Abstract
Genetic variation at 16 allozyme loci was studied in both American (Larix occidentalis Nutt., L. laricina (Du Roi) C. Koch, L. lyallii Parl.) and Eurasian (L. sibirica Ledeb., L. gmelinii Rupr., L. olgensis A. Henry, L. kaempferi (Lamb.) Carr. (=L. leptolepis (Sieb. et Zucc.) Endl.), L. kamtschatica (Rupr.) Carr. and L. decidua (Mill.)) larch species. Species with a limited range, such as L. olgensis and L. lyallii, had lower genetic variation than species with a wider range. Population differentiation within species was of the same order of magnitude among species. The resulting phylogeny indicates a clear separation between American and Eurasian species. This result is in agreement with recent palaeontological findings that suggest that gene flow between American and Eurasian species has been unlikely since the last glaciation.
Similar content being viewed by others
Introduction
The genus Larix (Pinaceae) comprises some 10 species of deciduous trees with varying geographical importance (Bobrov, 1972). For instance, L. sibirica and L. gmelinii are a main component of the Siberian forest, whereas the natural range of L. decidua is mostly limited to the mountainous areas of western and central Europe. Similarly, on the American continent L. laricina is found over most of the Canadian boreal forests, whereas L. lyallii and L. occidentalis are confined to the Rocky Mountains and the northern part of the Cascade Range. The phylogeny and phylogeography of Larix is still poorly known, and recent studies based on chloroplast DNA RFLPs (Kisanuki et al., 1995; Qian et al., 1995) did not yield an easily interpretable pattern. Not too surprisingly, given the weak morphological differentiation between species and the ease of artificial hybridization, the resulting phylogenies were not congruent with classical taxonomy based on cone bract morphology. Perhaps more surprisingly, nor was the species clustering always consistent with what could be expected based on current knowledge of glacial history. Some 60 years ago Hultén showed that the forests of Siberia and Alaska were so different that they could not have merged in the Pleistocene (Hultén, 1937). This was recently confirmed by Elias et al. (1996), who used under-sea cores of the Bering Strait to demonstrate that Beringia, the Bering land bridge that was submerged some 11 000 years ago, was a treeless Arctic tundra during the last glaciation, particularly during its maximum 22 000–14 000 years ago. This does not preclude gene flow between the two continents, through pollen or seeds, but it certainly reduces its likelihood. Furthermore, it seems consistent with divergence estimates obtained from molecular phylogenies of broad-leaved species, that indicate a divergence during the Miocene (Qiu et al., 1995; Hasebe et al., 1998; Wen et al., 1998). Hence, most probably, although Larix might have been continuously distributed over high latitudes some million years ago (LePage & Basinger, 1995), extant American larch populations originate from glacial refugia located south of the ice sheet (Whitlock, 1995) and, consequently, differentiation between them and Siberian species would be expected. Contrary to that expectation, L. laricina and L. occidentalis clustered with L. sibirica in Qian et al.’s (1995) phylogenetic analysis (the variation was too limited to draw any conclusion from Kisanuki et al.’s (1995) study).
The genetic variation at 16 allozyme loci in three North American and seven Eurasian Larix species was analysed. The genetic variation within species was assessed, the genetic distance between species estimated and a phylogenetic tree constructed. The use of allozyme data for phylogenetic reconstruction has a weak resolution compared with RFLP or sequence data because of the limited number of loci available. However, the possibility of analysing a large number of individuals makes this approach quite reasonable for a first, large scale, investigation (Qiu et al., 1995).
Materials and methods
Sampling
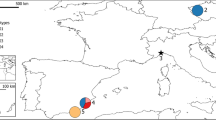
The species investigated, the number of populations per species and their locations are given in Table 1. Fig. 1 gives the distribution range of all Larix species. Seed samples were obtained from four populations of L. occidentalis, three populations of L. laricina and one population of L. lyallii. Each of the eight populations was characterized by seeds from at least nine trees. These seeds were planted and grown into seedlings, which were then used for protein electrophoresis. Samples from 15 populations of L. sibirica, six of L. gmelinii, three of L. olgensis, two of L. decidua and one each of L. kaempferi (=L. leptolepis) and L. kamtschatica were analysed. In Eurasian Larix species adult needles and seedlings were analysed. A full description of the Eurasian samples is given in Semerikov et al. (1998).
