Abstract
Aim:
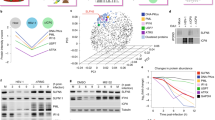
The present study examined the differential expression of proteins in HuH-7 cells and HuH-7 cells harboring in vitro-transcribed full-length hepatitis C virus 1b RNA (HuH-7-HCV), and elucidated the cellular responses to HCV replication.
Methods:
The protein profiles of matched pairs of HuH-7-HCV cells and HuH-7 mock cells were analyzed by 2-D electrophoresis (2DE). Solubilized proteins were separated in the first dimension by isoelectric focusing, and by 12.5% SDS-PAGE in the second dimension. The differential protein expression was analyzed by use of image analysis software to identify candidates for HCV infection-associated proteins.
Results:
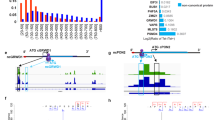
In total, 29 protein spots showed increases and 25 protein spots showed decreases in signal in HuH-7-HCV cell 2DE profiles as compared with HuH-7 mock cells. In the next step, the 10 spots showing the greatest increase and the 10 spots showing the greatest decrease were excised from gels and the proteins present were identified by Matrix-Assisted Laser Desorption/Ionization Time of Flight Mass Spectrometer (MALDI-TOF MS) or MALDI-TOF/TOF MS. In total, 13 proteins were identified successfully. The potential significance of the differential expression due to HCV replication was discussed.
Conclusion:
Our study identifies changes in the proteome of HuH-7 cells in the presence of HCV replication and yields information of the mechanism of HCV pathogenesis. These results will be useful for the identification of HCV infection-associated proteins that could be molecular targets for treatment.
Similar content being viewed by others
Article PDF
References
Alter MJ, Margolis HS, Krawczynski K, Judson FN, Mares A, Alexander WJ, et al. The sentinel counties chronic non-A NBHST: the natural history of community-acquired hepatitis C in the United States. N Engl J Med 1992; 327: 1899–905.
Choo Q, Richman KH, Han JH, Berger K, Lee C, Dong C, et al. 1991. Genetic organization and diversity of the hepatitis C virus. Proc Natl Acad Sci USA 1991; 88: 2451–5.
Reed KE, Rice CM . Overview of hepatitis C virus genome structure, polyprotein processing, and protein properties. Curr Top Microbiol Immunol 2000; 242: 55–84.
Hijikata M, Kato N, Ootsuyama Y, Nakagawa M, Shimotohno K . Gene mapping of the putative structural region of the hepatitis C virus genome by in vitro processing analysis. Proc Natl Acad Sci USA 1991; 88: 5547–51.
Blight KJ, Rice CM . Secondary structure determination of the conserved 98-base sequence at the terminus of hepatitis c virus genome RNA. J Virol 1997; 71: 7345–52.
Kolykhalov AA, Feinstone SM, Rice CM . Identification of a highly conserved sequence element at the 3′ terminus of hepatitis C virus genome RNA. J Virol 1996; 70: 3363–71.
Yanagi M, St Claire CM, Emerson SU, Purcell RH, Bukh J . In vivo analysis of the 39 untranslated region of the hepatitis C virus after in vitro mutagenesis of an infectious cDNA clone. Proc Natl Acad Sci USA 1991; 96: 2291–5.
Bartenschlager R, Lohmann V . Replication of hepatitis C virus. J Gen Virol 2000; 81: 1631–48.
Lohmann V, Körner F, Koch J, Herian U, Theilmann L, Bartenschlager R . Replication of subgenomic hepatitis C virus RNAs in a hepatoma cell line. Science 1999; 285: 110–3.
Blight KJ, McKeating JA, Marcotrigiano J, Rice CM . Efficient replication of hepatitis C virus genotype 1a RNAs in cell culture. J Virol 2003; 77: 3181–90.
Ikeda M, Yi M, Li K, Lemon SM . Selectable subgenomic and genome-length dicistronic RNAs derived from an infectious molecular clone of the HCV-N strain of hepatitis C virus replicate efficiently in cultured Huh7 cells. J Virol 2002; 76: 2997–3006.
Pietschmann T, Lohmann V, Kaul A, Krieger N, Rinck G, Rutter G, et al. Persistent and transient replication of full-length hepatitis C virus genomes in cell culture. J Virol 2002; 76: 4008–21.
Tang HL, Chu YL, Zhang SL, Guo WX, Inventors; Yangling, Daiying Biolog Engine (CN), assignee. The intact hepatitis C virus and the method for culturing it in in vitro cell culture. European Patent 1 424 390. 2004 Feb 06.
Yan JX, Wait R, Berkelman T, Harry RA, Westbrook JA, Wheeler CH, et al. A modified silver staining protocol for visualization of proteins compatible with matrix-assisted laser desorption/ionization and electrospray ionization-mass spectrometry. Electrophoresis 2000; 21: 3666–72.
Livak KJ, Schmittgen TD . Analysis of relative gene expression data using realtime quantitative PCR and the 2(-Delta Delta C (T)) Method. Methods 2001; 25: 402–8.
Gonias SL, Hembrough TA, Sankovic M . Cytokeratin 8 functions as a major plasminogen receptor in select epithelial and carcinoma cells. Front Biosci 2001; 1: 1403–11.
Omary MB, Ku NO, Toivola DM . Keratins: guardians of the liver. Hepatology 2002; 35: 251–7.
Ku NO, Darling JM, Krams SM, Esquivel CO, Keeffe EB, Sibley RK, et al. Keratin 8 and 18 mutations are risk factors for developing liver disease of multiple etiologies. Proc Natl Acad Sci USA 2003; 100: 6063–8.
