Abstract
Background:
Oesophageal squamous cell carcinoma (SCC) causes the highest number of cancer deaths in some regions of Northern China. Previously, we narrowed down a critical region at 9q33-34, identified to be associated with tumour-suppressive function of deleted in oesophageal cancer 1 (DEC1) in oesophageal SCC.
Methods:
We generated DEC1 antibody and constructed tissue microarrays (TMAs) utilising tissue specimens from Henan, a high-risk region for oesophageal SCC, to investigate the importance of DEC1 expression in this cancer.
Results:
Tissue microarray immunohistochemical staining reveals significant loss of DEC1 from hyperplasia, to tumour, and to lymph node metastasis. In addition, the loss of DEC1 in tumour is age-dependent. Interestingly, there is significant abrogation of DEC1 expression in patients with a family history of oesophageal SCC. Deleted in oesophageal cancer 1 localises to both the cytoplasm and nucleus. The vesicular pattern of DEC1 in the cytoplasm appears to localise at the Golgi and Golgi–endoplasmic reticulum intermediate compartment.
Conclusion:
This is the first TMA study to suggest a clinical association of DEC1 in lymph node metastatic oesophageal SCC, early onset oesophageal SCC and familial oesophageal SCC development. Subcellular localisation of DEC1 and its expression in oesophageal SCC tissues provide important insight for further deciphering the molecular mechanism of DEC1 in oesophageal SCC development.
Similar content being viewed by others
Main
Oesophageal carcinoma occurs with an especially high frequency in certain regions of Northern China, including the high-risk region of Henan (Li, 1982). Allelic losses on human chromosomes hallmark the presence of tumour-suppressor genes involved in cancer development. Frequent loss of heterozygosity (LOH) on chromosome 9 is reported in squamous cell carcinoma (SCC) of the lung and of the head and neck (Tsuchiya et al, 1992). In oesophageal SCC, this chromosome 9 loss is also commonly observed (Ko et al, 2001). Previously, we identified a critical region at 9q33-34 in oesophageal SCC and identified the importance of deleted in oesophageal cancer 1 (DEC1), which suppresses anchorage-independent growth in vitro and tumourigenesis in vivo (Nishiwaki et al, 2000; Yang et al, 2005). Reverse transcriptase–PCR analysis shows its frequent downregulation in an oesophageal SCC cell line panel and primary tumours, indicating that DEC1 may serve as an early detection biomarker for oesophageal cancer patients (Leung et al, 2008). Despite the tumour-suppressive role of DEC1, there are as yet no studies reported regarding either the role of the protein or its clinical relevance in oesophageal SCC development. A polyclonal antibody was generated against DEC1 and used to analyse a tissue microarray (TMA) constructed with tissues from the oesophageal SCC high-risk region of Henan, China. Immunohistochemical (IHC) staining was used to study the association of DEC1 expression according to status of cancer progression, age, and association with familial oesophageal cancer. Finally, to better understand the molecular mechanism for DEC1 function, we examined its subcellular protein cytolocalisation.
Materials and methods
Cell lines, culture conditions, and clinical specimens
The pcDNA 3.1(+)/DEC1 transfectants and pcDNA3.1(+) vector-alone control were cultured in growth medium containing 400 μg ml–1 G418 (CalBiochem, San Diego, CA, USA), as previously described (Leung et al, 2008). The SLMT-1 cell line was established from a Hong Kong Chinese oesophageal SCC patient (Tang et al, 2001). An immortalised normal oesophageal epithelial cell line (NE1) was cultured in keratinocyte serum-free medium (Invitrogen, Carlsbad, CA, USA), as previously described (Zhang et al, 2006). Oesophageal SCC patient tissues were collected from the Yaocun Esophageal Cancer Hospital, Linzhou, Henan, China from 2005 to 2007.