Distribution of Larix taxa. (1) L. decidua; (2) L. sibirica; (3) L. gmelinii; (4) L. cajanderi; (5) L. kamtschatica; (6) L. kaempferi (=L. leptolepis); (7) L. olgensis; (8) putative hybrid L. gmelinii × L. olgensis; (9) L. principis rupprechtii; (10) L. mastersiana; (11) L. potaninii; (12) L. griffithiana; (13) L. laricina; (14) L. occidentalis; (15) L. lyallii. Adapted from Bobrov (1972).
Allozymes
Twelve enzyme systems representing 16 protein loci were examined: glutamic-oxaloacetic transaminase (GOT, EC 2.6.1.1, two loci), glutamate dehydrogenase (GDH EC 1.4.1.2), isocitrate dehydrogenase (IDH, EC 1.1.1.42), diaphorase (DIA, EC 1.8.1.4), phosphoglucoisomerase (PGI, EC 5.3.1.9, two loci), phosphogluconate dehydrogenase (6-PGD, EC 1.1.1.44), shikimate dehydrogenase (SKDH, EC 1.1.1.25), glucose-6-phosphate dehydrogenase (G6PDH, EC 1.1.1.49), phosphoglucomutase (PGM, EC 5.4.2.2, two loci), superoxide dismutase (SOD, EC 1.15.1.1, two loci), fluorescent esterase (EST-F, EC 3.1.1.1) and formate dehydrogenase (FDH, EC 1.2.1.2). Mendelian segregation for these allozymes was shown in Larix species by Cheliak & Pitel (1985), Fins & Seeb (1986) and Lewandowski & Mejnartowicz (1990). In GOT and PGD, third and second zones of electrophoretic mobility, respectively, were observed, but they were not considered here as their reproducibility was weak. Finally, homology of the loci among species was established on the basis of the common alleles.
Protein extraction and polyacrylamide gel electrophoresis were carried out according to Shurkhal et al. (1992) and Semerikov & Matveev (1995). Needles (150 mg) and insoluble PVP (150 mg) were ground with liquid nitrogen in a mortar, and mixed with extraction buffer. The extraction buffer was composed of 1 M sucrose, 5.7 mM-ascorbic acid, 8.3 mL-cysteine, 0.02 M dithiothreitol, and 1.5 mM aminocapronic acid dissolved in electrode buffer diluted 1 : 1.7. One mL Tween-80 was added to 100 mL of this solution and after 2–14 h at 4°C the mixture of extraction buffer, ground needles and PVP was filtered through nylon filters. The supernatant was centrifuged after adding a small amount of CCl4 in order to improve the centrifugation of the supernatant from small tissue particles. Extracts from seedlings were prepared by grinding needles in 0.15 mL extraction buffer and centrifuging with CCl4. Electrophoresis in 7% polyacrylamide gel in the tris-EDTA-borate system was conducted. The electrode buffer (pH 8.0) was: 116 mM tris, 3.5 mM EDTA, 161 mM boric acid and the gel buffer (pH 8.6) was 118 mM tris, 3.5 mM EDTA, 118 mM boric acid. Histochemical staining was carried out using standard methods (Harris & Hopkinson, 1976).
Data analysis
The within-species genetic variation was analysed using GENEPOP (version 2; Raymond & Rousset, 1995) and FSTAT (version 1.2; Goudet, 1995).
Hardy–Weinberg expectations The fit of genotypic distributions to Hardy–Weinberg expectations was tested by the exact test proposed by Guo & Thompson (1992). The overall significance for each locus was estimated by Fisher’s combined probability test (Fisher, 1954). According to this test, if P-values are obtained for each locus separately under the null hypothesis, then −2∑i=1n log(Pi) is distributed according to a χ2 distribution with n degrees of freedom, where n is the number of loci and Pi is the P-value for the ith locus. Heterozygote deficiencies or excesses were tested using an exact test (Rousset & Raymond, 1995).
