl J Med 371 (22): 2140–2141.
Sauter DR, Novak I, Pedersen SF, Larsen EH, Hoffmann EK (2015) ANO1 (TMEM16A) in pancreatic ductal adenocarcinoma (PDAC). Pflugers Arch 467 (7): 1495–1508.
Schroeder BC, Cheng T, Jan YN, Jan LY (2008) Expression cloning of TMEM16A as a calcium-activated chloride channel subunit. Cell 134 (6): 1019–1029.
Siegel RL, Miller KD, Jemal A (2015) Cancer statistics, 2015. CA Cancer J Clin 65 (1): 5–29.
Simon S, Grabellus F, Ferrera L, Galietta L, Schwindenhammer B, Muhlenberg T, Taeger G, Eilers G, Treckmann J, Breitenbuecher F, Schuler M, Taguchi T, Fletcher JA, Bauer S (2013) DOG1 regulates growth and IGFBP5 in gastrointestinal stromal tumors. Cancer Res 73 (12): 3661–3670.
Stocken DD, Hassan AB, Altman DG, Billingham LJ, Bramhall SR, Johnson PJ, Freemantle N (2008) Modelling prognostic factors in advanced pancreatic cancer. Br J Cancer 99 (6): 883–893.
Uhlen M, Fagerberg L, Hallstrom BM, Lindskog C, Oksvold P, Mardinoglu A, Sivertsson A, Kampf C, Sjostedt E, Asplund A, Olsson I, Edlund K, Lundberg E, Navani S, Szigyarto CA, Odeberg J, Djureinovic D, Takanen JO, Hober S, Alm T, Edqvist PH, Berling H, Tegel H, Mulder J, Rockberg J, Nilsson P, Schwenk JM, Hamsten M, von Feilitzen K, Forsberg M, Persson L, Johansson F, Zwahlen M, von Heijne G, Nielsen J, Ponten F (2015) Proteomics. Tissue-based map of the human proteome. Science 347 (6220): 1260419.
Vermeer S, Hoischen A, Meijer RP, Gilissen C, Neveling K, Wieskamp N, de Brouwer A, Koenig M, Anheim M, Assoum M, Drouot N, Todorovic S, Milic-Rasic V, Lochmuller H, Stevanin G, Goizet C, David A, Durr A, Brice A, Kremer B, van de Warrenburg BP, Schijvenaars MM, Heister A, Kwint M, Arts P, van der Wijst J, Veltman J, Kamsteeg EJ, Scheffer H, Knoers N (2010) Targeted next-generation sequencing of a 12.5 Mb homozygous region reveals ANO10 mutations in patients with autosomal-recessive cerebellar ataxia. Am J Hum Genet 87 (6): 813–819.
Yang YD, Cho H, Koo JY, Tak MH, Cho Y, Shim WS, Park SP, Lee J, Lee B, Kim BM, Raouf R, Shin YK, Oh U (2008) TMEM16A confers receptor-activated calcium-dependent chloride conductance. Nature 455 (7217): 1210–1215.
Zhang Z, Duan Q, Zhao H, Liu T, Wu H, Shen Q, Wang C, Yin T (2016) Gemcitabine treatment promotes pancreatic cancer stemness through the Nox/ROS/NF-kappaB/STAT3 signaling cascade. Cancer Lett 382 (1): 53–63.
Zorde Khvalevsky E, Gabai R, Rachmut IH, Horwitz E, Brunschwig Z, Orbach A, Shemi A, Golan T, Domb AJ, Yavin E, Giladi H, Rivkin L, Simerzin A, Eliakim R, Khalaileh A, Hubert A, Lahav M, Kopelman Y, Goldin E, Dancour A, Hants Y, Arbel-Alon S, Abramovitch R, Galun E (2013) Mutant KRAS is a druggable target for pancreatic cancer. Proc Natl Acad Sci USA 110 (51): 20723–20728.
Acknowledgements
This study was supported by a grant from the National Research Foundation, the Ministry of Science, ICT and Future Planning, Republic of Korea (2013R1A3A2042197).
Author contributions
IJ and HSP: acquisition and analysis of data and drafting of the manuscript; HP, JWH, MJA, BGY, XZ, YHC, and YKS: acquisition and interpretation of experimental data; JIY, JJ, and HYG: study concept and design; JSP and DSY: acquisition and interpretation of clinical data; H-CJ and MGL: study concept and design, critical revision of the manuscript, and study supervision.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
This work is published under the standard license to publish agreement. After 12 months the work will become freely available and the license terms will switch to a Creative Commons Attribution-NonCommercial-Share Alike 4.0 Unported License.
Supplementary Information accompanies this paper on British Journal of Cancer website
Rights and permissions
From twelve months after its original publication, this work is licensed under the Creative Commons Attribution-NonCommercial-Share Alike 4.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/4.0/
About this article
Cite this article
Jun, I., Park, H., Piao, H. et al. ANO9/TMEM16J promotes tumourigenesis via EGFR and is a novel therapeutic target for pancreatic cancer. Br J Cancer 117, 1798–1809 (2017). https://doi.org/10.1038/bjc.2017.355
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/bjc.2017.355
Keywords
This article is cited by
-
Regulation of phospholipid distribution in the lipid bilayer by flippases and scramblases
Nature Reviews Molecular Cell Biology (2023)
-
Interleukin-4-induced FABP4 promotes lipogenesis in human skeletal muscle cells by activating the PPAR γ signaling pathway
Cell Biochemistry and Biophysics (2022)
-
MethylSPWNet and MethylCapsNet: Biologically Motivated Organization of DNAm Neural Networks, Inspired by Capsule Networks
npj Systems Biology and Applications (2021)
-
Comparison of ophthalmic toxicity of light-emitting diode and organic light-emitting diode light sources
Scientific Reports (2020)
-
Predicting Functional Modules of Liver Cancer Based on Differential Network Analysis
Interdisciplinary Sciences: Computational Life Sciences (2019)