ABSTRACT
Active host-pathogen interactions take place during infection of human immunodeficiency virus type 1 (HIV-1). Outcomes of these interactions determine the efficiency of viral infection and subsequent disease progression. HIV-infected cells respond to viral invasion with various defensive strategies such as innate, cellular and humoral immune antiviral mechanisms. On the other hand, the virus has also developed various offensive tactics to suppress these host cellular responses. Among many of the viral offensive strategies, HIV-1 viral auxiliary proteins (Tat, Rev, Nef, Vif, Vpr and Vpu) play important roles in the host-pathogen interaction and thus have significant impacts on the outcome of HIV infection. One of the best examples is the interaction of Vif with a host cytidine deaminase APOBEC3G. Although specific roles of other auxiliary proteins are not as well described as Vif-APOBEC3G interaction, it is the goal of this brief review to summarize some of the preliminary findings with the hope to stimulate further discussion and investigation in this exhilarating area of research.
Similar content being viewed by others
INTRODUCTION
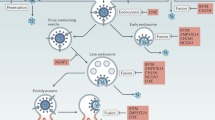
In addition to the prototypical retroviral Gag, Pol, and Env proteins, HIV-1 produces six additional proteins, i.e., Tat, Rev, Nef, Vif, Vpr and Vpu (Fig. 1, adapted from 1). While Tat and Rev are required for viral replication, Nef, Vif, Vpr and Vpu are usually dispensable for viral growth in many of the in vitro systems 2, 3, hence known as auxiliary proteins. However, these proteins are often necessary for viral replication and pathogenesis in vivo and they carry out many of the essential functions during the viral life cycle (Tab. 1). Consequently, presence or absence of these auxiliary proteins can significantly change the course and severity of the viral infection 4. In the followings, main functions of these auxiliary proteins in the process of HIV-1 infection and their roles in HIV-host interactions are briefly described.
Genome of HIV-1 (Adapted from 1).
Viral Protein U (Vpu)
Vpu is a small (9 kDa) membrane protein that enhances the release of progeny virions from infected cells and induces the degradation of the CD4 receptor. Vpu expressed in the ER interacts with a membrane-proximal domain of the cytoplasmic tail of CD4 and links it to h-βTrCP 5, a member of the F-box protein family first characterized as components of ubiquitin-ligase complexes 6. The CD4–Vpu–h-βTrCP ternary complex then recruits SKP1, another member of the ubiquitination machinery 7. As a result, CD4 is ubiquitinated and targeted to proteasomes for degradation. The ability of Vpu to increase progeny virus secretion from infected cells had been attributed initially to ion conductive membrane pore formation characteristic to cells over-expressing Vpu 8. However, a later report 9 showed that the requirement for Vpu is host cell-dependent, suggesting that Vpu may counteract an inhibitory factor expressed in some, but not the other, cells. This factor was identified recently as TASK-1, a widely expressed acid-sensitive K+ channel 10. TASK-1 is structurally homologous to Vpu, suggesting oligomerization as a possible mechanism of inactivation of ion channel activity of these proteins. However, the mechanism by which TASK-1 inhibits virion release is still unclear.
Viral Protein R (Vpr)
The viral protein R (Vpr) is a 96 amino acids small basic protein, and is well conserved in HIV-1, HIV-2 and SIV 11. Nuclear magnetic resonance (NMR) analysis suggests that Vpr protein of HIV-1 consists of an α-helix-turn-α-helix domain in the amino-terminal half from amino acids 17 to 46 and a long α-helix from 53 to 78 ended with an α turn in the carboxyl-terminal half 12, 13. The Vpr protein can be found in virions 14, cells, sera and cerebrospinal fluid of AIDS patients, indicating that it may exert its biological functions on many different targets. Despite its small size, Vpr has been shown to have multiple activities during virus replication, including effects on the nuclear import of the proviral DNA as a component of the pre-integration complex (PIC), cell cycle G2/M progression, regulation of apoptosis, and transactivation of the HIV-1 LTR as well as host cell genes.
One of the Vpr functions in the viral infection process is to mediate the nuclear import of HIV-1 PIC 15. In the cytoplasm, HIV viral RNA (in complex with several viral proteins) is reverse transcribed into DNA which then associates with the host cellular proteins to form PIC. Vpr is a component of this pre-integration complex 16, 17, 18. Vpr moves with PIC along cytoskeletal filaments and accumulates at the perinuclear region close to centrosomes 19. Though it is not yet known whether Vpr plays an active role during this movement of the PIC along microtubules, Vpr appears to participate in the subsequent steps, including the anchoring of the PIC to the nuclear envelope and the nuclear translocation of the viral DNA 15. Experiments in macrophages strongly suggest an important role for Vpr in mediating the nuclear import of HIV-1 PICs into the nucleus of nondividing cells 20. The mechanism of Vpr-mediated nuclear import is not clear, it is likely that Vpr interacts directly or indirectly with cellular machinery regulating the nucleo-cytoplasmic shuttling 21, 22, 23, 24, 25.
In addition to the effect in nuclear import, Vpr induces cell cycle G2 phase arrest in human and fission yeast cells suggesting a highly conserved effect of Vpr on cellular activities 26, 27, 28, 29, 30, 31, 32. Progression of cells from G2 phase of the cell cycle to mitosis is a tightly regulated cellular process that requires activation of the Cdc2 kinase, which determines onset of mitosis in all eukaryotic cells. In human and fission yeast cells, the activity of Cdc2 is regulated in part by the phosphorylation status of Cdc2, which is phosphorylated by Wee1 kinase during late G2 and is rapidly dephosphorylated by the Cdc25 tyrosine phosphatase to trigger entry into mitosis. These Cdc2 regulators are the downstream targets of two well-characterized G2/M checkpoint pathways which prevent cells from entering mitosis when cellular DNA is damaged or when DNA replication is inhibited. Vpr also inhibits Cdc2 through hyperphosphorylation 29, 32. However, the exact molecular mechanism leading to the hyper-phosphorylation of Cdc2 and G2 arrest is not yet clear. There are reports suggesting that Vpr induces G2 arrest by mimicking components of the DNA damage repair pathway involving ATR, Rad17 and Hus1 33, 34. However, other reports showed that Vpr modulates cell cycle G2/M transition through cellular mechanisms other than the classic mitotic checkpoints 32, 35, 36, 37. For example, Vpr-induced G2 arrest was shown to involve protein phosphatase 2A 32, 36 or a mitogen-activated protein kinase signal transduction pathway 37. It is possible that there are multiple mechanisms leading to Vpr-induced G2 arrest. Alternatively, vpr gene expression may trigger a type of cellular surveillance responses other than the well-characterized DNA damage or replication checkpoints but results in G2 arrest by impinging upon the same cellular targets, i.e., Cdc2. This premise certainly needs to be further evaluated. Biological significance of Vpr-induced G2 arrest during viral infection is also not well understood. However, HIV-1 LTR seems to be more active in the G2 phase, implying that Vpr-induced G2 arrest may confer a favorable cellular environment for efficient transcription of HIV-1 38.
Vpr also induces apoptosis in infected cells. Since a major mechanism for CD4+ T cell depletion in HIV-infected patients is apoptosis, which is induced by HIV through multiple pathways in both infected cells and non-infected “bystander” cells 39, it is expected that the apoptotic effect of Vpr may contribute to CD4+ T cell depletion. Although it is well accepted that Vpr induces apoptosis, there are studies suggesting that Vpr may also act as a negative regulator of T cell apoptosis 40, 41. In addition, it is debated whether Vpr-induced apoptosis is a result of G2 arrest. The activity of the cell cycle regulatory Wee-1 kinase associates with a decrease in Vpr-induced apoptosis, indicating a direct correlation between G2 arrest and apoptotic properties of Vpr 42. However, other reports suggested that these two Vpr activities can be separated 43, 44, 45, 46, 47. Even though the molecular mechanism of Vpr-induced apoptosis is elusive, most researchers favor the idea that Vpr induces apoptosis through mitochondria-dependent pathway 48. This intrinsic pathway for apoptosis is initiated by mitochondrial outer membrane permeabilization (MOMP) leading to release of the apoptotic factors from the space between the inner and outer mitochondrial membranes 49. Vpr binds to ANT (adenine nucleotide transporter) protein of the inner mitochondrial membrane 48, 50, 51, and can move across the outer mitochondrial membrane leading to depolarization of the inner mitochondrial membrane, swelling of the inner mitochondria and ultimately MOMP with release of the apoptosis factors. There is considerable evidence supporting this hypothesis, including the finding that the cell killing induced by Vpr can be reduced by the down-regulation of ANT levels 48, and that Vpr activates caspase-9 which initiates caspases of the intrinsic apoptotic pathway 52. However, other reports do not fit this hypothesis. Vpr was shown to locate predominantly in the nucleus or at the nuclear membrane 25, 53, 54, but not in the mitochondria. In addition, it has been reported that Vpr activates caspase-8 55, 56 which should not be activated in the intrinsic MOMP pathway.