Antibody generation and purification
Bacterial His-tag and GST-tag protein purification systems were used to purify the DEC1 protein. The DEC1 210 bp cDNA fragment was subcloned from pCR3.1-DEC1 into pGEX-4T-1 (Amersham Pharmacia Biotech, Uppsala, Sweden) at EcoRI and SalI restriction sites and into pET28a (Novagen, Madison, WI, USA) at EcoRI and XhoI sites. Ni-NTA agarose (Qiagen, Hilden, Germany) was used to purify His-tagged DEC1 protein according to the manufacturer's protocol. Purified proteins were dialysed and then emulsified in adjuvant (Sigma-Aldrich, St Louis, MO, USA). Subsequently, it was used to immunise New Zealand 2-month-old white rabbits by subcutaneous methods. Glutathione agarose (GE Healthcare, London, Canada) was used to purify GST-DEC1 protein according to the manufacturer's protocol. The purified GST-tagged DEC1 protein was transferred to a nitrocellulose membrane by SDS–PAGE. The membrane region containing the GST-tagged DEC1-sized protein was cut out and incubated with 10 ml of the rabbit serum at 4 °C overnight. Then 0.1 M glycine buffer (pH 2.0) was added to elute the antibody. One-tenth volume of Tris-Cl buffer (pH 8.5) was then added to neutralise the glycine buffer.
Western blot analysis
Preparation of cell lysates, protein electrophoresis, and transfer were as previously described (Wong et al, 2008). Primary antibody incubation was performed with DEC1 polyclonal antibody (1 : 1000). Ab-1 antibody (1 : 10 000; Calbiochem, Darmstadt, Germany) was used for detecting alpha-tubulin as a loading control.
TMA and IHC staining
A TMA was constructed using tissues of 196 oesophageal SCC patients and 32 non-oesophageal SCC patients from Henan provided by LD Wang. It comprises normal epithelium from 32 non-oesophageal SCC individuals and 35 matched normal-appearing oesophageal epithelia, 150 hyperplastic epithelia, 71 carcinoma in situ, 184 primary tumours, and 28 lymph node metastatic tumours from oesophageal SCC patients. Table 1 shows the clinical information of these patients. The family history-positive (FH+) cases were from individuals having two members or more with oesophageal SCC within three consecutive generations. Immunohistochemical staining was performed as previously described (Yuen et al, 2007) using DEC1 polyclonal antibody (1 : 100 dilution) as the primary antibody. The stained sections were examined by a pathologist (KW Chan), who had no previous knowledge of the clinico–pathological data of the patients. The intensity of staining was graded by an arbitrary scale that ranged from 0 to 3, representing negative (‘0’), weak (‘1’), moderate (‘2’), and strong (‘3’) staining, respectively. Staining values of 0 and 1 were classified as low expression, while 2 and 3 were classified as high expression.
Statistical analysis
Associations between clinical pathological information of oesophageal SCC patients and expression of DEC1 were analysed by χ2 test using SPSS. A P-value below 0.005 was considered statistically significant after Bonferroni adjustment to control for type I error from multiple comparisons (Figure 3C). The trend test for measuring the correlation of DEC1 expression status (high and low DEC1 staining) with ESCC progression states that include normal, hyperplasia, carcinoma in situ, tumour, and lymph node metastasis, was carried out by Gamma test. A P-value of <0.05 was considered statistically significant.
Subcellular fractionation
Subcellular fractionation was performed as previously described (Guillemin et al, 2005). Monolayer cultures of NE1 cells on 100 mm diameter culture dishes were harvested by trypsinisation and resuspended for 5 min on ice in 500 μl extraction buffer (10 mM HEPES, 10 mM NaCl, 1 mM KH2PO4, 5 mM NaHCO3, 5 mM EDTA, 1 mM CaCl2, and 0.5 mM MgCl2). A volume of 50 ml of 2.5 M sucrose was added to restore isotonic conditions. The lysate was centrifuged at 6300 g for 5 min. The supernatant was kept as the cytoplasmic fraction. The pellet was resuspended in 1 ml 500 μl extraction buffer with 2.5 M sucrose and then centrifuged again at 6300 g for 5 min. The pellet was considered the nuclear fraction. Anti-lamin A antibody (1 : 1000, Abcam, Cambridge, UK) was utilised to detect the nuclear fraction (1 : 100 dilution).
