Abstract
Aim:
To character the specific metabolomics profiles in the sera of Chinese patients with mild persistent asthma and to explore potential metabolic biomarkers.
Methods:
Seventeen Chinese patients with mild persistent asthma and age- and sex-matched healthy controls were enrolled. Serum samples were collected, and serum metabolites were analyzed using GC-MS coupled with a series of multivariate statistical analyses.
Results:
Clear intergroup separations existed between the asthmatic patients and control subjects. A list of differential metabolites and several top altered metabolic pathways were identified. The levels of succinate (an intermediate in tricarboxylic acid cycle) and inosine were highly upregulated in the asthmatic patients, suggesting a greater effort to breathe during exacerbation and hypoxic stress due to asthma. Other differential metabolites, such as 3,4-dihydroxybenzoic acid and phenylalanine, were also identified. Furthermore, the differential metabolites possessed higher values of area under the ROC curve (AUC), suggesting an excellent clinical ability for the prediction of asthma.
Conclusion:
Metabolic activity is significantly altered in the sera of Chinese patients with mild persistent asthma. The data might be helpful for identifying novel biomarkers and therapeutic targets for asthma.
Similar content being viewed by others
Introduction
Asthma is an inflammatory disease of the airways and one of the most common chronic illnesses1,2. It is a heterogeneous disease characterized by recurrent and reversible airflow obstruction with clinical classifications previously based mainly on patient symptoms, lung function measurements, and responses to therapy3,4. The symptoms are largely caused and sustained by abnormal airway inflammation and its effects on structural airway cells, including the epithelium and smooth muscle cells5,6,7. However, detecting airway inflammation is often difficult for clinicians. Physiological tests (ie, spirometry and peak flow measurements) and functional tests may not always reflect airway inflammation and are insensitive to small changes in the inflammatory status8. Accurate invasive airway measurements such as bronchoscopy are expensive and accompanied by adverse events8. To address these issues, research has focused on less invasive measurements of airway inflammation.
Numerous chemical tests can be used to improve the clinical diagnosis of diseases by quantifying the concentrations of a range of biomarkers9. Monitoring marker metabolite levels has become an important way to detect the pathogenesis of diseases. Biomarkers are typically measured in biofluids, such as urine, plasma, serum and exhaled breath condensate (EBC), which are commonly used to detect asthma due to the ease of collection and their minimally invasive nature10,11,12. However, many current biomarkers are not specific and lack adequate sensitivity in detection assays. Therefore, one major unmet need is to identify new biomarkers with high sensitivity and specificity that reflect the specific pathology and airway inflammation associated with asthma and can be collected in a less invasive manner.
Metabolomics represents an emerging discipline that is defined as the comprehensive assessment of low molecular weight (<1 kDa) endogenous metabolites generated by biochemical reactions under a given set of physiological conditions13,14. It utilizes fundamental analytical techniques to probe the chemical fingerprint of the samples and makes use of multivariate statistical analyses to search for disease-related potential biomarkers and metabolic pathways15,16,17,18. To date, both nuclear magnetic resonance (NMR)8,19,20,21,22 and mass spectrometry (MS)-based23,24 metabolomics techniques have been used to analyze respiratory diseases, including asthma, chronic obstructive pulmonary disease (COPD) and cystic fibrosis. However, studies have not documented specific metabolic alterations in the sera of asthmatic patients using gas chromatography mass spectrometry (GC-MS) methods.
In this study, we performed GC-TOF-MS using serum samples to detect potential metabolite changes associated with mild persistent asthma. We demonstrated clear separation between asthmatic patients and healthy controls after a series of multivariate statistical analyses. We also identified several novel metabolites that exhibited significant changes in the serum and potentially represented the top altered pathways.
Materials and methods
Human subjects
The patients (n=17) were recruited from the Respiratory Clinics at Peking University Third Hospital and were all confirmed to be diagnosed with asthma according to the Global Initiative for Asthma (GINA) guidelines. Only non-smoking, mildly persistent asthmatic patients were included in this study (Table 1). Subjects with other clinically relevant lung or medical illnesses, exacerbated asthma or respiratory infection that occurred within the 4 weeks prior to this study were excluded. We enrolled an additional 17 age- and sex-matched healthy controls who met the following criteria: (1) no history of respiratory or other diseases that might interfere with the results; (2) baseline forced expiratory volume in 1 s (FEV1) >80% predicted and FEV1/forced vital capacity (FVC) ratio >0.7; and (3) negative methacholine challenge. The parameters of lung function (ie, FEV1, FVC, and FEV1/FVC) were measured using a standard spirometry method. Demographics of the study population are briefly presented in Table 1. All healthy people and asthmatic patients provided informed consent prior to the collection of any data. This study was approved by the Ethics Committee of Peking University Third Hospital (2014071).