There are several reported host responses to Vpr. Vpr is targeted by the CD8+ T-lymphocytes during the acute phase of the viral infection 57, 58. Production of some heat shock proteins (HSPs) is also responsive to vpr gene expression 59, 60, 61. Furthermore, some of the heat shock proteins, such as yeast Hsp16 or human HSP70, exert effective protective effect against some or all of the Vpr activities 62, 63, 64. Conversely, Vpr suppresses cellular 65 and humoral immune responses through adjusting the cell proliferation 40, 66, 67 or the production of the cytokines (TNFα and IL12) and chemokines (RANTES, MIP-1α and MIP-1β) 40, 67. Thus there appears to be an active and antagonistic interaction between Vpr and host anti-Vpr responses. For detailed review on this subject see 68.
Virus Infectivity Factor (Vif)
Vif protein of HIV-1 is a 192 aa protein that expresses at high levels in the cytoplasm of infected cells. Vif was thought to be important because it is essential for the reproduction of HIV-1 in peripheral blood lymphocytes, macrophages, and certain cell lines known as ‘nonpermissive’ cells 69. Vif-deficient virions produced from ‘permissive’ cells can infect ‘nonpermissive’ cells, but the virus subsequently produced is not infectious. The molecular nature of permissivity and the exact function of Vif in infection of nonpermissive cells was not known until recently when a series of reports showed that a host cellular protein known as APOBEC3G (apolipoprotein B mRNA-editing enzyme catalytic polypeptide-like 3G) is a potent inhibitor of HIV infection in the nonpermissive cells 70, 71. APOBEC3G is a member of the cytidine deaminase family, which prevents viral cDNA synthesis via deaminating deoxycytidines (dC) in the minus-strand retroviral cDNA replication intermediate 72, 73, 74, 75, 76. As a result, it creates stop codons or G-A transitions in the newly synthesized viral cDNA which is then subjective to elimination by host DNA repair machinery 74, 76. APOBEC3G confers its antiviral effect by encapsidating into the virus particles through interaction with viral Gag protein 77, 78, 79, 80, 81. Thus, APOBEC3G represents an innate host defense mechanism against HIV infection. However, the virus has also developed an offensive strategy to suppress the antiviral effect of APOBEC3G through Vif. Vif binds directly to APOBEC3G and counteracts its anti-HIV activity by promoting its degradation. Vif-mediated APOBEC3G degradation involves the recruitment of a specific E3 ligase complex, which leads to the polyubiquitylation and proteasome-mediated degradation 82, 83, 84, 85, 86.
In addition to the counteracting effect of Vif on APOBEC3G, Vif protein is specifically packaged into virus particles, where it is processed by protease. Protease-processed Vif is believed to be an important step for production of infectious viruses 87. Vif also stabilizes viral nucleoprotein complex through direct interaction with 5′ region of HIV-1 genomic RNA 88, 89, 90, 91. Moreover, Vif modulates viral reverse transcriptase through its C-terminal domain either by stimulating the binding of RT and primer or increasing the polymerization rate of RT 92.
Negative Regulator Factor(Nef)
The HIV-1 Nef protein is a 27-kDa myristoylated protein that is abundantly produced during the early phase of viral replication cycle. It is highly conserved in all primate lentiviruses, suggesting that its function is essential for survival of these pathogens. Nevertheless, early publications reported a negative effect of Nef on viral replication, hence the name 'negative factor' or Nef 93, 94. Subsequent studies, however, demonstrated that Nef plays an important role in several steps of HIV replication. In addition, it appears to be a critical pathogenic factor, as Nef-deficient SIV and HIV are significantly less pathogenic than the wild-type viruses 95, 96, 97, whereas Nef-transgenic mice show many features characteristic to HIV disease 98, 99.
The role of Nef in HIV-1 replication and disease pathogenesis is determined by at least four independent activities of this protein. First, Nef affects the cell surface expression of several cellular proteins. It down-regulates CD4 100, CD8 101, CD28 102, major histocompatibility complex class I 103 and class II 104 proteins, but upregulates the invariant chain of MHC II (CD74) 104. To modulate cell surface receptor expression, Nef utilizes several strategies, linked to distinct regions within the Nef protein (reviewed in 105). For example, down-regulation of the CD4 and CD28 receptors involves a dileucine-based motif in the second disordered loop of Nef, which connects Nef to adaptor protein (AP) complex 106, which is a part of cellular endocytosis machinery. Nef also directly binds to CD4 and CD28 using overlapping sequences within its core structure 102, thus inducing accelerated endocytosis of these proteins via clathrin-coated pits followed by lysosomal degradation. Down-regulation of MHC class I involves Nef-mediated connection in the endosomes between MHC-I's cytoplasmic tail and the phosphofurin acidic cluster sorting protein-1 (PACS-1)-dependent protein-sorting pathway 107. Since all these receptors are essential for proper functions of the immune system, modulation of their surface expression by Nef has profound effects on anti-HIV immune responses. Down-regulation of MHC I protects HIV-infected cells from host CTL response, whereas down-modulation of CD28 and CD4 probably limits the adhesion of a Nef-expressing T cell to the antigen-presenting cell, thus promoting the movement of HIV-infected cells into circulation and the spread of the virus. Another benefit for the virus from CD4 down-modulation is abolishment of interaction between the receptor and the Env protein of the budding virus, which likely increases HIV release from infected cell as well as infectivity of viral particles.
Second, Nef interferes with cellular signal transduction pathways. Nef is myristoylated on its amino-terminus and exhibits a proline-rich SH3-binding domain, both of which mediate Nef association with lipid rafts, cholesterol-rich membrane microdomains that concentrate potent signaling mediators 108. Nef was found to complex with and activate serine/threonine protein kinase PAK-2 109, which may contribute to activation of infected cell. In vitro, HIV-infected T cells produce enhanced levels of interleukin-2 during activation 108. When expressed in macrophages, Nef intersects the CD40L signaling pathway inducing secretion of chemokines and other factors that attract resting T cells and promote their infection by HIV 110, 111.
Third, Nef enhances virion infectivity and viral replication 112. This effect is mediated by Nef present in HIV virions and is due, at least in part, to the ability of Nef to induce actin remodeling and facilitate the movement of the viral core past a potentially obstructive cortical actin barrier 113. In support of this model, the infectivity-enhancing properties of Nef are eliminated by disruption of actin cytoskeleton or pseudotyping of HIV virions with VSV-G glycoprotein, which targets viral entry to endocytosis-dependent pathway thus bypassing cortical actin.
Fourth, Nef regulates cholesterol trafficking in HIV-infected cells. Cholesterol plays an important role in the HIV life cycle, as HIV assembly and budding, as well as infection of target cells all depend on plasma membrane cholesterol. Depletion of cellular cholesterol markedly and specifically reduces HIV-1 particle production 114, and cholesterol-sequestering drugs, such as beta-cyclodextrin, render the virus incompetent for cell entry 115, 116. Nef has been shown to bind cholesterol via a cholesterol-recognition motif at its carboxyl-terminus and to transport newly synthesized cholesterol to the site of viral budding 117. In addition, Nef interferes with activity of cellular cholesterol efflux machinery (MB, unpublished result), thus effectively hijacking cholesterol transport in HIV-infected cell.
Regulator of Expression of the Virion (Rev)
Rev is a ∼116 aa sequence-specific RNA binding phosphoprotein that is expressed during the early stages of HIV-1 replication 118, 119. Rev transports to cytoplasm single-spliced and un-spliced viral mRNAs that are required for expression of HIV structural proteins and production of genomic RNA. Eukaryotes have evolved a special mechanism to retain the incompletely spliced RNAs in the nucleus. This mechanism is undoubtedly beneficial to the host cell, but presents HIV-1 with a serious problem. Since HIV only has one LTR promoter, it encodes a single, genome-length primary transcript. In order to express the various incompletely spliced viral transcripts, some of HIV-1 transcripts must be transported out of the nucleus without splicing. Rev fulfills this function 120.