Immunostaining and colocalisation study
NE1 cells were grown until 80% confluence in 35 mm culture dishes on coverslips. Cells were then fixed in 4% paraformaldehyde for 10 min at room temperature and permeabilised with 1% Triton X-100 in PBS for 10 min at room temperature. The procedure for immunostaining was the same as previously described (Leung et al, 2008). A nuclear stain, DAPI (Sigma-Aldrich), was applied at a concentration of 0.1 mg ml–1. Finally, the slides were then mounted with mounting reagent (DAKO, Hamburg, Germany) and observed by confocal microscopy (LSM 510, Carl Zeiss MicroImaging GmbH, Jena, Germany). Antibodies against GM130 (BD Biosciences, Labware, MA, USA), ERGIC53 (BD Biosciences), and Calnexin (Sigma, St Louis, MO, USA) were used to stain Golgi, Golgi–endoplasmic reticulum (ER) intermediate compartment, and ER, respectively, at a 1 : 100 dilution. The images were captured by confocal microscopy. The scatter plot of the images was generated and the colocalisation coefficient was measured by ImageJ (Available at http://rsbweb.nih.gov/ij/; Developed by National Institutes of Health, Bethesda, MD, USA). Pearson's correlation coefficient indicates the degree of colocalisation, with a value close to 1 meaning perfect colocalisation. Thresholded Mander's split colocalisation coefficient provides the proportion of signals from one channel colocalising with another, with a value of zero meaning no colocalisation and a value of one corresponding to perfect colocalisation (Adler and Parmryd, 2010).
Results
Generation and validation of antibodies
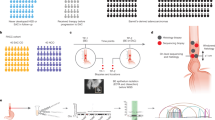
To further our study of DEC1 at the protein level, recombinant His-tagged DEC1 was purified, emulsified in adjuvant, and then used to immunise rabbits (Figure 1). The polyclonal DEC1 antibody can specifically detect GST-tagged DEC1, but not the GST protein (Figure 1B). The DEC1 antibody specifically recognises the mammalian-expressed GFP–DEC1 fusion protein, but not GFP (Figure 1B). By immunostaining using DEC1 antibody, DEC1–GFP transiently transfected HeLa cells can be detected (Figure 1C). In addition, ectopic expression of DEC1 can be detected in the DEC1 stable transfectant (C9) (Figure 1D). All experiments validated the epitope-affinity and specificity of the DEC1 antibody for western blot and immunostaining purposes. Thereafter, DEC1 antibodies were utilised to detect DEC1 protein in cell lines and tissues. Western blot analysis detected expression of both exogenous and endogenous DEC1 in DEC1 stable transfectants (SLMT-1 c4 and c9) and the immortalised oesophageal epithelial cell line, NE1 (Figure 2A). Loss of DEC1 was observed in oesophageal SCC tumour tissues compared with corresponding normal counterparts (Figure 2B). High expression of DEC1 was also detected in non-cancer normal individuals. Notably, the expression of DEC1 could also be detected by immunostaining. The DEC1 protein locates to both the cytoplasm and nucleus in NE1 and DEC1 stable transfectants (Figures 1C and 2C).
Generation and characterisation of DEC1 antibodies. (A) His-tagged DEC1 proteins were expressed and purified as an antigen to immunise rabbits. (B) Upper panel: DEC1 antibody specifically recognises recombinant GST–DEC1 fusion proteins, but not GST proteins. Lower panel: the antibody specifically recognises GFP–DEC1 fusion protein, but not GFP. (C) In immunostaining, DEC1 antibody specifically recognises GFP–DEC1 transiently transfected HeLa cells. Nonspecific IgG was utilised as a control. BF, bright field. (D) By immunostaining using DEC1 antibodies, higher expression of DEC1 is detected in stable transfectant (C9) than the vector-alone control (V1).
Endogenous DEC1 detection in primary tissues and cell lines. (A) Endogenous DEC1 expression in the immortalised epithelial cell line, NE1, and exogenous DEC1 protein in DEC1 stable transfectants (SLMT-1 c4 and c9) were detected by DEC1 antibodies. α-Tubulin serves as control for equal loading. (B) Downregulation of DEC1 is detected in tumour tissues compared with normal tissues and non-cancer normal individuals. ‘T’, tumour; ‘N’, normal-appearing oesophageal mucosa. (C) Upper panel: in NE1, immunostaining shows DEC1 localised in both nucleus and cytoplasm. A vesicular pattern was observed at the perinuclear region. Lower panel: immunostaining of NE1 with IgG primary antibody as a control. ‘DAPI’, nuclear stain (scale bar, 20 μm).