Serum samples
To avoid variation due to circadian rhythms, blood was drawn in the morning between 8:00 and 10:30 AM after overnight fasting (at least 8 h). Blood samples were transferred into serum gel tubes and gently inverted twice, followed by rest at room temperature for 30 min to obtain complete coagulation. The tubes were centrifuged at 2750 ×g for 10 min at 15 °C. Serum was aliquoted and stored at -80 °C prior to analysis.
Serum metabolites were analyzed with chemical derivatization according to the previously published procedure with minor modifications25,26. First, 100 μL aliquots of the thawed serum samples were transferred into 1.5-mL Eppendorf tubes, followed by the addition of 350 μL of methanol for extraction. A total of 50 μL of L-2-chlorophenylalanine (0.1 mg/mL stock in dH2O; Shanghai Hengbai Biotech Co Ltd, Shanghai, China) that acted as an internal standard was added to the tubes, which were then vortexed for 10 s. The samples were centrifuged at 12 000 rounds per minute for 10 min at 4 °C. An aliquot of 300 μL of supernatant was transferred to a 2 mL GC-MS glass vial and vacuum-dried at room temperature. The residue was derivatized using a two-step procedure. Then, 80 μL of methoxyamine hydrochloride (20 mg/mL in pyridine) was added to each vial, mixed gently and shaken for 2 h at 37 °C. Finally, 100 μL of bis-(trimethylsilyl)-trifluoroacetamide (BSTFA) plus 1% (v/v) trimethylchlorosilane (TMCS) (REGIS Technologies, Morton Grove, IL, USA) was added, and the tubes were shaken for 1 h at 70 °C. The derivatized samples were cooled to room temperature prior to GC-MS analysis.
GC-MS analysis
GC-MS analysis was performed using an Agilent 7890A gas chromatograph system (Agilent Technologies, Santa Clara, CA, USA) coupled with a Pegasus HT time-of-flight mass spectrometer (LECO, St Joseph, MI, USA). The system utilized a DB-5MS capillary column coated with 5% diphenyl cross-linked with 95% dimethylpolysiloxane (30.0 μm×250 μm inner diameter, 0.25 μm film thickness; J&W Scientific, Folsom, CA, USA). Briefly, each 1-μL aliquot of the derivatized sample was injected in splitless mode with helium as the carrier gas at a constant flow rate of 1.0 mL/min. Then, separation was achieved on a DB-5MS capillary column. The injector temperature was set at 280 °C. The column temperature was initially kept at 50 °C for 1 min and then increased to 330 °C at a rate of 10 °C/min, where it remained for 5 min. The transfer line temperature and ion source temperature were set at 280 °C and 250 °C, respectively. The energy was -70 eV in electron impact mode. The MS data were acquired in full-scan mode with a mass-to-charge ratio (m/z) range of 85–600 at a rate of 20 spectra per second after a solvent delay of 360 s.
Multivariate statistical analysis
The acquired raw GC-MS data were analyzed as previously described27. The Chroma TOF4.3X software (LECO) and LECO-Fiehn Rtx5 database were used for raw peak exacting, data baseline filtering and calibration, peak alignment, deconvolution analysis, peak identification and integration of the peak area. The missing values of the raw data were filled up by half of the minimum value; then, 296 peaks were detected, representing 272 metabolites, through the interquartile range denoising method. The retention time index (RI) method was employed for peak identification, and the RI tolerance was 5000. Additionally, the internal standard normalization method was used in this data analysis. The resulting normalized data involving the peak number, sample name and normalized peak area were fed into the SIMCA-P 13.0 software package (Umetrics, Umea, Sweden) for a series of multivariate statistical analyses.
Principal component analysis (PCA) was employed to visualize the dataset and display the similarities and differences. The partial least squares-discriminant analysis (PLS-DA) was used for cluster analysis, and linear regression was used for categorical variables in supervised learning. The R2 value was recorded to describe how well the data were mathematically reproduced; the values ranged between 0 and 1, with 1 indicating a model with perfect fitness. The Q2 value was recorded as the percent variation of the response predicted by the model, or how accurately the model could predict new data. The PLS-DA model was validated by performing permutation tests (n=200) to check its validity and then was converted into corresponding orthogonal projection to latent structures-discriminant analysis (OPLS-DA) models.
Metabolite identification, ROC curve and pathway analysis
We identified the discriminating compounds by matching the RI with data from the LECO/Fiehn Metabolomics Library27, which gave a similarity value for the compound identification accuracy. Peaks with similarities greater than 700 were assigned compound names, while those with similarities less than 200 were named “analyte”. If the similarity was between 200 and 700, the compound name was a putative annotation.