Population differentiation Wright’s F-statistics, FIS, FIT and FST, were estimated according to Weir & Cockerham (1984) and a 95% confidence interval was estimated by bootstrapping over loci. FIS and FIT are the correlations between two uniting gametes relative to the subpopulation and relative to the total population, respectively, and FST is the correlation between two gametes drawn at random from each subpopulation and measures the degree of genetic differentiation of subpopulations (Nei, 1987).
Phylogeny The software PHYLIP 3.5 (Felsenstein, 1993) was used for phylogenetic analysis. Following Hillis et al.’s (1996) recommendations we used Cavalli-Sforza & Edwards’s (1967) genetic distance (other distances provided by the program GENDIST give roughly the same result). The Fitch–Margoliash method (Fitch & Margoliash, 1967) was used for tree building (procedure KITCH). Bootstrap values were calculated by generating 500 samples with the program SEQBOOT, which were used as input for the program KITCH. A consensus tree was obtained from these 500 bootstrap replicates with the program CONSENSE. The tree was drawn using TREEVIEW (Page, 1996).
Results
The locus G6pdh of L. laricina gave an unusual zymogram pattern: in addition to the bands common for other Larix taxa some additional indistinct bands were also produced. Similar problems were encountered by Cheliak & Pitel (1985) who observed “non-genetic modification” of one of the alleles after storage of the seeds. Diffuse bands were found not only on zymograms of megagametophytes, but also when analysing seedlings, embryos and needles of young plants. Because the proteins in the leaves of the young plants could not have been produced before the seed germination, we concluded that the modification of the G6pdh enzyme was not produced by seed storage. G6pdh was not used in the phylogenetic analysis. A high frequency of ‘null-alleles’ was discovered at the Skdh locus of L. occidentalis, and their frequency was estimated from the homozygote frequency assuming Hardy–Weinberg equilibrium and using the EM algorithm (Lange, 1997).
Genetic variability, Hardy–Weinberg and within-species differentiation
The mean number of alleles, the percentage of polymorphic loci and the observed and expected heterozygosities are given in Table 2 (the allozyme frequency table is available from the first author). All three quantities were lowest in species with more restricted distribution areas, such as L. lyallii and L. olgensis. A significant heterozygote deficit was found at loci G6pdh (P = 0.035), Skdh (P < 0.0001) and Idh (P < 0.0001) in L. occidentalis, Pgm-B (P = 0.0091) in L. laricina and Dia (P = 0.0025) in L. lyallii. The deficit of heterozygotes at Skdh can be explained by the incorrect scoring of null-allele heterozygotes as homozygotes of active alleles. However, heterozygote deficit at other loci cannot be attributed to the presence of null-alleles with certainty, because these were not observed among the genotyped individuals. Even if this explanation seems to be the most plausible, other explanations such as the Wahlund effect cannot a priori be ruled out. Wright’s F-statistics for the different species are given in Table 3. With FST values lower than 0.1, population differentiation was generally not very pronounced. Population differentiation was, however, more conspicuous in L. occidentalis and L. sibirica (0.100 and 0.079) than in L. laricina and L. gmelinii (0.026 and 0.021).
Phylogeny
A consensus tree was constructed from 500 Fitch–Margoliash dendrograms (Fig. 2) and the bootstrap method was used to evaluate the support for the different groupings. Because a rather limited number of loci was analysed (only 15), results should be interpreted with care even if some of the groupings are strongly supported. The American larches are clearly separated from the Eurasian ones, the Japanese larch being more closely related to the latter than to the former. Larix occidentalis and L. lyallii always grouped together, regardless of the distance or tree-building method used and the consensus tree constructed on the data, after excluding the Skdh and Idh loci, gave a similar topology to that obtained when these loci were included (results not shown). In both phylogenetic trees American larches are grouped together, separately from other species.