DEC1 expression is associated with lymph node metastasis, early onset of oesophageal SCC, and familial oesophageal SCC development
Tissue microarray analysis was performed using normal epithelium from 32 non-oesophageal SCC patients and tissues from 196 oesophageal SCC patients. First and foremost, the trend test was performed using Gamma test. A value of −0.404 (P-value <0.001), indicated a significant trend to show that the cancer progression is negatively correlated with DEC1 expression. Later cancer stages were correlated with lower DEC1 expression. The percentage of high DEC1 staining in hyperplasia is higher than that in normal, but the difference is not statistically significant. Figure 3A shows that expression of DEC1 was attenuated from normal/hyperplasia, to carcinoma in situ, to tumour, and to lymph node metastasis progression. Thereafter, a χ2 test comparing portion of high/low DEC1 staining in all levels of cancer progression indicated that significant difference of DEC1 staining exists between at least two levels of cancer progression (P-value <0.001). In order to identify which two levels of cancer progression contained significant differences of DEC1 staining, χ2 tests were performed to compare DEC1 staining between different levels of cancer progression two by two (i.e., normal vs hyperplasia, normal vs tumour, etc.). Expression of DEC1 was significantly abated in primary tumours compared with tissues of the normal oesophagus, hyperplasia, and carcinoma in situ (P-value⩽0.001) (Figure 3C). This is consistent with our previous in vivo and in vitro functional studies identifying DEC1 as a tumour suppressor of oesophageal SCC (Yang et al, 2005; Leung et al, 2008). Intriguingly, expression of DEC1 was significantly abrogated in tissues of lymph node metastasis compared with tissues of normal oesophagus and hyperplasia (P-value⩽0.001) (Figure 3C), suggesting a clinical association of DEC1 expression with dissemination of oesophageal SCC.
Evaluation of clinical significance of DEC1 by TMA. (A) A trend test was performed to correlate DEC1 staining in different levels of cancer progression. The expression of DEC1 continuously decreased from normal/hyperplasia, to carcinoma in situ, to tumour, and to lymph node metastasis (Gamma test, −0.44, P<0.001). ‘Normal’ includes oesophageal tissues from non-oesophageal SCC patients and normal-appearing oesophageal epithelium from oesophageal SCC patients. (B) Representative images in TMA study. High expression of DEC1 in normal and hyperplastic tissues and low expression of DEC1 in tumour and lymph node metastatic tissues are shown. Normal: tissue from non-cancer normal individuals. (Scale bar, 100 μm). (C) Chi-square analysis was performed to compare the staining of DEC1 between different levels of cancer progression. A P-value <0.005 was regarded as a significant difference after Bonferroni adjustment to control for type I error and is highlighted and bolded.
Interestingly, when specimens were analysed according to age groups, the reduced DEC1 expression levels observed in tumours compared with hyperplastic tissues is significant only for younger (30–44 years) and middle-aged (45–69) patients, but not for older (70–79 years) patients (Table 2). The loss of DEC1 expression in younger patients is more significant than in middle-aged ones (P=0.001 for younger patients and P=0.006 for middle-aged patients). These results suggest DEC1 may have a critical role in early onset of oesophageal SCC development. To determine whether loss of DEC1 is involved in familial oesophageal SCC development, we compared DEC1 expression in oesophageal SCC patients with and without a FH of oesophageal SCC. Significantly lower expression of DEC1 was observed in both hyperplastic and tumour tissues of oesophageal cancer FH+ patients, as compared with oesophageal cancer FH– patients (P=0.002 for hyperplasia; P=0.006 for tumour) (Table 2). Taken together, the TMA analysis reveals an essential role of DEC1 in not only early onset of oesophageal SCC malignancies, but also familial oesophageal SCC development.
Subcellular localisation of DEC1
Identifying the subcellular localisation of a protein provides added insight which may be useful in deciphering its function. The protein localisation program PSORT (www.psort.org) predicts that DEC1 is a cytoplasmic and nuclear protein, while pTARGET (bioapps.rit.albany.edu/pTARGET/) predicts the localisation of DEC1 at the Golgi. We confirmed the subcellular location of DEC1 after subcellular fractionation using lysates from NE1. Deleted in oesophageal cancer 1 was detected in both cytoplasmic and nuclear fractions (Figure 4A). Consistent with this result, immunostaining recognised DEC1 in both the cytoplasm and nucleus. A vesicular pattern of DEC1 staining in the cytoplasm was also detected at the perinuclear region of interphase cells (Figure 4B). Although no specific pattern of DEC1 staining was observed in mitosis (Figure 4B), the vesicular cytoplasmic DEC1 appears to localise in the Golgi and Golgi–ER intermediate compartment (Figure 4C). Organelle-specific markers, GM130 for Golgi; Calnexin for ER; and ERGIC53 for the ER–Golgi intermediate compartment, were utilised. Immunostaining with the organelle markers GM130 and ERGIC53 showed a large overlap with DEC1 (Rp=0.671 for GM130 and Rp=0.437 for ERGIC53; Figure 4C). This was not the case for Calnexin and DEC1 staining and colocalisation is not obvious (Rp=0.287; Figure 4C). In summary, DEC1 localises to the cytoplasm and nucleus and the vesicular pattern of DEC1 in the cytoplasm appears to localise at the Golgi and ER–Golgi intermediate compartment.