After completion of the OPLS-DA analysis, the differentially expressed metabolites between the control and asthmatic subjects could be distinguished. To choose the most accurate list of metabolites, potential candidates were chosen based on the contribution of Variable Importance for the Projection (VIP) that was extracted from the first principal component of the OPLS-DA analysis. The greater the consistent difference in metabolite levels between groups, the more important a metabolite would become in creating the final model, which is reflected by a VIP value. VIP values exceeding 1.0 were first selected as significantly changed metabolites. Then, the remaining variables were assessed by Student's t-test, with P values <0.05 considered to be statistically significant; variables that were not significantly changed were discarded.
The levels of differential metabolites were quantified by measuring the peak area after normalization and analyzed using R (the R Project). ROC curves and the area under the ROC curve (AUC) were computed using the 'pROC' package in R28. Differential metabolites of asthmatic patients were further identified and validated by searching online databases, including the Kyoto Encyclopedia of Genes and Genomes (KEGG), PubChem Compound, Chemical Entities of Biological Interest (ChEBI), Japan Chemical Substance Dictionary Web (NIKKAJI) and Chemical Abstracts Service (CAS). Then, each differential metabolite was cross-listed with the pathways in KEGG, and the top altered pathways were identified and finally constructed according to the potential functional analysis.
Results
Characterization of GC-MS data
For an efficient evaluation of the metabolic variability in the serum samples, the acquired GC-MS data were normalized and exported into the Chroma TOF4.3X software and LECO-Fiehn Rtx5 database for analysis. A total of 296 peaks were detected, and 272 metabolites were selected through the interquartile range denoising method. To better visualize the subtle similarities and differences among the complex data sets, multivariate statistical methods including PCA, PLS-DA and OPLS-DA analysis were applied.
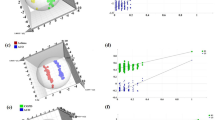
The unsupervised segregation was checked by PCA, which mainly showed the distribution of the original data. PCA reduces the dimensionality of data and summarizes the similarities and differences between multiple MS spectra using score plots. As shown in the plot, the unsupervised PCA basically demonstrated a clear separation between the asthma and control groups, except for one abnormal control sample (Figure 1A). Similarly, asthma and control subjects were also separated in the three-dimensional (3-D) PCA score plot (Figure 1B). The PCA analysis suggests that metabolic alterations indeed occur in the sera of asthmatic patients.
PCA analysis of GC-MS metabolite profiles. (A) The PCA score plot showed that the asthma samples and control subjects were scattered into two different regions. The X-axis, t[1], and Y-axis, t[2], indicated the first and second principal components, respectively. (B) 3-D score plot of the PCA analysis.
To obtain an improved separation and gain a better understanding of the variables responsible for the classification, a supervised clustering PLS-DA model analysis was applied. This model had an R2Y value of 0.955 (ie, the model explained 95.5% of the variation observed within the data) and a Q2Y value of 0.862, suggesting that the model had a very good predictive capability. The score plot of PLS-DA analysis showed a distinct separation between the asthma and control groups (Figure 2A). Furthermore, a leave-one-out cross-validation (LOOCV) was used to estimate the robustness and predictive ability of this model, and thus a permutation test was applied. The R2 and Q2 intercept values were 0.726 and -0.17, respectively, after 200 permutations (Figure 2B). The low value of the Q2 intercept indicated the robustness of the model and thus showed a low risk of over-fitting.
PLS-DA and OPLS-DA analysis of GC-MS metabolite profiles. (A) The PLS-DA score plot showed that the asthma samples and control subjects were scattered into two different regions. The X-axis, t[1], and Y-axis, t[2], indicated the first and second principal components, respectively. (B) A permutation test was performed to validate the PLS-DA model. The R2 and Q2 intercept values were 0.726 and -0.17, respectively after 200 permutations. (C) The OPLS-DA score plot showed that the asthma samples and control subjects were scattered into two different regions. The X-axis, t[1], and Y-axis, t[2], indicated the first and second principal components, respectively. (D) The OPLS-DA loading plot was constructed to display the relationship between the X-variables and the Y-variables for the first predictive component and the first Y-orthogonal component. The horizontal axis represented the X-loadings p and the Y-loadings q of the predictive component. The vertical axis represented the X-loadings p(o) and the Y-loadings s(o) for the Y-orthogonal component. X-variables situated in the vicinity of the dummy Y-variables have the highest discriminatory power between the classes.
To refine the PLS-DA analysis, OPLS-DA analysis was performed to maximize the differences between groups in the model. As shown in the score plot, asthma subjects were appreciably separated from the controls (Figure 2C). A loading plot was constructed based on the OPLS analysis that manifested the contribution of variables to the differences between the two groups. The plot showed that the important variables were situated far from the origin, which had the highest discriminatory power between the groups (Figure 2D). Taken together, these results demonstrated that the levels of metabolic components in the sera of asthmatic patients were significantly altered, which was visualized using a series of multivariate statistical methods.