Rev contains at least three functional domains 119, 121. An arginine-rich domain which mediates both specific RNA binding and nuclear/nucleolar localization 122, 123, a nuclear export signal (NES) 119, 124, and a homomultimerization domain 125, 126. Homo-multimerized Rev interacts directly with importin β and the nucleolar phosphopprotein B23 via its NLS domain 127, 128. The Rev-importin β-B23 complex is recruited to the nuclear pore by the direct importin β-nucleoporin interaction. GTPase known as Ran plays a key role during the transporting process 129. In the cytoplasm, Ran presents in a Ran-GDP form allowing Rev binding to importin β. Once the importin β-Rev complex reaches the nucleus, where Ran-GTP predominates due to high concentration of Ran-GEF (Ran-specific guaninenucleotide-exchange factor) and RCC1 (regulator of chromosomal condensation 1), the interaction of importin β with Ran-GTP results in the disassembly of the Rev-importin β-B23 complex and the release of Rev cargo. In the nucleus, Rev binds to a special 234-basepair region of complex HIV RNA secondary structure called the Rev Response Element (RRE), which is located within the second intron of HIV 120. The high-affinity binding of first Rev monomer to its primary site in the RRE structure is followed by the binding of additional copies of Rev to form multimerized Rev 130, 131, 132, 133. The Rev protein, with its RNA cargo, will then bind to CRM1, also known as exportin-1 134, 135, 136, through it's nuclear export signal (NES) domain. CRM-1 forms a complex with the GTP-bound Ran and the leucine-rich NES mediating the export of the NES-containing protein from the nucleus through the nuclear pore 135. In the cytoplasm, a Ran-specific GTPase-activating protein (Ran-GAP) converts Ran-GTP to Ran-GDP, resulting in a Ran-GTP gradient across the nuclear membrane. Upon binding of RanBP1 (Ran binding protein 1) to Ran-GTP, the Crm1-Rev-Ran-GTP complex is disassembled and the Rev/RNA cargo is released. Asymmetric distribution of Ran-GEF and Ran-GAP between nucleus and cytoplasm ensures a constant Ran-GTP/GDP gradient to facilitate Crm1 recycling and continued Rev/RRE nuclear export 137, 138, 139, 140. This cycle of continuous protein shuttling between the nucleus and the cytoplasm generates a system where the small amounts of Rev present in an HIV-infected cell have the capacity to mediate the export of significant amounts of intron-containing HIV RNAs.
Besides Crm1, a number of other cellular proteins also participate in nuclear transport activity of Rev including eIF-5A (enkaryotic initiantion factor 5A) 141, 142, Sam68 (68 kDa Src-associated protein) 143, certain DEAD box protein RNA helicases (DDX3, DDX1) and hRIP (human Rev-interacting protein). The eIF-5A plays a crucial role in the nuclear export of Rev-RRE complexes and mutant eIF-5A inhibits HIV-1 replication in lymphocytes 141, 142. The precise mechanism of eIF-5A activity in Rev function remains to be defined. However, it was proposed that eIF-5A acts as an adapter that targets the Rev-NES to the nucleoplasmic face of the NPC and mediates efficient binding to Crm1 139, 144. Sam68 promotes the nuclear export of Rev in astrocytes 145. Sam68 is also required for Rev function and HIV-1 production in HeLa cells 146. In 293T, Jurkat cells and peripheral blood mononuclear cells, down-modulation of endogenous Sam68 significantly lowers HIV expression by inhibiting the CRM1-mediated export of nuclear Rev, resulting in the nuclear retention of both Rev and Crm1147. Sam68 might function through enhancement of HIV-1 RNA 3′ end processing 148. Recent research showed that certain DEAD box (Asp-Glu-Ala-Asp) protein RNA helicases (DDX3 and DDX1) and hRIP (human Rev-interacting protein) play important roles in Rev functions and HIV-1 replication. DDX3 acts as a nucleo-cytoplasmic shuttling protein, which binds CRM1 and localizes to nuclear membrane pores 149. DDX1 is a critical co-factor for Rev function, which helps maintain the proper subcellular distribution of Rev and functions through the Rev-RRE axis 150, 151. The hRIP is an essential Rev cofactor required for virus replication. Ablation of hRIP activity by a dominant-negative mutant or RNA interference, inhibits virus production by mis-localizing Rev-directed RNAs to the nuclear periphery whereas reintroduction of hRIP protein restores virus production 152, 153.
In addition to facilitating nuclear export, Rev has several additional effects on HIV RNA. Rev increases stability and translation of HIV RNA 154, 155. With the expression of Rev, the half-life of HIV RNAs in the nucleus of a T-cell line infected with HIV increases significantly 156. If Rev function is inhibited by LMB, a nuclear export inhibitor, nuclear pool of RRE-containing RNA decreases even in the presence of Rev 157. HIV-infected cells exert special mechanisms to counteract the function of Rev; the 16.4.1 protein is one of anti-Vif cellular proteins that also counteracts Rev activity. Overexpression of 16.4.1 inhibits Rev, whereas downregulation of 16.4.1 by siRNA stimulates Rev 158.
Transactivator of Transcription (Tat)
Tat is a small protein (101 amino acids in most clinical HIV-1 isolates, 86 amino acids in the laboratory HIV-1 HXB2 strain) which is essential for efficient transcription of viral genes and for viral replication. Tat potently trans-activates LTR-driven transcription, resulting in a remarkable increase of viral gene expression 159, 160, 161.
Tat increases the transcriptional rate in three different ways. First, Tat modifies chromatin conformation at the proviral integration site and makes it more suitable to viral transcription. Tat binds to a structured RNA element (TAR, transactivation-responsive region) present at the 5′ -end of viral leader mRNAs (nucleotide position +1 to +59 163) via cyclin T1 bridging between the activation domain of Tat and the TAR loop 164. Through this interaction, Tat recruits a series of transcriptional complexes, including enzymes with histone and factor acetyl transferase (HAT and FAT) activities, which modify chromatin at the proviral integration site and make it more suitable to transcription. With Tat protein, long polyadenylated RNA and increased gene expression ensue 159, 160, 161, 165.
Second, Tat recruits P-TEFb to adjust the activity of polymerase II. In mammalian cells, RNA polymerase II activity is controlled by the phosphorylation status of its carboxyl-terminal domain (CTD). Hypophosphorylation of the CTD on Ser2 correlates with low processivity, whereas hyperphosphorylation increases the processivity of the enzyme complex 166. In the absence of Tat, transcription from the HIV-1 LTR produces predominantly short RNA because hypophosphorylated RNAPII is arrested prematurely following the actions of negative elongation factors, including DSIF (5, 6-dichloro-1-beta-D-ribofuranosylbenzimidazolesensitivity-inducing factor) and NELF (negative elongation factor complex) 167. P-TEFb, one of the kinase complexes that can phosphorylate the CTD of RNA Pol II in the mammalian cells, is composed of CKD9 kinase and its cyclin partner, cyclin T 168, 169. Cdk9 kinase activity is naturally suppressed by interaction with 7SK RNA, hexamethylene bisacetamide-induced protein1 170 and indirubin-3′-monoxime. Tat binds to the TAR structure on the viral RNA and recruits P-TEFb through binding to cyclin T1 164, 169, 171. Recruitment of P-TEFb to TAR stimulates RNAPII Ser2 phosphorylation by Cdk9 170, and alters the substrate specificity of Cdk9 to include Ser5 phosphorylation of the CTD 172, resulting in the dissociation of DSIF and NELF. Recent studies demonstrated that human splicing factor SKIP (the splicing-associated c-Ski-interacting protein 173) and PP1 (protein phosphatase-1) are also required in this step 174. As a result, Tat facilitates the transcription initiation. On the other hand, Tat also facilitates transcription elongation. Acetylation of Tat at Lys50 caused by p300 or hGCN5 dissociates cyclinT1 and Tat from TAR RNA 175, 176, 177 and transfers Tat to the elongating RNAPII complex where it recruits PCAF (p300/CREB binding protein-associated factor) via the PCAF bromodomain and enhances the transcriptional elongation of HIV-1 178, 179, 180, 181. It was proposed that arginine methylation within the arginine-rich motif of HIV-1 Tat by PRTM6 (protein arginine methyltransferases) triggers the dissociation of acetylated Tat from the polymerase complex and PCAF at the end of the transcription cycle 182, 183, and the ubiquitination and dimethylation of arginines mark Tat for degradation 184. In addition, monomethylation can be reversed by the action of a Tat peptidyl arginine deaminase, and ubiquitinated Tat can be recycled after deacetylation by SIRT1 (the class III deacetylase sirtuin 1) into the transcription cycle 183, 185.
Third, Tat transactivates HIV-1 RNAs through the activation of NF-κB 186. Protein members of the Rel/NF-κB family bind to the enhancer element of the viral LTR 187, 188. In the un-stimulated normal mammalian cells, NF-κB is retained in the cytoplasm by its inhibitor protein IκB-α. Tat promotes NF-κB activation through a change in the redox state of the cell and IκB-α degradation.