Identifying the localisation of DEC1. (A) Subcellular fractionation preparations from NE1 lysates show DEC1 in both cytoplasmic and nuclear fractions. α-Tubulin serves as a control for the cytoplasmic fraction. Laminin-A serves as a control for the nuclear fraction. (B) Localisation pattern of DEC1 was analysed by immunostaining during cell division by DEC1 antibody. DAPI stain for DNA. (Scale bar, 10 μm) (C) Colocalisation of DEC1 with different organelle markers analysed by confocal microscopy. Immunostaining was performed using the immortalised oesophageal epithelial cell line, NE1, with the ERGIC, GM130, Calnexin, and DEC1 antibodies. Upper panel: immunostaining with ERGIC and DEC1 antibodies. ERGIC is a marker for the ER-Golgi intermediate compartment. Colocalisation of DEC1 with ERGIC was observed (arrow). Middle panel: immunostaining with GM130 and DEC1 antibodies. GM130 is a marker for the Golgi. Colocalisation of DEC1 with GM130 was observed (arrow). Lower panel: immunostaining with Calnexin and DEC1 antibodies. Calnexin is an ER marker. No colocalisation of DEC1 with Calnexin was observed. Scale bar, 20 μm. Colocalisations (indicated by arrows) are shown under higher magnification and the region shown in the insert was analysed by ImageJ. The Pearson's correlation coefficient of DEC1 with GM130, ERGIC, and Calnexin is 0.671, 0.437, and 0.287, respectively. Thresholded Mander's split colocalisation coefficient and scatterplot are shown in Supplementary Figure 1.
Discussion
The 5-year survival rates for oesophageal SCC are 37.1% for primary cancer, 18.5% for regionally spread cancers, and 3.1% for metastatic cancer (Horner et al, 2009). This highlights the importance of cancer research in identifying biomarkers for early diagnosis, to improve survival rates of EC patients. In oesophageal SCC, LOH at marker D9S910 on 9q was associated with metastasis (Hu et al, 2000). The 9q31 locus shows an early high frequency of LOH even in low-grade dysplasia of oesophageal SCC (Mori et al, 1994). In line with these studies, we previously identified a critical 9q33-34 tumour-suppressive region and found that DEC1 is frequently downregulated at the transcriptional level and functionally involved in suppressing oesophageal SCC tumour formation in vivo (Nishiwaki et al, 2000; Yang et al, 2005; Leung et al, 2008). Evidence suggests that DEC1 may serve as a good biomarker for detecting oesophageal SCC. Henan is a high-risk oesophageal SCC area, where the oesophageal SCC incidence rate is as high as 121 per 100 000 population (Xibib et al, 2003). This TMA study using tissues from Henan patients revealed a trend of increasingly reduced DEC1 expression with progression from normal, to hyperplasia, to carcinoma, to tumour, and to lymph node metastasis. Not only does this study confirm the crucial tumour-suppressive role of DEC1 in oesophageal SCC, but it also provides a novel finding of a clinical association of DEC1 expression with oesophageal SCC malignancies and metastasis. Interestingly, DEC1 expression is abrogated in tumour tissues from younger oesophageal SCC patients, suggesting its potential utility as an oesophageal SCC detection marker for early onset of oesophageal SCC development.
Having a FH of oesophageal cancer is one of the significant risk factors for oesophageal SCC. This is also associated with several clinical features, including higher prevalence rate of double primary oesophageal SCC and having a worse prognosis than for sporadic cases (Wen et al, 2009). Socioeconomic status, living environment, and dietary habits are associated with familial clustering of EC (De Groot, 2006). Not much is known regarding the molecular alterations underlying familial oesophageal SCC, although our recent genome-wide association study has now identified susceptibility loci at PLCE1 and C20orf53 (Wang et al, 2010). Downregulation of TGF-β signalling (SMAD1) is reported in tumour tissues of familial oesophageal SCC patients (Chattopadhyay et al, 2009). In China, allelic loss of chromosome 13 regions was significantly found in patients with a positive FH of oesophageal SCC compared with negative history (Hu et al, 2003). Oesophageal SCC family members show aberrant methylation on the promoter of p16 (Abbaszadegan et al, 2005). Here, we document one more molecular alteration associated with this risk factor. Decreased expression of DEC1 in tissues of oesophageal SCC patients is significantly correlated with oesophageal SCC FH status. The statistically significant lower DEC1 staining observed in the FH+ hyperplasia vs that in the FH– hyperplasia suggests that loss or reduced DEC1 expression appears to be an early event in ESCC development in FH+ patients. Further study with larger sample sizes is needed for substantiation of the current result. The mechanistic explanation for this observation warrants further investigation.