Differential metabolites in asthma
We determined which differentially expressed metabolites played the greatest role in separating the two groups. We obtained VIP values from the OPLS-DA analysis; differential metabolites were selected when the VIP values were more than 1.0 and P values were less than 0.05. Based on these criteria, we found a total of 30 metabolites that were present at different abundances in asthmatic patients compared with the controls (Table 2). By ranking the VIP values according to their significance in the model, we determined that 14 specific metabolites were remarkably altered in the sera of asthmatic patients (Figure 3). The levels of differential expressed metabolites were quantified by measuring the peak area after normalization. Among them, the levels of 2-ketovaleric acid, 3,4-dihydroxybenzoic acid, 5-aminovaleric acid, ascorbate, dehydroascorbic acid, inosine, phenylalanine, and succinic acid (succinate) were significantly higher in serum samples from asthmatic patients than the healthy controls (Figure 3A–3H). In contrast, the levels of β-glycerophosphoric acid, maleamate, maleic acid, monoolein, ribose, and trans-4-hydroxy-L-proline were significantly reduced in the sera of asthmatic patients (Figure 3I–3N).
Expression levels of the significantly changed metabolites. (A) 2-ketovaleric acid; (B) 3,4-dihydroxybenzoic acid; (C) 5-aminovaleric acid; (D) ascorbate; (E) dehydroascorbic acid; (F) inosine; (G) phenylalanine; (H) succinic acid; (I) β-glycerophosphoric acid; (J) maleamate; (K) maleic acid; (L) monoolein; (M) ribose; (N) trans-4-hydroxy-L-proline; (O) ranking of VIP values.
Furthermore, we graphed ROC curves to illustrate and evaluate the prognostic performance of the differential metabolites. The closer the apex of the curve toward the upper left corner, the greater the discriminatory ability and the higher the AUC value achieved. An AUC value of 1 represents a perfect test, while an AUC value of 0.5 represents a worthless test. The AUC value is used to measure discrimination between disease and healthy subjects and manifests excellent sensitivity and specificity that have great clinical applications in the diagnosis of diseases. Interestingly, there was actually a large number of differential metabolites that presented nice ROC curves with very high AUC values (Table 2; Figure 4). Among them, three metabolites (succinate, 3,4-dihydroxybenzoic acid and inosine) possessed the top three AUC values (all more than 0.96), indicating an excellent discriminatory ability (Table 2; Figure 4). These results suggest that many, although not all, of the differential metabolites have the potential to be effective clinical indicators of asthma.
ROC graphs of metabolites with the highest AUC values.
Metabolic pathways and function analysis
It is informative to map the metabolites that describe asthma onto the top altered metabolic pathways in order to develop an improved understanding of the underlying metabolic perturbations. By searching the KEGG database, the differential metabolites were identified to be involved in several key metabolic pathways, including the tricarboxylic acid (TCA) cycle, nitrogen metabolism, glutamine and glutamate metabolism, ribose metabolism, and phenylalanine metabolism. A detailed construction of the altered pathways was generated using the reference maps from KEGG (Figure 5).
Pathway analysis of metabolomics alterations associated with asthma. The KEGG database was used to search for each differential metabolites. The illustration was generated using the reference maps from KEGG to construct the altered TCA cycle, urea cycle and amino acid metabolic pathway in asthmatic patients.
Specifically, we found that the TCA cycle was involved in the metabolic changes associated with asthma. The level of succinate (the most effective substrate following oxygen consumption) was demonstrated to be increased in the asthma group (Figure 5). The enhancement of TCA cycle metabolism with the increased abundance of TCA-cycle intermediates may result from a greater effort to breathe during exacerbation and hypoxic stress due to poor oxygenation.
We also documented the involvement of altered nitrogen metabolism (ie, the urea cycle) in asthma. Levels of L-ornithine and L-citrulline were significantly lower in the asthma group compared to the healthy controls, suggesting that the urea cycle and nitrogen metabolism were greatly dampened (Figure 5). Reduced levels of L-glutamine and L-asparagine were also observed in asthma samples, indicating alterations in amino acid metabolism (Figure 5).
Additionally, the levels of inosine, 3,4-dihydroxybenzoic acid and phenylalanine were dramatically upregulated in asthma; this result was not previously reported in metabolomics studies (Figure 3F). Inosine (a breakdown product of adenosine) can penetrate into cells to enhance the activity of many enzymes, particularly coenzyme A (CoA) and pyruvate oxidase, thereby facilitating cell metabolism under hypoxic conditions. The enhanced inosine expression suggests that the asthmatic human body may be under hypoxic circumstances with less oxygen. Notably, succinate and inosine possessed the highest VIP values, underpinning the important roles of the TCA cycle and hypoxic metabolism in asthma (Table 2 and Figure 3O). Although we did not assess any pathways related to the other differential metabolites (ie, inosine, 3,4-dihydroxybenzoic acid and phenylalanine), they all possessed the highest VIP or AUC values and might also be of great importance in monitoring asthma progression.