In addition to its crucial role in activating viral transcription, Tat is associated with a number of additional activities 189. Extracellular Tat induces production of cytokines such as transforming growth factor beta, IL-2, or IL-6 190, 191, 192, 192, 193. Tat causes neurotoxicity in the central nervous system 194, 195, 196, 197, 198, 199, 200 and apoptosis in cultured peripheral blood mononuclear cells and some CD4 T-cell lines 201, 202, 203, 204. Another report demonstrated that Tat contributes to cell survival through up-regulation of the anti-apoptotic gene Bcl-2 205. Recently, Benasser and co-workers 206 demonstrated that Tat plays important role in abrogating nucleic acid-based adaptive immunity, RNA silencing. It is suggested that Tat impairs the cell's RNA-silencing defense by inhibiting the ability of Dicer to process precursor double-stranded RNAs into siRNAs 206.
References
Frankel AD, Young JA . HIV-1: fifteen proteins and an RNA. Annu Rev Biochem. 1998; 67:1–25.
Strebel K, Klimkait T, Martin MA . A novel gene of HIV-1, vpu, and its 16-kilodalton product. Science 1988; 241:1221–23.
Fan L, Peden K . Cell-free transmission of Vif mutants of HIV-1. Virology. Sep 1992; 190:19–29.
Bour S, Strebel K . HIV accessory proteins: multifunctional components of a complex system. Adv Pharmacol 2000; 48:75–120.
Margottin F, Bour SP, Durand H, et al. A novel human WD protein, h-beta TrCp, that interacts with HIV-1 Vpu connects CD4 to the ER degradation pathway through an F-box motif. Mol Cell 1998; 1:565–74.
Kipreos ET, Pagano M . The F-box protein family. Genome Biol 2000; 1:REVIEWS3002.
West CM . Evolutionary and functional implications of the complex glycosylation of Skp1, a cytoplasmic/nuclear glycoprotein associated with polyubiquitination. Cell Mol Life Sci 2003; 60:229–240.
Bour S, Strebel K . The HIV-1 Vpu protein: a multifunctional enhancer of viral particle release. Microbes Infect 2003; 5:1029–39.
Varthakavi V, Smith RM, Bour SP, Strebel K, Spearman P . Viral protein U counteracts a human host cell restriction that inhibits HIV-1 particle production. Proc Natl Acad Sci U S A 2003; 100:15154–9.
Hsu K, Seharaseyon J, Dong P, Bour S, Marban E . Mutual functional destruction of HIV-1 Vpu and host TASK-1 channel. Mol Cell 23 2004; 14:259–67.
Tristem M, Marshall C, Karpas A, Hill F . Evolution of the primate lentiviruses: evidence from vpx and vpr. EMBO J 1992; 11:3405–12.
Schuler W, Wecker K, de Rocquigny H, et al. NMR structure of the (52-96) C-terminal domain of the HIV-1 regulatory protein Vpr: molecular insights into its biological functions. J Mol Biol 1999; 285:2105–17.
Wecker K, Roques BP . NMR structure of the (1–51) N-terminal domain of the HIV-1 regulatory protein Vpr. Eur J Biochem 1999; 266:359–69.
Paxton W, Connor RI, Landau NR . Incorporation of Vpr into human immunodeficiency virus type 1 virions: requirement for the p6 region of gag and mutational analysis. J Virol 1993; 67:7229–7237.
Heinzinger NK, Bukinsky MI, Haggerty SA, et al. The Vpr protein of human immunodeficiency virus type 1 influences nuclear localization of viral nucleic acids in nondividing host cells. Proc Natl Acad Sci U S A 1994; 91:7311–5.
Fouchier RA, Malim MH . Nuclear import of human immunodeficiency virus type-1 preintegration complexes. Adv Virus Res 1999; 52:275–99.
Cullen BR . Journey to the center of the cell. Cell 2001; 105:697–700.
Bukrinsky M, Adzhubei A . Viral protein R of HIV-1. Rev Med Virol 1999; 9:39–49.
McDonald D, Vodicka MA, Lucero G, et al. Visualization of the intracellular behavior of HIV in living cells. J Cell Biol 2002; 159:441–52.
Connor RI, Chen BK, Choe S, Landau NR . Vpr is required for efficient replication of human immunodeficiency virus type-1 in mononuclear phagocytes. Virology 1995; 206:935–44.
Fouchier RA, Meyer BE, Simon JH, et al. Interaction of the human immunodeficiency virus type 1 Vpr protein with the nuclear pore complex. J Virol 1998; 72:6004–13.
Le Rouzic E, Mousnier A, Rustum C, et al. Docking of HIV-1 Vpr to the nuclear envelope is mediated by the interaction with the nucleoporin hCG1. J Biol Chem 2002; 277:45091–8.
Popov S, Rexach M, Ratner L, Blobel G, Bukrinsky M . Viral protein R regulates docking of the HIV-1 preintegration complex to the nuclear pore complex. J Biol Chem 1998; 273:13347–52.
Popov S, Rexach M, Zybarth G, et al. Viral protein R regulates nuclear import of the HIV-1 pre-integration complex. EMBO J 1998; 17:909–17.
Vodicka MA, Koepp DM, Silver PA, Emerman M . HIV-1 Vpr interacts with the nuclear transport pathway to promote macrophage infection. Genes Dev 1998; 12:175–85.
Di Marzio P, Choe S, Ebright M, Knoblauch R, Landau NR . Mutational analysis of cell cycle arrest, nuclear localization and virion packaging of human immunodeficiency virus type 1 Vpr. J Virol 1995; 69:7909–16.
Jowett JB, Planelles V, Poon B, Shah NP, Chen ML, Chen IS . The human immunodeficiency virus type 1 vpr gene arrests infected T cells in the G2 + M phase of the cell cycle. J Virol 1995; 69:6304–13.
He J, Choe S, Walker R, et al. Human immunodeficiency virus type 1 viral protein R (Vpr) arrests cells in the G2 phase of the cell cycle by inhibiting p34cdc2 activity. J Virol 1995; 69:6705–11.
Re F, Braaten D, Franke EK, Luban J . Human immunodeficiency virus type 1 Vpr arrests the cell cycle in G2 by inhibiting the activation of p34cdc2-cyclin B. J Virol 1995; 69:6859–64.
Bartz SR, Rogel ME, Emerman M . Human immunodeficiency virus type 1 cell cycle control: Vpr is cytostatic and mediates G2 accumulation by a mechanism which differs from DNA damage checkpoint control. J Virol 1996; 70:2324–31.
Planelles V, Jowett JB, Li QX, et al. Vpr-induced cell cycle arrest is conserved among primate lentiviruses. J Virol 1996; 70:2516–24.
Zhao Y, Cao J, O'Gorman MR, Yu M, Yogev R . Effect of human immunodeficiency virus type 1 protein R (vpr) gene expression on basic cellular function of fission yeast Schizosaccharomyces pombe. J Virol 1996; 70:5821–6.
Roshal M, Kim B, Zhu Y, Nghiem P, Planelles V . Activation of the ATR-mediated DNA damage response by the HIV-1 viral protein R. J Biol Chem 2003; 278:25879–86.
Zimmerman ES, Chen J, Andersen JL, et al. Human immunodeficiency virus type 1 Vpr-mediated G2 arrest requires Rad17 and Hus1 and induces nuclear BRCA1 and gamma-H2AX focus formation. Mol Cell Biol 2004; 24:9286–94.
Elder RT, Benko Z, Zhao Y . HIV-1 VPR modulates cell cycle G2/M transition through an alternative cellular mechanism other than the classic mitotic checkpoints. Front Biosci 2002; 7:d349–57.
Masuda M, Nagai Y, Oshima N, et al. Genetic studies with the fission yeast Schizosaccharomyces pombe suggest involvement of wee1, ppa2, and rad24 in induction of cell cycle arrest by human immunodeficiency virus type 1 Vpr. J Virol 2000; 74:2636–46.
Yoshizuka N, Yoshizuka-Chadani Y, Krishnan V, Zeichner SL . Human immunodeficiency virus type 1 Vpr-dependent cell cycle arrest through a mitogen-activated protein kinase signal transduction pathway. J Virol 2005; 79:11366–81.
Goh WC, Rogel ME, Kinsey CM, et al. HIV-1 Vpr increases viral expression by manipulation of the cell cycle: a mechanism for selection of Vpr in vivo. Nat Med 1998; 4:65–71.
Alimonti JB, Ball TB, Fowke KR . Mechanisms of CD4+ T lymphocyte cell death in human immunodeficiency virus infection and AIDS. J Gen Virol 2003; 84:1649–61.
Ayyavoo V, Mahboubi A, Mahalingam S, et al. HIV-1 Vpr suppresses immune activation and apoptosis through regulation of nuclear factor kappa B. Nat Med 1997; 3:1117–23.