Three independent in silico protein analysis programs, ROSETTA (http://boinc.bakerlab.org/rosetta/), SMART (http://smart.embl-heidelberg.de/), and DisEMBL 1.5 (http://dis.embl.de/) identified intrinsic disorder regions at around 10 residues at the N-terminus and the C-terminus of DEC1. Intrinsic disorder regions frequently act as sites of posttranslational modifications and are common among cell signalling and cancer-associated proteins (Iakoucheva et al, 2002). Its localisation in the cytoplasm and nuclear compartments, as observed in immunostaining and subcellular fractionation experiments, is consistent with its possible role in cellular signalling and signal transduction.
Not only does the Golgi apparatus have a well-characterised function in the secretory pathway, but it also controls centrosome organisation, cell cycle progression, cell polarisation, and cell migration. The Golgi re-orientation towards invadopodia is important for directing cell invasion (Buccione et al, 2009). Interestingly, the DEC1 antibody detects vesicular DEC1 located at the Golgi and ER–Golgi intermediate compartments. DEC1 colocalises with GM130, a cis-Golgi protein regulating cell migration (Preisinger et al, 2004). Indeed, stable expression of DEC1 in oesophageal SCC cell lines upregulates dual-specificity phosphatase 6 (Leung et al, 2008), a tumour- and cell invasion- suppressor gene that is associated with patient survival in oesophageal SCC (Wong et al, 2011). Further investigations are required to elucidate the molecular mechanism of DEC1 in oesophageal SCC.
Taken together, this TMA study reveals the important clinical relevance of DEC1 in lymph node metastatic oesophageal SCC, in early onset oesophageal SCC and familial oesophageal SCC development, further solidifying the crucial role of DEC1 in oesophageal SCC malignancies. This finding adds a novel candidate to the current repertoire of oesophageal SCC diagnostic markers. Moreover, these studies on the subcellular localisation of DEC1 show that it localises to both the cytoplasm and nucleus. Cytoplasmic vesicular DEC1 proteins appear to localise to the Golgi and ER–Golgi intermediate compartment, providing a pivotal clue for further study into the detailed molecular mechanism of DEC1 in oesophageal SCC development.
Change history
29 March 2012
This paper was modified 12 months after initial publication to switch to Creative Commons licence terms, as noted at publication
References
Abbaszadegan MR, Raziee HR, Ghafarzadegan K, Shakeri MT, Afsharnezhad S, Ghavamnasiry MR (2005) Aberrant p16 methylation, a possible epigenetic risk factor in familial esophageal squamous cell carcinoma. Int J Gastrointest Cancer 36 (1): 47–54
Adler J, Parmryd I (2010) Quantifying colocalisation by correlation: the Pearson correlation coefficient is superior to the Mander's overlap coefficient. Cytometry A 77 (8): 733–742
Buccione R, Caldieri G, Ayala I (2009) Invadopodia: specialized tumor cell structures for the focal degradation of the extracellular matrix. Cancer Metastasis Rev 28 (1–2): 137–149
Chattopadhyay I, Phukan R, Singh A, Vasudevan M, Purkayastha J, Hewitt S, Kataki A, Mahanta J, Kapur S, Saxena S (2009) Molecular profiling to identify molecular mechanism in esophageal cancer with familial clustering. Oncol Rep 21 (5): 1135–1146
De Groot KM (2006) Cancer of the oesophagus—quo vadis? S Afr Med J 96 (3): 197–198
Guillemin I, Becker M, Ociepka K, Friauf E, Nothwang HG (2005) A subcellular prefractionation protocol for minute amounts of mammalian cell cultures and tissue. Proteomics 5 (1): 35–45