Collectively, we speculate that asthma is tightly associated with abnormalities in the TCA cycle, hypoxia metabolism, the urea cycle and amino acid metabolism due to the great effort to breathe and the presence of hypoxic stress.
Discussion
Respiratory diseases are a major cause of global morbidity and mortality that greatly impair the quality of life and represent a burden to the healthcare system throughout adult life14,29. Biomarkers play an ever-increasing role in the diagnosis and prognosis of respiratory diseases, the evaluation of the effects of a chosen therapy, and monitoring disease progression. The repertoire of small molecule metabolites represents a number of attractive candidates to understand disease phenotypes. To gain new insights into asthma pathogenesis, it is necessary to clarify the global metabolic alterations that characterize its progression. Fortunately, metabolomics technology has enabled us to explore particular metabolites, potential prognostic and diagnostic biomarkers, and novel pathways. Metabolomics is based on an unbiased approach that simultaneously considers a large number of metabolites in a given sample, resulting in fairly comprehensive coverage of the central pathways of primary metabolism3. To date, many initial metabolomics studies in the respiratory field were conducted with NMR due to its ease of application and non-destructive nature, but MS is now increasingly used due to its improved sensitivity and specificity30.
Among MS methods, GC-MS has been recognized as a robust metabolomics tool and has been widely applied in metabolite identification and quantification due to its high sensitivity, peak resolution, and reproducibility31. In our study, we employed GC-MS with high resolution and successfully discriminated mild persistent asthmatic patients from healthy people. The GC-MS data showed stable RI with minor fluctuations, which was greatly beneficial for matching and extracting the co-eluting peaks. Following multivariate statistical analysis of the metabolites, a clear separation of the asthma group and control group was achieved. The values of R2 (0.955) and Q2 (0.862) showed that the analysis model was stable, with good fitness and prediction capability. Notably, an abnormal point was detected in the control group after PCA or PLS-DA analysis that we considered to represent an interpersonal variation in the human samples. Because the samples were derivatized prior to GC-MS analysis, further investigation with a larger number of patients should be performed to outline the metabolic origins of the observed metabolic changes and define a possible association with mild persistent asthma.
Although the abnormal metabolic activity of asthma is primarily localized in the lung, impaired lung functions may disturb systemic metabolism. Serum is an integrated biofluid that offers the simultaneous advantage of reflecting both localized and systemic changes. More importantly, serum is commonly used for biomarker detection because its compositions are relatively well documented and its collection is less invasive. The results from our study demonstrated that serum samples were reliable for metabolomics analysis in asthma. However, it should be noted that a number of potential metabolomic pitfalls lurk in the serum, including age, gender, smoking, sampling time, nutritional status, environment, and exercise32, which might be reflected in the metabolite composition and could potentially mask molecular changes caused by asthma. Therefore, the selection of age- and sex-matched controls was extremely important in this study. Many strategies have been used to correct for interpersonal variability. For example, fasting prior to blood collection can reduce the influence of diet-related effects33; additionally, standardization of serum collection protocols with different strategies is also of great help34.
Pathway and network analyses have both been applied to metabolomics analysis, thereby vastly extending its clinical relevance and effects. We identified several top altered metabolic pathways associated with asthma. The TCA cycle is a series of enzymatic reactions that are used by aerobic organisms to generate energy and involves the oxidation of acetate derived from carbohydrates, fats or proteins. Succinate is an intermediate in this cycle and is greatly enriched in the asthma samples; this finding was consistent with previous reports of changes in the urine8 or serum22 of asthmatic patients by NMR-based metabolomics analysis. The study by Saude et al8 showed that five metabolites acting in the TCA cycle (succinate, fumarate, oxaloacetate, cis-aconitate and 2-oxoglutarate) were present at higher abundances in urine in asthmatic patients who had recently suffered an exacerbation. The upregulation of the succinate level was highly consistent with the result in our study, although we detected metabolomics alterations in different biofluids.
Notably, the changes in other intermediates from the TCA cycle were not detected in our study. This may be due to many variables. We propose that the severity of asthma may be a major cause. The study by Saude et al8 examined urine from asthma patients who recently suffered an exacerbation; thus, these patients can be considered to be affected with more severe asthma than our subjects with mild persistent asthma without exacerbation. We assume that the change in succinate in our study reflects a very early stage of metabolic alteration along the long-term course of asthma. When the asthma becomes severe, the metabolism of the whole TCA cycle will change. Additionally, there are many other contributing causes (eg, the use of urine vs serum, children vs adults, and stable asthma vs exacerbated asthma). Metabolism is complicated among different systems, organs, and individuals; therefore, metabolic profiles may also be affected to different extents.