Sorom AJ, Nyberg SL, Gores GJ . Keratin, fas, and cryptogenic liver failure. Liver Transpl 2002; 8: 1195–7.
Bantel H, Schulze-Osthoff K . Apoptosis in hepatitis C virus infection. Cell Death Differ 2003; 10: 48–58.
Canbay A, Friedman S, Gores GJ . Apoptosis: the nexus of liver injury and fibrosis. Hepatology 2004; 39: 273–8.
Egger D, Wölk B, Gosert R, Bianchi L, Blum HE, Moradpour D, et al. Expression of hepatitis C virus proteins induces distinct membrane alterations including a candidate viral replication complex. J Virol 2002; 76: 5974–84.
Schröder M, Kaufman RJ . ER stress and the unfolded protein response. Mutat Res 2005; 569: 29–63.
Misra UK, Gonzalez-Gronow M, Gawdi G, Pizzo SV . The Role of MTJ-1 in cell surface translocation of GRP78, a receptor for α2-macroglobulin-dependent signaling. J Immunol 2005; 174: 2092–7.
Liao J, Price D, Omary MB . Association of glucose-regulated protein (grp78) with human keratin 8. FEBS Lett 1997; 417: 316–20.
Boyd-Kimball D, Sultana R, Poon HF, Lynn BC, Casamenti F, Pepeu G, et al. Proteomic identification of proteins specifically oxidized by intracerebral injection of amyloid betapeptide (1-42) into rat brain: implications for Alzheimer's disease. Neuroscience 2005; 132: 313–24.
Cappello F, Di Stefano A, David S, Rappa F, Anzalone R, La Rocca G, et al. Hsp60 and Hsp10 down-regulation predicts bronchial epithelial carcinogenesis in smokers with chronic obstructive pulmonary disease. Cancer 2006 15; 107: 2417–24.
Caldas H, Jiang Y, Holloway MP, Fangusaro J, Mahotka C, Conway EM, et al. Survivin splice variants regulate the balance between proliferation and cell death. Oncogene 2005; 24: 1994–2007.
Dohi T, Xia F, Altieri DC . Compartmentalized phosphorylation of IAP by protein kinase A regulates cytoprotection. Mol Cell 2007; 27: 17–28.
Mizzen LA, Kabiling AN, Welch WJ . The two mammalian mito-chondrial stress proteins, grp 75 and hsp 58, transiently interact with newly synthesized mitochondrial proteins. Cell Regul 1991; 2: 165–79.
Manara GC, Sansoni P, Badiali-De Giorgi L, Gallinella G, Ferrari C, Brianti V, et al. New insights suggesting a possible role of heat shock protein 70-kD familyrelated protein in antigen processing/presentation phenomenon in humans. Blood 1993; 82: 2865–71.
Sprenger RR, Speijer D, Back JW, De Koster CG, Pannekoek H, Horrevoets AJ . Comparative proteomics of human endothelial cell caveolae and raftsusing two-dimensional gel electrophoresis and mass spectrometry. Electrophoresis 2004; 25: 156–72.
Zhang L, Ding F, Cao W, Liu Z, Liu W, Yu Z, et al. Stomatin-like protein 2 is overexpressed in cancer and involved in regulating cell growth and cell adhesion in human esophageal squamous cell carcinoma. Clin Cancer Res 2006; 12: 1639–46.
Mori T, Ishigami A, Seyama K, Onai R, Kubo S, Shimizu K, et al. Senescence marker protein-30 knockout mouse as a novel murine model of senile lung. Pathol Int 2004; 54: 167–73.
Sar P, Rath B, Subudhi U, Chainy GB, Supakar PC . Alterations in expression of senescence marker protein-30 gene by 3,3′,5-triiodo-L-thyronine (T3). Mol Cell Biochem 2007; 303: 239–42.
Raynal P, Pollard HB . Annexins: the problem of assessing the biological role for a gene family of multifunctional calcium and phospholipid-binding proteins. Biochim Biophys Acta 1994; 1197: 63–93.
Camors E, Charue D, Trouvé P, Monceau V, Loyer X, Russo-Marie F, et al. Association of annexin A5 with Na+/Ca2+ exchanger and caveolin3 in non-failing and failing human heart. J Mol Cell Cardiol 2005; 40: 47–55.
Friedman DB, Hill S, Keller JW, Merchant NB, Levy SE, Coffey RJ, et al. Proteome analysis of human colon cancer by two-dimensional difference gel electrophoresis and mass spectrometry. Proteomics 2004; 4: 793–811.
Kondo H, Rabouille C, Newman R, Levine TP, Pappin D, Freemont P, et al. p47 is a cofactor for p97-mediated membrane fusion. Nature 1997; 388: 75–8.
Author information
Authors and Affiliations
Corresponding author
Additional information
This project was supported by grants from the National Natural Science Foundation of China (No 30470089) and the National High Technology Research and Development Program of China (863 Program, No 2005A A214160).
Rights and permissions
About this article
Cite this article
Xun, M., Zhao, Sh., Cao, Cx. et al. Proteomic analysis of HuH-7 cells harboring in vitro-transcribed full-length hepatitis C virus 1b RNA. Acta Pharmacol Sin 29, 720–727 (2008). https://doi.org/10.1111/j.1745-7254.2008.00789.x
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1111/j.1745-7254.2008.00789.x
Keywords
This article is cited by
-
Activity-based protein profiling of the hepatitis C virus replication in Huh-7 hepatoma cells using a non-directed active site probe
Proteome Science (2010)
-
Proteomic analysis of mitochondria reveals a metabolic switch from fatty acid oxidation to glycolysis in the failing heart
Science in China Series C: Life Sciences (2009)