Horner MJRL, Krapcho M, Neyman N, Aminou R, Howlader N, Altekruse SF, Feuer EJ, Huang L, Mariotto A, Miller BA, Lewis DR, Eisner MP, Stinchcomb DG, Edwards BK (eds) (2009) SEER Cancer Statistics Review, 1975–2006. External Web Site Icon National Cancer Institute: Bethesda, MD, based on November 2008 SEER data submission, posted to the SEER Web site
Hu N, Goldstein AM, Albert PS, Giffen C, Tang ZZ, Ding T, Taylor PR, Emmert-Buck MR (2003) Evidence for a familial esophageal cancer susceptibility gene on chromosome 13. Cancer Epidemiol Biomarkers Prev 12 (10): 1112–1115
Hu N, Roth MJ, Polymeropolous M, Tang ZZ, Emmert-Buck MR, Wang QH, Goldstein AM, Feng SS, Dawsey SM, Ding T, Zhuang ZP, Han XY, Ried T, Giffen C, Taylor PR (2000) Identification of novel regions of allelic loss from a genomewide scan of esophageal squamous-cell carcinoma in a high-risk Chinese population. Genes Chromosomes Cancer 27 (3): 217–228
Iakoucheva LM, Brown CJ, Lawson JD, Obradovic Z, Dunker AK (2002) Intrinsic disorder in cell-signaling and cancer-associated proteins. J Mol Biol 323 (3): 573–584
Ko J, Wong C, Tang C, Lau K, Lung M (2001) Frequent loss of heterozygosity on multiple chromosomes in Chinese esophageal squamous cell carcinomas. Cancer Lett 170 (2): 131–138
Leung AC, Wong VC, Yang LC, Chan PL, Daigo Y, Nakamura Y, Qi RZ, Miller LD, Liu ET, Wang LD, Li JL, Law S, Tsao SW, Lung ML (2008) Frequent decreased expression of candidate tumor suppressor gene, DEC1, and its anchorage-independent growth properties and impact on global gene expression in esophageal carcinoma. Int J Cancer 122 (3): 587–594
Li J (1982) Epidemiology of esophageal cancer in China. Natl Cancer Inst Monogr 62: 113–120
Mori T, Yanagisawa A, Kato Y, Miura K, Nishihira T, Mori S, Nakamura Y (1994) Accumulation of genetic alterations during esophageal carcinogenesis. Hum Mol Genet 3 (11): 1969–1971
Nishiwaki T, Daigo Y, Kawasoe T, Nakamura Y (2000) Isolation and mutational analysis of a novel human cDNA, DEC1 (deleted in esophageal cancer 1), derived from the tumor suppressor locus in 9q32. Genes Chromosomes Cancer 27 (2): 169–176
Preisinger C, Short B, De Corte V, Bruyneel E, Haas A, Kopajtich R, Gettemans J, Barr FA (2004) YSK1 is activated by the Golgi matrix protein GM130 and plays a role in cell migration through its substrate 14-3-3zeta. J Cell Biol 164 (7): 1009–1020
Tang J, Wan T, Wong N, Pang E, Lam K, Law S, Chow L, Ma E, Chan L, Wong J, Srivastava G (2001) Establishment and characterization of a new xenograft-derived human esophageal squamous cell carcinoma cell line SLMT-1 of Chinese origin. Cancer Genet Cytogenet 124 (1): 36–41
Tsuchiya E, Nakamura Y, Weng S, Nakagawa K, Tsuchiya S, Sugano H, Kitagawa T (1992) Allelotype of non-small cell lung carcinoma--comparison between loss of heterozygosity in squamous cell carcinoma and adenocarcinoma. Cancer Res 52 (9): 2478–2481
Wang LD, Zhou FY, Li XM, Sun LD, Song X, Jin Y, Li JM, Kong GQ, Qi H, Cui J, Zhang LQ, Yang JZ, Li JL, Li XC, Ren JL, Liu ZC, Gao WJ, Yuan L, Wei W, Zhang YR, Wang WP, Sheyhidin I, Li F, Chen BP, Ren SW, Liu B, Li D, Ku JW, Fan ZM, Zhou SL, Guo ZG, Zhao XK, Liu N, Ai YH, Shen FF, Cui WY, Song S, Guo T, Huang J, Yuan C, Wu Y, Yue WB, Feng CW, Li HL, Wang Y, Tian JY, Lu Y, Yuan Y, Zhu WL, Liu M, Fu WJ, Yang X, Wang HJ, Han SL, Chen J, Han M, Wang HY, Zhang P, Dong JC, Xing GL, Wang R, Guo M, Chang ZW, Liu HL, Guo L, Yuan ZQ, Liu H, Lu Q, Yang LQ, Zhu FG, Yang