Another study by Jung et al22 profiled the serum of asthmatic patients using NMR and showed an increase in succinate; however, glutamine was also shown to be increased, which is opposite of our results. Nevertheless, the increased TCA-cycle metabolism may suggest an enhanced requirement for energy due to the greater effort required to breathe during exacerbation and most likely is a response to hypoxic stress due to poor oxygenation. Similar shifts in TCA-cycle metabolism have also been observed during exercise35, supporting the hypothesis that elevated levels of these metabolites may be a result of enhanced breathing and hypoxic metabolism.
In addition to succinate, we also identified several novel differential metabolites (ie, inosine, 3,4-dihydroxybenzoic acid and phenylalanine) that were not previously reported to be changed in asthma36. However, we did not detect changes in aldehydes and alkanes, which were demonstrated in urinary metabolic profiles to be linked to asthma exacerbation by GC-MS in other studies37. Moreover, our results did not show unique oxidative stress-associated metabolomics profiles, such as those reported for children with severe asthma38. Inosine, a naturally occurring purine product that results from the breakdown of adenosine, is associated with inflammation, hypoxia and tissue injury39. Inosine acting on A2A or A3 adenosine receptors can regulate ovalbumin-induced allergic lung inflammation and is also an endogenous modulator of inflammatory processes observed in the lungs of asthmatic patients40. Elevated inosine in the serum appears to respond to hypoxia and may also help modulate the inflammatory process in asthma.
Because hypoxic stress is also prevalent in COPD and cystic fibrosis, the metabolic changes that occur in asthma are highly likely to be observed in those two diseases. Indeed, the changes in formate, phenylalanine and glutamine previously reported22 were also observed in COPD21,41. Similarly, inosine change was also reported in cystic fibrosis24. Thus, because it is not easy to use a single marker (inosine or succinate) to specifically or accurately predict any one respiratory disease, a combination of differential metabolites or a unique metabolic profile may be required.
The potential functions of other observed differential metabolites (ie, 3,4-dihydroxybenzoic acid and phenylalanine) in the development of asthma are not understood; however, they all possess high AUC values and may be clinically effective for the diagnosis of asthma. Further studies might be needed to focus on the functions of those metabolites in detail and exclude the possibility of false-positives.
The urea cycle was dampened in asthmatic patients, with lower levels of L-ornithine and L-citrulline, indicating the alteration of nitrogen metabolism. In the arginine metabolism pathways, L-arginine can also be catalyzed into L-citrulline and nitric oxide (NO) by nitric oxide synthase (NOS)42. It is interesting to speculate that NO production might also be reduced in the serum. However, by estimating the fraction of exhaled NO (FeNO), we know that the NO level is increased in the exhaled air of asthmatic patients, which reflects the moderate eosinophilic-mediated inflammatory pathways in the central and/or peripheral airway sites43,44. These findings may not be contradictory, because we detected FeNO mainly produced from the epithelial cells of the airways, whereas in the serum, the NO level was most likely reduced to maintain the balance of nitrogen metabolism within the human body.
Notably, because we only enrolled mild persistent subjects, our study may only provide a unique metabolic profile for mild persistent asthma. Because asthma is a very heterogeneous inflammatory disease, a larger sample size may be required to explore metabolic changes in various different asthma phenotypes.
Conclusion
Our GC-MS analysis demonstrates that metabolic alterations indeed occur in asthmatic patients, specifically patients with mild persistent asthma. Differential metabolites in the serum, such as succinate, inosine, 3,4-dihydroxybenzoic acid and phenylalanine, that possess the highest VIP or AUC values might be of great value for the diagnosis of asthma. The pathway and function analysis might provide a comprehensive understanding of asthma disease etiology and novel biomarkers for asthma assessment and treatment.
Author contribution
Chun CHANG and Wan-zhen YAO designed this study; Chun CHANG, Zhi-guo GUO and Bei HE collected and analyzed the data; Chun CHANG wrote the draft of the manuscript; Wan-zhen YAO assisted with the data analysis and interpretation and revised the manuscript.
References
Adamko DJ, Sykes BD, Rowe BH . The metabolomics of asthma: novel diagnostic potential. Chest 2012; 141: 1295–302.
Leung TF, Ko FW, Wong GW . Recent advances in asthma biomarker research. Ther Adv Respir Dis 2013; 7: 297–308.
Carraro S, Giordano G, Reniero F, Carpi D, Stocchero M, Sterk PJ, et al. Asthma severity in childhood and metabolomic profiling of breath condensate. Allergy 2013; 68: 110–7.
Holgate ST . Pathogenesis of asthma. Clin Exp Allergy 2008; 38: 872–97.
Green RH, Brightling CE, Bradding P . The reclassification of asthma based on subphenotypes. Curr Opin Allergy Clin Immunol 2007; 7: 43–50.
Holgate ST . The airway epithelium is central to the pathogenesis of asthma. Allergol Int 2008; 57: 1–10.