Discussion
Genetic variability and population differentiation
The genetic variation found in the L. laricina populations in the present study (Hexp = 0.048–0.100) was lower than those reported previously (Hexp = 0.220, Cheliak et al., 1988; Hexp = 0.151, Ying & Morgenstern, 1991; Hexp = 0.104, Liu & Knowles, 1991). This difference may be a consequence of the location of the populations at the eastern limit of the distribution range. A different picture might have been obtained had populations from the western part of the range been added to the present study. The low genetic variation observed in L. lyallii and L. olgensis could be explained by the narrowness of their ecological niche. Larix lyallii grows on poor and rocky sites, in cold, snowy and generally moist climates at high altitudes (upper subalpine to timberline ecotone) (Schmidt, 1995). Similarly, L. olgensis is found in small populations on rocky, steep, north-facing slopes. In either case, small population sizes and narrow ecological niches could easily be the cause of the low genetic diversity. Genetic differentiation was lower in L. laricina and L. gmelinii than in L. occidentalis and L. sibirica. The agreement with previous estimates is generally good (L. occidentalis, Fins & Seeb, 1986), except for L. laricina for which Cheliak et al. (1988), using populations sampled over a larger range, found a FST value of 5%. As for genetic diversity estimates, the discrepancy may simply be the consequence of the limited geographical range of our samples. Indeed Ying & Morgenstern (1991) also observed a low FST among populations restricted to New Brunswick.
Phylogeny
The main results of our study are: (i) the clear separation of American and Eurasian larches; (ii) the consistent grouping of L. laricina and L. occidentalis; and (iii) the lack of differentiation among Eurasian species. As was expected, based on recent palaeontological results (Elias et al., 1996) showing that Beringia was a tundra, and thus a priori not conducive to the migration of trees such as larch, American and European larch species are clearly genetically differentiated, based on variation at isozyme loci. Furthermore, L. lyallii and L. occidentalis form a single and separated group, regardless of the genetic distance and tree building method used. The present range of these two species is not in contact with other species and they probably had refugia south of their present range during the last glaciation (Whitlock, 1995). In fact, the divergence between these two species and Eurasian ones is of the same order of magnitude as that observed between several eastern Asia-eastern North America disjunct broad-leaved species based on isozyme loci (table 6 in Qiu et al., 1995). The corresponding divergence time, using Nei’s method (Nei, 1987, p. 237), is ≈ 3–4 Myr. The differentiation between L. laricina and Eurasian species is somewhat less pronounced, which was expected, considering the present distribution range of the species. In that case the divergence could have occurred during the Pleistocene. An even less pronounced difference would probably have been observed had samples from Alaska been included in the study. These results might seem to conflict with the phylogeny based on bract morphology, for which L. kaempferi clusters with L. lyallii and L. occidentalis, whereas L. laricina clusters with other Eurasian species (LePage & Basinger, 1995). However, the two can easily be reconciled, if one assumes that allele frequencies at isozyme loci reflect recent demographic events, whereas bract morphology reflects more ancient history. Furthermore, LePage & Basinger (1995) observe that short-bract species commonly occupy low-altitude habitats, whereas long-bract species are found at high altitudes. Bract morphology would thus be related to fitness and evolve slowly. The fossil record from the high Arctic indicates that larches could have dispersed between North America and northern Eurasia along the Beringia corridor some 40 Myr ago (LePage & Basinger, 1995). Our data do not allow us to draw any conclusions on the direction of dispersal, although migration from Eurasia into North America seems more likely, because L. griffithiana is apparently the oldest of the extant taxa of Larix (Sukaczew, 1924). No strong inference can be made within Eurasian species, the differentiation between them being too weak. We note, however, that L. kaempferi differs from others, notably L. decidua. Although they used 66 RFLP probe–enzyme combinations, Qian et al. (1995) observed no difference between L. decidua, L. kaempferi and L. gmelinii, but they detected differences between this group and L. sibirica. However, the overall differentiation was very small, the maximum number of nucleotide substitutions per site (d) being only 0.0096. In larch, mitochondrial DNA is maternally inherited, and thus dispersed only through seeds (Deverno et al., 1993; D. Prat, pers. comm.). A more pronounced genetic differentiation pattern is therefore expected. In a follow-up study the same populations, together with additional ones, will be analysed at mitochondrial DNA markers.