Conti L, Rainaldi G, Matarrese P, et al. The HIV-1 vpr protein acts as a negative regulator of apoptosis in a human lymphoblastoid T cell line: possible implications for the pathogenesis of AIDS. J Exp Med 1998; 187:403–13.
Yuan H, Xie YM, Chen IS . Depletion of Wee-1 kinase is necessary for both human immunodeficiency virus type 1 Vpr- and gamma irradiation-induced apoptosis. J Virol 2003; 77:2063–2070.
Waldhuber MG, Bateson M, Tan J, Greenway AL, McPhee DA . Studies with GFP-Vpr fusion proteins: induction of apoptosis but ablation of cell-cycle arrest despite nuclear membrane or nuclear localization. Virology 2003; 313:91–104.
Depienne C, Roques P, Creminon C, et al. Cellular distribution and karyophilic properties of matrix, integrase, and Vpr proteins from the human and simian immunodeficiency viruses. Exp Cell Res 2000; 260:387–95.
Nishizawa M, Kamata M, Katsumata R, Aida Y . A carboxy-terminally truncated form of the human immunodeficiency virus type 1 Vpr protein induces apoptosis via G(1) cell cycle arrest. J Virol 2000; 74:6058–67.
Elder RT, Yu M, Chen M, Edelson S, Zhao Y . Cell cycle G2 arrest induced by HIV-1 Vpr in fission yeast (Schizosaccharomyces pombe) is independent of cell death and early genes in the DNA damage checkpoint. Virus Res 2000; 68:161–73.
Nishizawa M, Kamata M, Mojin T, Nakai Y, Aida Y . Induction of apoptosis by the Vpr protein of human immunodeficiency virus type 1 occurs independently of G(2) arrest of the cell cycle. Virology 2000; 276:16–26.
Jacotot E, Ravagnan L, Loeffler M, et al. The HIV-1 viral protein R induces apoptosis via a direct effect on the mitochondrial permeability transition pore. J Exp Med 2000; 191:33–46.
Green DR, Kroemer G . The pathophysiology of mitochondrial cell death. Science 2004; 305:626–9.
Brenner C, Kroemer G . The mitochondriotoxic domain of Vpr determines HIV-1 virulence. J Clin Invest 2003; 111:1455–7.
Jacotot E, Ferri KF, El Hamel C, et al. Control of mitochondrial membrane permeabilization by adenine nucleotide translocator interacting with HIV-1 viral protein rR and Bcl-2. J Exp Med 2001; 193:509–19.
Muthumani K, Hwang DS, Desai BM, et al. HIV-1 Vpr induces apoptosis through caspase 9 in T cells and peripheral blood mononuclear cells. J Biol Chem 2002; 277:37820–31.
Mahalingam S, Collman RG, Patel M, Monken CE, Srinivasan A . Functional analysis of HIV-1 Vpr: identification of determinants essential for subcellular localization. Virology 1995; 212:331–9.
Chen M, Elder RT, Yu M, et al. Mutational analysis of Vpr-induced G2 arrest, nuclear localization, and cell death in fission yeast. J Virol 1999; 73:3236–45.
Lum JJ, Cohen OJ, Nie Z, et al. Vpr R77Q is associated with long-term nonprogressive HIV infection and impaired induction of apoptosis. J Clin Invest 2003; 111:1547–54.
Patel CA, Mukhtar M, Pomerantz RJ . Human immunodeficiency virus type 1 Vpr induces apoptosis in human neuronal cells. J Virol 2000; 74:9717–26.
Altfeld M, Addo MM, Eldridge RL, et al. Vpr is preferentially targeted by CTL during HIV-1 infection. J Immunol 2001; 167:2743–52.
Mothe BR, Horton H, Carter DK, et al. Dominance of CD8 responses specific for epitopes bound by a single major histocompatibility complex class I molecule during the acute phase of viral infection. J Virol 2002; 76:875–84.
Brenner BG, Tao Y, Pearson E, Remer I, Wainberg MA . Altered constitutive and stress-regulated heat shock protein 27 expression in HIV type 1-infected cell lines. AIDS Res Hum Retroviruses 1995; 11:713–7.
Wainberg Z, Oliveira M, Lerner S, Tao Y, Brenner BG . Modulation of stress protein (hsp27 and hsp70) expression in CD4+ lymphocytic cells following acute infection with human immunodeficiency virus type-1. Virology 1997; 233:364–73.
Zhao Y, Benko Z, Liang D, et al. Small heat shock proteins as innate antiviral factors counteracted by HIV-1 viral protein R. Paper presented at: The Eleventh Conference on Retroviruses and Opportunistic Infections, San Francisco, 2004.
Benko Z, Liang D, Agbottah E, et al. Anti-Vpr activity of a yeast chaperone protein. J Virol 2004; 78:11016–29.
Iordanskiy S, Zhao Y, DiMarzio P, et al. Heat-shock protein 70 exerts opposing effects on Vpr-dependent and Vpr-independent HIV-1 replication in macrophages. Blood 2004; 104:1867–72.
Iordanskiy S, Zhao Y, Dubrovsky L, et al. Heat shock protein 70 protects cells from cell cycle arrest and apoptosis induced by human immunodeficiency virus type 1 viral protein R. J Virol. Sep 2004; 78:9697–704.
Ayyavoo V, Muthumani K, Kudchodkar S, et al. HIV-1 viral protein R compromises cellular immune function in vivo. Int Immunol 2002; 14:13–22.
Poon B, Grovit-Ferbas K, Stewart SA, Chen IS . Cell cycle arrest by Vpr in HIV-1 virions and insensitivity to antiretroviral agents. Science 1998; 281:266–9.
Refaeli Y, Levy DN, Weiner DB . The glucocorticoid receptor type II complex is a target of the HIV-1 vpr gene product. Proc Natl Acad Sci U S A 1995; 92:3621–5.
Zhao RY, Bukrinsky M, Elder RT . HIV-1 viral protein R (Vpr) & host cellular responses. Indian J Med Res 2005; 121:270–86.
Strebel K, Daugherty D, Clouse K, Cohen D, Folks T, Martin MA . The HIV ‘A’ (sor) gene product is essential for virus infectivity. Nature 1987; 328:728–30.
Jarmuz A, Chester A, Bayliss J, et al. An anthropoid-specific locus of orphan C to U RNA-editing enzymes on chromosome 22. Genomics 2002; 79:285–96.
Harris RS, Petersen-Mahrt SK, Neuberger MS . RNA editing enzyme APOBEC1 and some of its homologs can act as DNA mutators. Mol Cell 2002; 10:1247–53.
Harris RS, Bishop KN, Sheehy AM, et al. DNA deamination mediates innate immunity to retroviral infection. Cell 2003; 113:803–89.
Mangeat B, Turelli P, Caron G, et al. Broad antiretroviral defence by human APOBEC3G through lethal editing of nascent reverse transcripts. Nature 2003; 424:99–103.
Yu Q, Konig R, Pillai S, et al. Single-strand specificity of APOBEC3G accounts for minus-strand deamination of the HIV genome. Nat Struct Mol Biol 2004; 11:435–442.
Lecossier D, Bouchonnet F, Clavel F, Hance AJ . Hypermutation of HIV-1 DNA in the absence of the Vif protein. Science 2003; 300:1112.
Zhang H, Yang B, Pomerantz RJ, et al. The cytidine deaminase CEM15 induces hypermutation in newly synthesized HIV-1 DNA. Nature 2003; 424:94–8.
Alce TM, Popik W . APOBEC3G is incorporated into virus-like particles by a direct interaction with HIV-1 Gag nucleocapsid protein. J Biol Chem 2004; 279:34083–6.
Cen S, Guo F, Niu M, Saadatmand J, Deflassieux J, Kleiman L . The interaction between HIV-1 Gag and APOBEC3G. J Biol Chem 2004; 279:33177–84.
Douaisi M, Dussart S, Courcoul M, et al. HIV-1 and MLV Gag proteins are sufficient to recruit APOBEC3G into virus-like particles. Biochem Biophys Res Commun 2004; 321:566–73.
Luo K, Liu B, Xiao Z, et al. Amino-terminal region of the human immunodeficiency virus type 1 nucleocapsid is required for human APOBEC3G packaging. J Virol 2004; 78:11841–52.
Schafer A, Bogerd HP, Cullen BR . Specific packaging of APOBEC3G into HIV-1 virions is mediated by the nucleocapsid domain of the gag polyprotein precursor. Virology 2004; 328:163–8.
Marin M, Rose KM, Kozak SL, Kabat D . HIV-1 Vif protein binds the editing enzyme APOBEC3G and induces its degradation. Nat Med 2003; 9:1398–403.