Fixman ED, Stewart A, Martin JG . Basic mechanisms of development of airway structural changes in asthma. Eur Respir J 2007; 29: 379–89.
Saude EJ, Skappak CD, Regush S, Cook K, Ben-Zvi A, Becker A, et al. Metabolomic profiling of asthma: diagnostic utility of urine nuclear magnetic resonance spectroscopy. J Allergy Clin Immunol 2011; 127: 757–64.
Atzei A, Atzori L, Moretti C, Barberini L, Noto A, Ottonello G, et al. Metabolomics in paediatric respiratory diseases and bronchiolitis. J Matern Fetal Neonatal Med 2011; 24: 59–62.
Dunn WB, Broadhurst D, Begley P, Zelena E, Francis-McIntyre S, Anderson N, et al. Procedures for large-scale metabolic profiling of serum and plasma using gas chromatography and liquid chromatography coupled to mass spectrometry. Nat Protoc 2011; 6: 1060–83.
Fatemi F, Sadroddiny E, Gheibi A, Mohammadi Farsani T, Kardar GA . Biomolecular markers in assessment and treatment of asthma. Respirology 2014; 19: 514–23.
Carraro S, Rezzi S, Reniero F, Heberger K, Giordano G, Zanconato S, et al. Metabolomics applied to exhaled breath condensate in childhood asthma. Am J Respir Crit Care Med 2007; 175: 986–90.
Fiehn O . Combining genomics, metabolome analysis, and biochemical modelling to understand metabolic networks. Comp Funct Genomics 2001; 2: 155–68.
Snowden S, Dahlen SE, Wheelock CE . Application of metabolomics approaches to the study of respiratory diseases. Bioanalysis 2012; 4: 2265–90.
Nicholson JK, Connelly J, Lindon JC, Holmes E . Metabonomics: a platform for studying drug toxicity and gene function. Nat Rev Drug Discov 2002; 1: 153–61.
Bogdanov M, Matson WR, Wang L, Matson T, Saunders-Pullman R, Bressman SS, et al. Metabolomic profiling to develop blood biomarkers for Parkinson's disease. Brain 2008; 131: 389–96.
Saito K, Matsuda F . Metabolomics for functional genomics, systems biology, and biotechnology. Annu Rev Plant Biol 2010; 61: 463–89.
Wu H, Liu T, Ma C, Xue R, Deng C, Zeng H, et al. GC/MS-based metabolomic approach to validate the role of urinary sarcosine and target biomarkers for human prostate cancer by microwave-assisted derivatization. Anal Bioanal Chem 2011; 401: 635–46.
Sofia M, Maniscalco M, de Laurentiis G, Paris D, Melck D, Motta A . Exploring airway diseases by NMR-based metabonomics: a review of application to exhaled breath condensate. J Biomed Biotechnol 2011; 2011: 403260.
Motta A, Paris D, D'Amato M, Melck D, Calabrese C, Vitale C, et al. NMR metabolomic analysis of exhaled breath condensate of asthmatic patients at two different temperatures. J Proteome Res 2014; 13: 6107–20.
Ubhi BK, Riley JH, Shaw PA, Lomas DA, Tal-Singer R, MacNee W, et al. Metabolic profiling detects biomarkers of protein degradation in COPD patients. Eur Respir J 2012; 40: 345–55.
Jung J, Kim SH, Lee HS, Choi GS, Jung YS, Ryu DH, et al. Serum metabolomics reveals pathways and biomarkers associated with asthma pathogenesis. Clin Exp Allergy 2013; 43: 425–33.
Mattarucchi E, Baraldi E, Guillou C . Metabolomics applied to urine samples in childhood asthma; differentiation between asthma phenotypes and identification of relevant metabolites. Biomed Chromatogr 2012; 26: 89–94.
Wetmore DR, Joseloff E, Pilewski J, Lee DP, Lawton KA, Mitchell MW, et al. Metabolomic profiling reveals biochemical pathways and biomarkers associated with pathogenesis in cystic fibrosis cells. J Biol Chem 2010; 285: 30516–22.
Qiu Y, Cai G, Su M, Chen T, Zheng X, Xu Y, et al. Serum metabolite profiling of human colorectal cancer using GC-TOFMS and UPLC-QTOFMS. J Proteome Res 2009; 8: 4844–50.
Chen T, Xie G, Wang X, Fan J, Qiu Y, Zheng X, et al. Serum and urine metabolite profiling reveals potential biomarkers of human hepatocellular carcinoma. Mol Cell Proteomics 2011; 10: M110.004945.
Kind T, Wohlgemuth G, Lee do Y, Lu Y, Palazoglu M, Shahbaz S, et al. FiehnLib: mass spectral and retention index libraries for metabolomics based on quadrupole and time-of-flight gas chromatography/mass spectrometry. Anal Chem 2009; 81: 10038–48.