References
Bobrov, E. G. (1972). History and Systematics of Larix. Nauka, Leningrad (in Russian).
Cavalli-Sforza, L. L. and Edwards, A. W. F. (1967). Phylogenetic analysis: models and estimation procedures. Am J Hum Genet, 19: 223–257.
Cheliak, W. M. and Pitel, J. A. (1985). Inheritance and linkage of allozymes in Larix laricina. Silvae Genet, 34: 142–148.
Cheliak, W. M., Wang, J. and Pitel, J. A. (1988). Population structure and genetic diversity in tamarack, L. laricina (Du Roi) K. Koch. Can J Forest Res, 18: 1318–1324.
Deverno, L. L., Charest, P. J. and Bonen, L. (1993). Inheritance of mitochondrial DNA in the conifer Larix. Theor Appl Genet, 86: 383–388.
Elias, S. A., Short, S. K., Nelson, C. H. and Birks, H. H. (1996). Life and time of the Bering land bridge. Nature. 382: 60–63.
Felsenstein, J. (1993). Phylip (Phylogeny inference package), version 3.5p. Distributed by the author, Department of Genetics, University of Washington, Seattle.
Fins, L. and Seeb, L. W. (1986). Genetic variation in allozymes of western larch. Can J Forest Res, 16: 1013–1018.
Fisher, R. A. (1954). Statistical Methods for Research Workers 12th edn. Oliver & Boyd, Edinburgh.
Fitch, W. M. and Margoliash, E. (1967). Construction of phylogenetic trees. A method based on mutation distances as estimated from cytochrome c sequences is of general applicability. Science. 155: 279–284.
Goudet, J. (1995). Fstat v-1.2: a computer program to calculate F-statistics. J Hered, 86: 485–486.
Guo, S. W. and Thompson, E. A. (1992). Performing the exact test of Hardy–Weinberg proportions for multiple alleles. Biometrics. 48: 361–372.
Harris, H. and Hopkinson, D. A. (1976). Handbook of Enzyme Electrophoresis North-Holland Publishers, Amsterdam.
Hasebe, M., Ando, T. and Iwatsuki, K. (1998). Intrageneric relationships of maple trees based on chloroplast DNA restriction fragment length polymorphisms. J Plant Res, 111: 441–451.
Hillis, D. M., Moritz, C. and Mable, B. K. (1996). Molecular Systematics 2nd edn. Sinauer Associates, Sunderland, MA.
Hultén, E. (1937). Outline of the History of Arctic and Boreal Biota During the Quaternary Period Bokförlags Aktiebolaget Thule, Stockholm.
Kisanuki, H., Kurahashi, A., Kato, H., Terauchi, R., Kawano, S., Ide, Y. and Watanabe, S. (1995). Interspecific relationship of genus Larix inferred from the RFLPs of chloroplast DNA. J Jap For Soc, 77: 83–85.
Lange, K. (1997). Mathematical and Statistical Methods for Genetic Analysis. Springer, New York.
Lepage, B. A. and Basinger, J. F. (1995). The evolutionary history of the genus Larix (Pinaceae). In: Ecology and Management of Larix Forests: A Look Ahead Proceedings of an international symposium, 5–9 October 1992, pp. 19–29. Whitefish, MT, U.S.A..
Lewandowski, A. and Mejnartowicz, L. (1990). Inheritance of allozymes in Larix decidua Mill. Silvae Genet, 39: 184–188.
Liu, Z. and Knowles, P. (1991). Patterns of allozyme variation in tamarack (Larix laricina) from northern Ontario. Can J Bot, 69: 2469–2474.
Nei, M. (1987). Molecular Evolutionary Genetics, Columbia University Press, New York.