Stopak K, de Noronha C, Yonemoto W, Greene WC . HIV-1 Vif blocks the antiviral activity of APOBEC3G by impairing both its translation and intracellular stability. Mol Cell 2003; 12:591–601.
Sheehy AM, Gaddis NC, Malim MH . The antiretroviral enzyme APOBEC3G is degraded by the proteasome in response to HIV-1 Vif. Nat Med 2003; 9:1404–7.
Yu X, Yu Y, Liu B, et al. Induction of APOBEC3G ubiquitination and degradation by an HIV-1 Vif-Cul5-SCF complex. Science 2003; 302:1056–60.
Conticello SG, Harris RS, Neuberger MS . The Vif protein of HIV triggers degradation of the human antiretroviral DNA deaminase APOBEC3G. Curr Biol 2003; 13:2009–13.
Khan MA, Akari H, Kao S, et al. Intravirion processing of the human immunodeficiency virus type 1 Vif protein by the viral protease may be correlated with Vif function. J Virol 2002; 76:9112–23.
Hoglund S, Ohagen A, Lawrence K, Gabuzda D . Role of vif during packing of the core of HIV-1. Virology 1994; 201:349–55.
Ohagen A, Gabuzda D . Role of Vif in stability of the human immunodeficiency virus type 1 core. J Virol 2000; 74:11055–66.
Simon JH, Malim MH . The human immunodeficiency virus type 1 Vif protein modulates the postpenetration stability of viral nucleoprotein complexes. J Virol 1996; 70:5297–305.
Henriet S, Richer D, Bernacchi S, et al. Cooperative and Specific Binding of Vif to the 5′ Region of HIV-1 Genomic RNA. J Mol Biol 2005.
Cancio R, Spadari S, Maga G . Vif is an auxiliary factor of the HIV-1 reverse transcriptase and facilitates abasic site bypass. Biochem J 2004; 383:475–82.
Ahmad N, Venkatesan S . Nef protein of HIV-1 is a transcriptional repressor of HIV-1 LTR. Science 1988; 241:1481–5.
Cheng-Mayer C, Iannello P, Shaw K, Luciw PA, Levy JA . Differential effects of nef on HIV replication: implications for viral pathogenesis in the host. Science 1989; 246:1629–1632.
Daniel MD, Kirchhoff F, Czajak SC, Sehgal PK, Desrosiers RC . Protective effects of a live attenuated SIV vaccine with a deletion in the nef gene. Science 1992; 258:1938–41.
Hofmann-Lehmann R, Vlasak J, Williams AL, et al. Live attenuated, nef-deleted SIV is pathogenic in most adult macaques after prolonged observation. AIDS 2003; 17:157–66.
Dyer WB, Geczy AF, Kent SJ, et al. Lymphoproliferative immune function in the Sydney Blood Bank Cohort, infected with natural nef/long terminal repeat mutants, and in other long-term survivors of transfusion-acquired HIV-1 infection. AIDS 1997; 11:1565–74.
Hanna Z, Kay DG, Cool M, et al. Transgenic mice expressing human immunodeficiency virus type 1 in immune cells develop a severe AIDS-like disease. J Virol 1998; 72:121–132.
Hanna Z, Kay DG, Rebai N, Guimond A, Jothy S, Jolicoeur P . Nef harbors a major determinant of pathogenicity for an AIDS-like disease induced by HIV-1 in transgenic mice. Cell 1998; 95:163–75.
Garcia JV, Miller AD . Serine phosphorylation-independent downregulation of cell-surface CD4 by nef. Nature 1991; 350:508–11.
Stove V, Van de Walle I, Naessens E, et al. Human immunodeficiency virus Nef induces rapid internalization of the T-cell coreceptor CD8alphabeta. J Virol 2005; 79:11422–33.
Swigut T, Shohdy N, Skowronski J . Mechanism for down-regulation of CD28 by Nef. EMBO J 2001; 20:1593–604.
Schwartz O, Marechal V, Le Gall S, Lemonnier F, Heard JM . Endocytosis of major histocompatibility complex class I molecules is induced by the HIV-1 Nef protein. Nat Med 1996; 2:338–42.
Schindler M, Wurfl S, Benaroch P, et al. Down-modulation of mature major histocompatibility complex class II and up-regulation of invariant chain cell surface expression are well-conserved functions of human and simian immunodeficiency virus nef alleles. J Virol 2003; 77:10548–56.
Geyer M, Fackler OT, Peterlin BM . Structure—function relationships in HIV-1 Nef. EMBO Rep 2001; 2:580–5.
Janvier K, Craig H, Hitchin D, et al. HIV-1 Nef stabilizes the association of adaptor protein complexes with membranes. J Biol Chem 2003; 278:8725–32.
Piguet V, Wan L, Borel C, et al. HIV-1 Nef protein binds to the cellular protein PACS-1 to downregulate class I major histocompatibility complexes. Nat Cell Biol 2000; 2:163–7.
Wang JK, Kiyokawa E, Verdin E, Trono D . The Nef protein of HIV-1 associates with rafts and primes T cells for activation. Proc Natl Acad Sci U S A 2000; 97:394–9.
Raney A, Kuo LS, Baugh LL, Foster JL, Garcia JV . Reconstitution and molecular analysis of an active human immunodeficiency virus type 1 Nef/p21-activated kinase 2 complex. J Virol 2005; 79:12732–41.
Schmidtmayerova H, Nottet HS, Nuovo G, et al. Human immunodeficiency virus type 1 infection alters chemokine beta peptide expression in human monocytes: implications for recruitment of leukocytes into brain and lymph nodes. Proc Natl Acad Sci U S A 1996; 93:700–4.
Swingler S, Brichacek B, Jacque JM, et al. HIV-1 Nef intersects the macrophage CD40L signalling pathway to promote resting-cell infection. Nature 2003; 424:213–9.
Chowers MY, Spina CA, Kwoh TJ, et al. Optimal infectivity in vitro of human immunodeficiency virus type 1 requires an intact nef gene. J Viroly 1994; 68:2906–2914.
Campbell EM, Nunez R, Hope TJ . Disruption of the actin cytoskeleton can complement the ability of Nef to enhance human immunodeficiency virus type 1 infectivity. J Virol 2004; 78:5745–55.
Maziere JC, Landureau JC, Giral P, et al. Lovastatin inhibits HIV-1 expression in H9 human T lymphocytes cultured in cholesterol-poor medium. Biomed Pharmacother 1994; 48:63–7.
Campbell SM, Crowe SM, Mak J . Virion-associated cholesterol is critical for the maintenance of HIV-1 structure and infectivity. AIDS 2002; 16:2253–61.
Guyader M, Kiyokawa E, Abrami L, Turelli P, Trono D . Role for human immunodeficiency virus type 1 membrane cholesterol in viral internalization. J Virol 2002; 76:10356–64.
Zheng YH, Plemenitas A, Fielding CJ, Peterlin BM . Nef increases the synthesis of and transports cholesterol to lipid rafts and HIV-1 progeny virions. Proc Natl Acad Sci U S A 2003; 100:8460–5.
Cochrane A, Kramer R, Ruben S, Levine J, Rosen CA . The human immunodeficiency virus rev protein is a nuclear phosphoprotein. Virology 1989; 171:264–6.
Malim MH, Bohnlein S, Hauber J, Cullen BR . Functional dissection of the HIV-1 Rev trans-activator—derivation of a trans-dominant repressor of Rev function. Cell 1989; 58:205–14.
Malim MH, Hauber J, Le SY, Maizel JV, Cullen BR . The HIV-1 rev trans-activator acts through a structured target sequence to activate nuclear export of unspliced viral mRNA. Nature 1989; 338:254–7.
Hope TJ, McDonald D, Huang XJ, Low J, Parslow TG . Mutational analysis of the human immunodeficiency virus type 1 Rev transactivator: essential residues near the amino terminus. J Virol 1990; 64:5360–6.
Cochrane AW, Perkins A, Rosen CA . Identification of sequences important in the nucleolar localization of human immunodeficiency virus Rev: relevance of nucleolar localization to function. J Virol 1990; 64:881–5.
Kjems J, Calnan BJ, Frankel AD, Sharp PA . Specific binding of a basic peptide from HIV-1 Rev. EMBO J 1992; 11:1119–1129.
Fischer U, Huber J, Boelens WC, Mattaj IW, Luhrmann R . The HIV-1 Rev activation domain is a nuclear export signal that accesses an export pathway used by specific cellular RNAs. Cell 1995; 82:475–83.
Hope TJ, Klein NP, Elder ME, Parslow TG . trans-dominant inhibition of human immunodeficiency virus type 1 Rev occurs through formation of inactive protein complexes. J Virol 1992; 66:1849–55.