Robin X, Turck N, Hainard A, Tiberti N, Lisacek F, Sanchez JC, et al. pROC: an open-source package for R and S+ to analyze and compare ROC curves. BMC Bioinformatics 2011; 12: 77.
Mukherjee AB, Zhang Z . Allergic asthma: influence of genetic and environmental factors. J Biol Chem 2011; 286: 32883–9.
Wheelock CE, Goss VM, Balgoma D, Nicholas B, Brandsma J, Skipp PJ, et al. Application of 'omics technologies to biomarker discovery in inflammatory lung diseases. Eur Respir J 2013; 42: 802–25.
Lisec J, Schauer N, Kopka J, Willmitzer L, Fernie AR . Gas chromatography mass spectrometry-based metabolite profiling in plants. Nat Protoc 2006; 1: 387–96.
Nordstrom A, Lewensohn R . Metabolomics: moving to the clinic. J Neuroimmune Pharmacol 2010; 5: 4–17.
Lloyd AJ, Beckmann M, Fave G, Mathers JC, Draper J . Proline betaine and its biotransformation products in fasting urine samples are potential biomarkers of habitual citrus fruit consumption. Br J Nutr 2011; 106: 812–24.
Walsh MC, Brennan L, Malthouse JP, Roche HM, Gibney MJ . Effect of acute dietary standardization on the urinary, plasma, and salivary metabolomic profiles of healthy humans. Am J Clin Nutr 2006; 84: 531–9.
Pechlivanis A, Kostidis S, Saraslanidis P, Petridou A, Tsalis G, Mougios V, et al. 1H NMR-based metabonomic investigation of the effect of two different exercise sessions on the metabolic fingerprint of human urine. J Proteome Res 2010; 9: 6405–16.
Amaral AF . Metabolomics of asthma. J Allergy Clin Immunol 2014; 133: 1497–9.
Loureiro CC, Duarte IF, Gomes J, Carrola J, Barros AS, Gil AM, et al. Urinary metabolomic changes as a predictive biomarker of asthma exacerbation. J Allergy Clin Immunol 2014; 133: 261–3.
Fitzpatrick AM, Park Y, Brown LA, Jones DP . Children with severe asthma have unique oxidative stress-associated metabolomic profiles. J Allergy Clin Immunol 2014; 133: 258–61.
Eltzschig HK . Adenosine: an old drug newly discovered. Anesthesiology 2009; 111: 904–15.
da Rocha Lapa F, de Oliveira AP, Accetturi BG, de Oliveira Martins I, Domingos HV, de Almeida Cabrini D, et al. Anti-inflammatory effects of inosine in allergic lung inflammation in mice: evidence for the participation of adenosine A2A and A3 receptors. Purinergic Signal 2013; 9: 325–36.
McClay JL, Adkins DE, Isern NG, O'Connell TM, Wooten JB, Zedler BK, et al. 1H Nuclear magnetic resonance metabolomics analysis identifies novel urinary biomarkers for lung function. J Proteome Res 2010; 9: 3083–90.
Bronte V, Zanovello P . Regulation of immune responses by L-arginine metabolism. Nat Rev Immunol 2005; 5: 641–54.
Barnes PJ, Dweik RA, Gelb AF, Gibson PG, George SC, Grasemann H, et al. Exhaled nitric oxide in pulmonary diseases: a comprehensive review. Chest 2010; 138: 682–92.
Hatziagorou E, Tsanakas J . Assessment of airway inflammation with exhaled NO measurement. Hippokratia 2007; 11: 51–62.
Acknowledgements
This study was supported by the Special Fund of the Chinese Medical Association for the Scientific Research of Chronic Respiratory Tract Diseases (07010440052) and National Natural Science Foundation of China-Youth Fund Project (81400017). We are grateful to Dr Yu TANG and Mr Jun-liang DENG (Biotree Bio-technology Co, Ltd, Shanghai, China) for providing help with the data analysis.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Chang, C., Guo, Zg., He, B. et al. Metabolic alterations in the sera of Chinese patients with mild persistent asthma: a GC-MS-based metabolomics analysis. Acta Pharmacol Sin 36, 1356–1366 (2015). https://doi.org/10.1038/aps.2015.102
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/aps.2015.102
Keywords
This article is cited by
-
Association of lower plasma citric acid with prolonged cough: the Nagahama study
Scientific Reports (2023)
-
The combination of machine learning and untargeted metabolomics identifies the lipid metabolism -related gene CH25H as a potential biomarker in asthma
Inflammation Research (2023)
-
Metabolomics of bronchoalveolar lavage in children with persistent wheezing
Respiratory Research (2022)
-
Effects of carbon black nanoparticles and high humidity on the lung metabolome in Balb/c mice with established allergic asthma
Environmental Science and Pollution Research (2022)
-
Histological chorioamnionitis is associated with an increased risk of wheezing in preterm children less than 34 gestational weeks
BMC Pediatrics (2021)