Page, R. D. M. (1996). Treeview: an application to display phylogenetic trees on personal computers. Comp Appl Biosci, 12: 357–358.
Qian, T., Ennos, R. A. and Helgason, T. (1995). Genetic relationships among larch species based on analysis of restriction fragment variation for chloroplast DNA. Can J Forest Res, 25: 1197–1202.
Qiu, Y. -L., Parks, C. R. and Chase, M. W. (1995). Molecular divergence in the eastern Asia–eastern North America disjunct section Rytidospermum of Magnolia (Magnoliaceae). Am J Bot, 82: 1589–1598.
Raymond, M. and Rousset, F. (1995). Genepop (version 1.2): a population genetics software for exact tests and ecumenicism. J Hered, 86: 248–249.
Rousset, F. and Raymond, M. (1995). Testing heterozygote excess and deficiency. Genetics. 140: 1413–1419.
Schmidt, W. C. (1995). Around the world with Larix: an introduction. In: Ecology and Management of Larix Forests: A Look Ahead Proceedings of an international symposium, 5–9 October 1992, pp. 6–17. Whitefish, MT, U.S.A.
Semerikov, V. L. and Matveev, A. V. (1995). Investigation of genetic variation of allozyme loci in Siberian larch. Russian J Genet, 31: 944–949.
Semerikov, V. L., Semerikov, L. F. and Lascoux, M. (1998). Intra- and interspecific allozyme variability in Eurasian Larix Mill. species. Heredity. 82: 193–204.
Shurkhal, A. V., Podogas, A. V. and Zhivotovsky, L. A. (1992). Allozyme differentiation in the genus Pinus. Silvae Genet, 41: 105–109.
Sukaczew, V. N. (1924). On the Evolutionary History of Larix. People’s Commissariat of Agriculture, Leningrad (in Russian).
Weir, B. S. and Cockerham, C. C. (1984). Estimating F -statistics for the analysis of population structure. Evolution. 38: 1358–1370.
Wen, J., Shi, S., Jansen, R. K. and Zimmer, E. A. (1998). Phylogeny and biogeography of Aralia sect. Aralia (Araliaceae). Am J Bot, 85: 866–875.
Whitlock, C. (1995). The history of Larix occidentalis during the last 20,000 years of environmental changes. In: Ecology and Management of Larix Forests: A Look Ahead Proceedings of an international symposium, 5–9 October, 1992, pp. 83–90. Whitefish, MT, U.S.A..
Ying, L. and Morgenstern, E. K. (1991). The population structure of Larix laricina in New Brunswick, Canada. Silvae Genet, 40: 180–184.
Acknowledgements
The first author is very grateful to all the people that helped him by collecting seed material in Eurasia. Seeds of L. occidentalis and L. lyallii were supplied by Dr B. C. Jaquish and B. Wilson from the British Columbia Forest Service. The L. laricina seeds were provided by Dr Y. S. Park from New Brunswick, Canadian Forest Service and Prof. F. Telewsky, Beal Botanical Garden, Michigan State University. This research was supported by the Russian Scientific Foundation (grant 94-04-1282-a). V. Semerikov was supported by the Swedish Institute and the Swedish Royal Academy of Sciences.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Semerikov, V., Lascoux, M. Genetic relationship among Eurasian and American Larix species based on allozymes. Heredity 83, 62–70 (1999). https://doi.org/10.1038/sj.hdy.6885310
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.hdy.6885310
Keywords
This article is cited by
-
Larix species range dynamics in Siberia since the Last Glacial captured from sedimentary ancient DNA
Communications Biology (2022)
-
Range-wide genetic diversity in natural populations of Larix principis-rupprechtii Mayr.
Journal of Forestry Research (2021)
-
Does larch arabinogalactan enhance immune function? A review of mechanistic and clinical trials
Nutrition & Metabolism (2016)
-
Rates of spontaneous hybridization and hybrid recruitment in co-existing exotic and native mature larch populations
Tree Genetics & Genomes (2014)
-
The variability of different larch clone provenances on the response to the attack by its main pests and fungal diseases
Trees (2013)