XF, Feng XS, Wang Z, Li Y, Gao SG, Qige Q, Bai LT, Yang WJ, Lei GY, Shen ZY, Chen LQ, Li EM, Xu LY, Wu ZY, Cao WK, Wang JP, Bao ZQ, Chen JL, Ding GC, Zhuang X, Zhou YF, Zheng HF, Zhang Z, Zuo XB, Dong ZM, Fan DM, He X, Wang J, Zhou Q, Zhang QX, Jiao XY, Lian SY, Ji AF, Lu XM, Wang JS, Chang FB, Lu CD, Chen ZG, Miao JJ, Fan ZL, Lin RB, Liu TJ, Wei JC, Kong QP, Lan Y, Fan YJ, Gao FS, Wang TY, Xie D, Chen SQ, Yang WC, Hong JY, Wang L, Qiu SL, Cai ZM, Zhang XJ (2010) Genome-wide association study of esophageal squamous cell carcinoma in Chinese subjects identifies susceptibility loci at PLCE1 and C20orf54. Nat Genet 42 (9): 759–763
Wen D, Wang S, Zhang L, Wei L, Zhou W, Peng Q (2009) Early onset, multiple primary malignancies, and poor prognosis are indicative of an inherited predisposition to esophageal squamous cell carcinoma for the familial as opposed to the sporadic cases--an update on over 14-year survival. Eur J Med Genet 52 (6): 381–385
Wong VC, Chen H, Ko MY, Chan KW, Chan YP, Law S, Chua D, Kwong DL, Srivastava G, Tang JC, Tsao SW, Zabarovsky ER, Stanbridge EJ, Lung ML (2011) Tumor suppressor Dual-Specificity Phosphatase 6 (DUSP6) impairs cell invasion and epithelial-mesenchymal transition (EMT)-associated phenotype. Int J Cancer; doi:10.1002/ijc.25970
Wong VC, Chan PL, Bernabeu C, Law S, Wang LD, Li JL, Tsao SW, Srivastava G, Lung ML (2008) Identification of an invasion and tumor-suppressing gene, Endoglin (ENG), silenced by both epigenetic inactivation and allelic loss in esophageal squamous cell carcinoma. Int J Cancer 123 (12): 2816–2823
Xibib S, Meilan H, Moller H, Evans H, Dixin D, Wenjie D, Jianbang L (2003) Risk factors for oesophageal cancer in Linzhou, China: a case-control study. Asian Pac J Cancer Prev 4 (2): 119–124
Yang L, Leung AC, Ko JM, Lo PH, Tang JC, Srivastava G, Oshimura M, Stanbridge EJ, Daigo Y, Nakamura Y, Tang CM, Lau KW, Law S, Lung ML (2005) Tumor suppressive role of a 2.4 Mb 9q33-q34 critical region and DEC1 in esophageal squamous cell carcinoma. Oncogene 24 (4): 697–705
Yuen HF, Chan YP, Chan KK, Chu YY, Wong ML, Law SY, Srivastava G, Wong YC, Wang X, Chan KW (2007) Id-1 and Id-2 are markers for metastasis and prognosis in oesophageal squamous cell carcinoma. Br J Cancer 97 (10): 1409–1415
Zhang H, Jin Y, Chen X, Jin C, Law S, Tsao SW, Kwong YL (2006) Cytogenetic aberrations in immortalization of esophageal epithelial cells. Cancer Genet Cytogenet 165 (1): 25–35
Acknowledgements
We acknowledge the Research Grants Council of Hong Kong Special Administrative Region, People's Republic of China for funding support to MLL. We acknowledge the University of Hong Kong Faculty of Medicine Core Facility for use of the confocal microscope.
Author information
Authors and Affiliations
Corresponding author
Additional information
Supplementary Information accompanies the paper on British Journal of Cancer website
Supplementary information
Rights and permissions
From twelve months after its original publication, this work is licensed under the Creative Commons Attribution-NonCommercial-Share Alike 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/3.0/
About this article
Cite this article
Wong, V., Ko, J., Qi, R. et al. Abrogated expression of DEC1 during oesophageal squamous cell carcinoma progression is age- and family history-related and significantly associated with lymph node metastasis. Br J Cancer 104, 841–849 (2011). https://doi.org/10.1038/bjc.2011.25
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/bjc.2011.25