Zapp ML, Hope TJ, Parslow TG, Green MR . Oligomerization and RNA binding domains of the type 1 human immunodeficiency virus Rev protein: a dual function for an arginine-rich binding motif. Proc Natl Acad Sci U S A 1991; 88:7734–8.
Szebeni A, Mehrotra B, Baumann A, et al. Nucleolar protein B23 stimulates nuclear import of the HIV-1 Rev protein and NLS-conjugated albumin. Biochemistry 1997; 36:3941–9.
Henderson BR, Percipalle P . Interactions between HIV Rev and nuclear import and export factors: the Rev nuclear localisation signal mediates specific binding to human importin-beta. J Mol Biol 1997; 274:693–707.
Izaurralde E, Kutay U, von Kobbe C, Mattaj IW, Gorlich D . The asymmetric distribution of the constituents of the Ran system is essential for transport into and out of the nucleus. EMBO J 1997; 16:6535–47.
Malim MH, Tiley LS, McCarn DF, Rusche JR, Hauber J, Cullen BR . HIV-1 structural gene expression requires binding of the Rev trans-activator to its RNA target sequence. Cell 1990; 60:675–83.
Iwai S, Pritchard C, Mann DA, Karn J, Gait MJ . Recognition of the high affinity binding site in rev-response element RNA by the human immunodeficiency virus type-1 rev protein. Nucleic Acids Res 1992; 20:6465–72.
Tiley LS, Malim MH, Tewary HK, Stockley PG, Cullen BR . Identification of a high-affinity RNA-binding site for the human immunodeficiency virus type 1 Rev protein. Proc Natl Acad Sci U S A 1992; 89:758–62.
Jain C, Belasco JG . Structural model for the cooperative assembly of HIV-1 Rev multimers on the RRE as deduced from analysis of assembly-defective mutants. Mol Cell 2001; 7:603–14.
Stade K, Ford CS, Guthrie C, Weis K . Exportin 1 (Crm1p) is an essential nuclear export factor. Cell 1997; 90:1041–50.
Fornerod M, Ohno M, Yoshida M, Mattaj IW . CRM1 is an export receptor for leucine-rich nuclear export signals. Cell 1997; 90:1051–60.
Fukuda M, Asano S, Nakamura T, et al. CRM1 is responsible for intracellular transport mediated by the nuclear export signal. Nature 1997; 390:308–11.
Pollard VW, Malim MH . The HIV-1 Rev protein. Annu Rev Microbiol. 1998;52:491–532.
Cullen BR . Nuclear RNA export. J Cell Sci 2003; 116:587–97.
Strebel K . Virus-host interactions: role of HIV proteins Vif, Tat, and Rev. AIDS 2003; 17 Suppl 4:S25–34.
Dayton AI . Within you, without you: HIV-1 Rev and RNA export. Retrovirology 2004; 1:35.
Bevec D, Jaksche H, Oft M, et al. Inhibition of HIV-1 replication in lymphocytes by mutants of the Rev cofactor eIF-5A. Science 1996; 271:1858–60.
Elfgang C, Rosorius O, Hofer L, Jaksche H, Hauber J, Bevec D . Evidence for specific nucleocytoplasmic transport pathways used by leucine-rich nuclear export signals. Proc Natl Acad Sci U S A 1999; 96:6229–34.
Fumagalli S, Totty NF, Hsuan JJ, Courtneidge SA . A target for Src in mitosis. Nature 1994; 368:871–4.
Hofmann W, Reichart B, Ewald A, et al. Cofactor requirements for nuclear export of Rev response element (RRE)- and constitutive transport element (CTE)-containing retroviral RNAs. An unexpected role for actin. J Cell Biol 2001; 152:895–910.
Li J, Liu Y, Park IW, He JJ . Expression of exogenous Sam68, the 68-kilodalton SRC-associated protein in mitosis, is able to alleviate impaired Rev function in astrocytes. J Virol 2002; 76:4526–35.
Modem S, Badri KR, Holland TC, Reddy TR . Sam68 is absolutely required for Rev function and HIV-1 production. Nucleic Acids Res 2005; 33:873–9.
Li J, Liu Y, Kim BO, He JJ . Direct participation of Sam68, the 68-kilodalton Src-associated protein in mitosis, in the CRM1-mediated Rev nuclear export pathway. J Virol 2002; 76:8374–82.
McLaren M, Asai K, Cochrane A . A novel function for Sam68: enhancement of HIV-1 RNA 3′ end processing. RNA 2004; 10:1119–29.
Yedavalli VS, Neuveut C, Chi YH, Kleiman L, Jeang KT . Requirement of DDX3 DEAD box RNA helicase for HIV-1 Rev-RRE export function. Cell 2004; 119:381–92.
Fang J, Kubota S, Yang B, et al. A DEAD box protein facilitates HIV-1 replication as a cellular co-factor of Rev. Virology 2004; 330:471–80.
Fang J, Acheampong E, Dave R, Wang F, Mukhtar M, Pomerantz RJ . The RNA helicase DDX1 is involved in restricted HIV-1 Rev function in human astrocytes. Virology 2005; 336:299–307.
Sanchez-Velar N, Udofia EB, Yu Z, Zapp ML . hRIP, a cellular cofactor for Rev function, promotes release of HIV RNAs from the perinuclear region. Genes Dev 2004; 18:23–34.
Yu Z, Sanchez-Velar N, Catrina IE, Kittler EL, Udofia EB, Zapp ML . The cellular HIV-1 Rev cofactor hRIP is required for viral replication. Proc Natl Acad Sci U S A 2005; 102:4027–32.
Arrigo SJ, Chen IS . Rev is necessary for translation but not cytoplasmic accumulation of HIV-1 vif, vpr, and env/vpu 2 RNAs. Genes Dev 1991; 5:808–19.
Felber BK, Hadzopoulou-Cladaras M, Cladaras C, Copeland T, Pavlakis GN . rev protein of human immunodeficiency virus type 1 affects the stability and transport of the viral mRNA. Proc Natl Acad Sci U S A 1989; 86:1495–9.
Malim MH, Cullen BR . Rev and the fate of pre-mRNA in the nucleus: implications for the regulation of RNA processing in eukaryotes. Mol Cell Biol 1993; 13:6180–9.
Otero GC, Harris ME, Donello JE, Hope TJ . Leptomycin B inhibits equine infectious anemia virus Rev and feline immunodeficiency virus rev function but not the function of the hepatitis B virus posttranscriptional regulatory element. J Virol 1998; 72:7593–7.
Kramer-Hammerle S, Ceccherini-Silberstein F, Bickel C, et al. Identification of a novel Rev-interacting cellular protein. BMC Cell Biol 2005; 6:20.
Ratnasabapathy R, Sheldon M, Johal L, Hernandez N . The HIV-1 long terminal repeat contains an unusual element that induces the synthesis of short RNAs from various mRNA and snRNA promoters. Genes Dev 1990; 4:2061–74.
Kessler M, Mathews MB . Premature termination and processing of human immunodeficiency virus type 1-promoted transcripts. J Virol 1992; 66:4488–96.
Zhou Q, Sharp PA . Novel mechanism and factor for regulation by HIV-1 Tat. EMBO J 1995; 14:321–8.
Marcello A, Zoppe M, Giacca M . Multiple modes of transcriptional regulation by the HIV-1 Tat transactivator. IUBMB Life 2001; 51:175–81.
Berkhout B, Silverman RH, Jeang KT . Tat trans-activates the human immunodeficiency virus through a nascent RNA target. Cell 1989; 59:273–82.
Wei P, Garber ME, Fang SM, Fischer WH, Jones KA . A novel CDK9-associated C-type cyclin interacts directly with HIV-1 Tat and mediates its high-affinity, loop-specific binding to TAR RNA. Cell 1998; 92:451–62.
Marshall NF, Price DH . Control of formation of two distinct classes of RNA polymerase II elongation complexes. Mol Cell Biol 1992; 12:2078–90.
Shilatifard A, Conaway RC, Conaway JW . The RNA polymerase II elongation complex. Annu Rev Biochem 2003; 72:693–715.
Kim DK, Yamaguchi Y, Wada T, Handa H . The regulation of elongation by eukaryotic RNA polymerase II: a recent view. Mol Cells 2001; 11:267–74.
Gold MO, Yang X, Herrmann CH, Rice AP . PITALRE, the catalytic subunit of TAK, is required for human immunodeficiency virus Tat transactivation in vivo. J Virol 1998; 72:4448–53.
Peng J, Zhu Y, Milton JT, Price DH . Identification of multiple cyclin subunits of human P-TEFb. Genes Dev 1998; 12:755–62.
Barboric M, Peterlin BM . A new paradigm in eukaryotic biology: HIV Tat and the control of transcriptional elongation. PLoS Biol 2005; 3:e76.
Bieniasz PD, Grdina TA, Bogerd HP, Cullen BR . Recruitment of a protein complex containing Tat and cyclin T1 to TAR governs the species specificity of HIV-1 Tat. EMBO J1998; 17:7056–65.
Garber ME, Mayall TP, Suess EM, Meisenhelder J, Thompson NE, Jones KA . CDK9 autophosphorylation regulates high-affinity binding of the human immunodeficiency virus type 1 tat-P-TEFb complex to TAR RNA. Mol Cell Biol 2000; 20:6958–69.
Bres V, Gomes N, Pickle L, Jones KA . A human splicing factor, SKIP, associates with P-TEFb and enhances transcription elongation by HIV-1 Tat. Genes Dev 2005; 19:1211–26.
Ammosova T, Jerebtsova M, Beullens M, et al. Nuclear Targeting of Protein Phosphatase-1 by HIV-1 Tat Protein. J Biol Chem 2005; 280:36364–71.
Kiernan RE, Vanhulle C, Schiltz L, et al. HIV-1 tat transcriptional activity is regulated by acetylation. EMBO J 1999; 18:6106–18.
Ott M, Schnolzer M, Garnica J, et al. Acetylation of the HIV-1 Tat protein by p300 is important for its transcriptional activity. Curr Biol 1999; 9:1489–92.
Col E, Caron C, Seigneurin-Berny D, et al. The histone acetyltransferase, hGCN5, interacts with and acetylates the HIV transactivator, Tat. J Biol Chem 2001; 276:28179–84.
Bres V, Kiernan R, Emiliani S, Benkirane M . Tat acetyl-acceptor lysines are important for human immunodeficiency virus type-1 replication. J Biol Chem 2002; 277:22215–21.
Dorr A, Kiermer V, Pedal A, et al. Transcriptional synergy between Tat and PCAF is dependent on the binding of acetylated Tat to the PCAF bromodomain. EMBO J 2002; 21:2715–23.
Mujtaba S, He Y, Zeng L, et al. Structural basis of lysine-acetylated HIV-1 Tat recognition by PCAF bromodomain. Mol Cell 2002; 9:575–86.
Kaehlcke K, Dorr A, Hetzer-Egger C, et al. Acetylation of Tat defines a cyclinT1-independent step in HIV transactivation. Mol Cell 2003; 12:167–76.
Boulanger MC, Liang C, Russell RS, et al. Methylation of Tat by PRMT6 regulates human immunodeficiency virus type 1 gene expression. J Virol 2005; 79:124–31.
Hetzer C, Dormeyer W, Schnolzer M, Ott M . Decoding Tat: the biology of HIV Tat posttranslational modifications. Microbes Infect 2005.
Bres V, Kiernan RE, Linares LK, et al. A non-proteolytic role for ubiquitin in Tat-mediated transactivation of the HIV-1 promoter. Nat Cell Biol 2003; 5:754–61.
Pagans S, Pedal A, North BJ, et al. SIRT1 regulates HIV transcription via Tat deacetylation. PLoS Biol 2005; 3:e41.
Demarchi F, d'Adda di Fagagna F, Falaschi A, Giacca M . Activation of transcription factor NF-kappaB by the Tat protein of human immunodeficiency virus type 1. J Virol 1996; 70:4427–37.
Pahl HL . Activators and target genes of Rel/NF-kappaB transcription factors. Oncogene 1999; 18:6853–66.
Hiscott J, Kwon H, Genin P . Hostile takeovers: viral appropriation of the NF-kappaB pathway. J Clin Invest 2001; 107:143–51.
Gallo RC . Tat as one key to HIV-induced immune pathogenesis and Tat (correction of Pat) toxoid as an important component of a vaccine. Proc Natl Acad Sci U S A 1999; 96:8324–6.
Zauli G, Davis BR, Re MC, et al. tat protein stimulates production of transforming growth factor-beta 1 by marrow macrophages: a potential mechanism for human immunodeficiency virus-1-induced hematopoietic suppression. Blood 1992; 80:3036–43.
Westendorp MO, Li-Weber M, Frank RW, Krammer PH . Human immunodeficiency virus type 1 Tat upregulates interleukin-2 secretion in activated T cells. J Virol 1994; 68:4177–85.
Scala G, Ruocco MR, Ambrosino C, et al. The expression of the interleukin 6 gene is induced by the human immunodeficiency virus 1 TAT protein. J Exp Med 1994; 179:961–71.
Lotz M, Clark-Lewis I, Ganu V . HIV-1 transactivator protein Tat induces proliferation and TGF beta expression in human articular chondrocytes. J Cell Biol 1994; 124:365–71.
Sabatier JM, Vives E, Mabrouk K, et al. Evidence for neurotoxic activity of tat from human immunodeficiency virus type 1. J Virol 1991; 65:961–7.
Hofman FM, Dohadwala MM, Wright AD, Hinton DR, Walker SM . Exogenous tat protein activates central nervous system-derived endothelial cells. J Neuroimmunol 1994; 54:19–28.
Nath A, Psooy K, Martin C, et al. Identification of a human immunodeficiency virus type 1 Tat epitope that is neuroexcitatory and neurotoxic. J Virol 1996; 70:1475–80.
Philippon V, Vellutini C, Gambarelli D, et al. The basic domain of the lentiviral Tat protein is responsible for damages in mouse brain: involvement of cytokines. Virology 1994; 205:519–29.
Weeks BS, Lieberman DM, Johnson B, et al. Neurotoxicity of the human immunodeficiency virus type 1 tat transactivator to PC12 cells requires the Tat amino acid 49–58 basic domain. J Neurosci Res 1995; 42:34–40.
Kim TA, Avraham HK, Koh YH, et al. HIV-1 Tat-mediated apoptosis in human brain microvascular endothelial cells. J Immunol 2003; 170:2629–37.
Haughey NJ, Mattson MP . Calcium dysregulation and neuronal apoptosis by the HIV-1 proteins Tat and gp120. J Acquir Immune Defic Syndr 2002; 31 Suppl 2:S55–61.
Li CJ, Friedman DJ, Wang C, Metelev V, Pardee AB . Induction of apoptosis in uninfected lymphocytes by HIV-1 Tat protein. Science 1995; 268:429–31.
Purvis SF, Jacobberger JW, Sramkoski RM, Patki AH, Lederman MM . HIV type 1 Tat protein induces apoptosis and death in Jurkat cells. AIDS Res Hum Retroviruses 1995; 11:443–50.
Westendorp MO, Shatrov VA, Schulze-Osthoff K, et al. HIV-1 Tat potentiates TNF-induced NF-kappa B activation and cytotoxicity by altering the cellular redox state. EMBO J 1995; 14:546–54.
Westendorp MO, Frank R, Ochsenbauer C, et al. Sensitization of T cells to CD95-mediated apoptosis by HIV-1 Tat and gp120. Nature 1995; 375:497–500.
Zhang M, Li X, Pang X, et al. Bcl-2 upregulation by HIV-1 Tat during infection of primary human macrophages in culture. J Biomed Sci 2002; 9:133–9.
Bennasser Y, Le SY, Benkirane M, Jeang KT . Evidence that HIV-1 encodes an siRNA and a suppressor of RNA silencing. Immunity 2005; 22:607–19.
Acknowledgements
The work was supported in part by grants from the National Institute of Health GM89630 and AI63080 (RYZ) and AI33776 (MB). Lin LI is a fellow sponsored by the UIC-UMB Forgarty AITRP program and a Ph.D. student jointly sponsored by UMB and the Chinese Military Academy of Medical Sciences.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
LI, L., LI, H., PAUZA, C. et al. Roles of HIV-1 auxiliary proteins in viral pathogenesis and host-pathogen interactions. Cell Res 15, 923–934 (2005). https://doi.org/10.1038/sj.cr.7290370
Issue Date:
DOI: https://doi.org/10.1038/sj.cr.7290370
Keywords
This article is cited by
-
Bioactive bone scaffolds manufactured by 3D printing and sacrificial templating of poly(ε-caprolactone) composites as filler for bone tissue engineering
Journal of Materials Science (2023)
-
Determination of Sodium Carboxymethyl Cellulose in Dairy Products by Resonance Rayleigh Scattering Spectrometry
Food Analytical Methods (2022)
-
Impact of HIV-1 Vpr manipulation of the DNA repair enzyme UNG2 on B lymphocyte class switch recombination
Journal of Translational Medicine (2020)
-
Premature Stop Codon at Residue 101 within HIV-1 Rev Does Not Influence Viral Replication of Clade BC but Severely Reduces Viral Fitness of Clade B
Virologica Sinica (2020)
-
Molecular characterization of HIV-1 genome in fission yeast Schizosaccharomyces pombe
Cell & Bioscience (2015)