Abstract
Islands are choice settings for experimental studies of vector control strategies based on transgenic insects. Before considering this approach, knowledge of the population structure of the vector is essential. Genetic variation at 12 microsatellite loci was therefore studied in samples of the malaria vector Anopheles gambiae s.s., collected from six localities of São Tomé island (West Africa). The objectives were (i) to assess the demographic stability and effective population size of A. gambiae from these sites, (ii) to determine population differentiation and (iii) to relate the observed patterns of population structure with geographic, ecological and historical aspects of the vector on the island. Significant population differentiation, revealed by FST and RST statistics, was found between the southernmost site, Porto Alegre, and northern localities. The observed patterns of population substructure are probably a result of restrictions to gene flow in the less inhabited, more densely forested and mountainous south. In all localities surveyed, A. gambiae appeared to be experiencing a demographic expansion, consistent with a relatively recent (ca. 500 years) founder effect. The results are discussed with respect to current and future prospects of malaria vector control.
Similar content being viewed by others
Introduction
Studies on the genetic structure of malaria vector populations can be used to infer the likely success of vector control strategies. They can highlight issues such as the impact of conventional methods (eg, insecticide spraying) in reducing vector abundance, the reintroduction of vectors into formerly controlled areas, the spread of insecticide-resistance genes or the control of vectors by means of transgenic technology (Collins et al, 2000). The latter is a promising novel control strategy, although still subject to intense debate (Curtis et al, 1999). Should transgenic-based control be attempted, a natural first step would be to test its efficacy on islands, where confounding effects such as migration are expected to be lesser problems than on the continent.
The islands of the Gulf of Guinea, in particular São Tomé, are potential sites that might benefit from malaria control strategies based on transgenic vectors. In São Tomé, malaria is responsible for most hospital admissions and is the leading cause of child mortality. The only vector present is the forest cytoform of Anopheles gambiae s.s. (Pinto et al, 2000). Its origin on the island is likely to be contemporary with the first human settlements in the 15th century (Baptista, 1996). The vector is largely confined to coastal areas, being virtually absent at altitudes above 200 m. It is intimately associated with humans, feeding predominately on them or their domestic animals, and breeding in sites located close to houses (Pinto et al, 2000; Sousa et al, 2001). In 1980–1982, the island was subjected to a malaria eradication programme based on intensive intradomicilliary DDT spraying and mass drug (chloroquine) distribution (Ceita, 1986). During this period, both malaria prevalence and A. gambiae densities inside houses were dramatically reduced. The subsequent disruption of the programme led to a severe epidemic with many fatalities and present malariological parameters have reached or even exceeded preintervention levels (Pinto et al, 2000).
In a previous study, populations of A. gambiae from the islands of São Tomé and Príncipe did not show evidence of departures from mutation-drift equilibrium (MDE), in spite of their relatively recent founding and the possible impact of vector control in the 1980s (Pinto et al, 2002). Furthermore, despite the island's relatively small size, there was significant genetic differentiation between two samples from extreme locations within São Tomé. Whether this was due to physical barriers to gene flow or was a consequence of geographic distance between two extremes of a continuous vector population distribution remained unknown.
The aims of the present study were therefore (i) to determine whether the DDT spraying intervention of the 1980s has had an impact on the effective population size of the vector, (ii) to assess the present population stability and effective population size, in order to infer whether there are likely to be optimal periods for transgenic releases and, (iii) to determine patterns of population substructure, in order to infer possible restrictions to gene flow that may interfere with the spread of refractory genes. Results are discussed in relation to geographic, ecological and historical aspects of the vector populations, as well as to current and future prospects of vector control in the island.
Material and methods
Collection sites
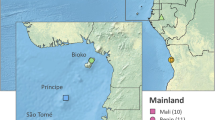
The island of São Tomé (836 km2) is part of a volcanic chain located in the Gulf of Guinea, ca. 240 km off the coast of Gabon (Figure 1). It has a mountainous topography, particularly in the west-central and southern parts, where a number of peaks rise over 1500 m (Denny and Ray, 1989). The southern part of the island is heavily forested, while the northern part is flatter and humid-savannah-like landscapes prevail.
The island of São Tomé. Geographic location, topography, demography and collection sites. (1) Neves: capital of Lembá district, where the Port for fuel importation is located. (2) Riboque: one of the main suburbs of São Tomé city, the capital of the Rep. Dem. São Tomé and Príncipe. (3) Bobo Forro: peripheral village of São Tomé city, in the most populated Água Grande district. (4) Ribeira Afonso: main town of Cantagalo district. (5) Angolares: capital of Caué, the largest but least populated district of the island. (6) Porto Alegre: the most extreme southern human settlement, a former colonial farm (Roça). With the exception of Porto Alegre, where most houses are ground-level and brick-built, wood-houses built on stilts up to 2 m prevail in all other collection sites.
The climate is equatorial with an average annual temperature of 25°C and average relative humidity of 80%. Average annual rainfall varies between 500–2000 mm3 in the north and 3000–7000 mm3 in the south (Denny and Ray, 1989). There are two dry seasons, gravana (June–August) and gravanito (January), when rainfall is reduced but rarely absent.
The human population is unevenly distributed throughout the island with over 60% of the 140 000 inhabitants living within 10 km of the capital city, São Tomé (Figure 1). Most of the remaining population lives to the northeast of the capital in small villages along main roads close to the coast (Pinto et al, 2000). In the mountainous and forested south and southeast, human settlements are scarce or absent, and the southwestern part of the island is almost completely uninhabited.
Mosquitoes were collected by human baited landing catches between March and May 1998, in six localities of São Tomé (Figure 1). Samples of 60 females each were also collected in May 1997 and May 1999 from Riboque using CDC light traps. Both methods are considered comparable and equally representative for A. gambiae sampling (Davies et al, 1995). Female A. gambiae were individually stored in silica-gel-filled tubes and kept at −20°C until further processing.
Species identification and microsatellite genotyping
The DNA from individual A. gambiae was extracted by a phenol–chloroform protocol (Ballinger-Crabtree et al, 1992) and species identification was made by PCR (Scott et al, 1993). Samples were further identified as belonging to the M/S molecular form according to the PCR protocol described in Favia et al (2001). Microsatellite DNA analysis was performed for 11 dinucleotide loci (GT repeats) and one trinucleotide locus (AGC repeat). Details on the cytological location for each locus are given in Table 1. Genotyping procedures with fluorescence technology using an ABI 373 automatic sequencer (Applied Biosystems) were as described in Donnelly et al (1999).
Data analysis
Estimates of expected heterozygosity were made using Nei's unbiased estimator (Nei, 1987). Differences in heterozygosity among samples and mean number of alleles were tested, pairwise, by Wilcoxon signed-ranks tests and overall by the Friedman test. Tests of deviation from Hardy–Weinberg proportions at each locus and of linkage disequilibrium between pairs of loci were performed using GENEPOP v3.3. (Raymond and Rousset, 1995).
Two independent estimates of effective population size (Ne) were produced. Current Ne was estimated for Riboque, based on the temporal variation in allele frequencies for three samples from May 1997, 1998 and 1999, respectively. The standardised variance of allele frequency change, F, was calculated for 1997–1998, 1998–1999 and the two-year interval 1997–1999, according to Pollak (1983), using equation (9) in Waples (1989). Estimates of Ne assumed that individuals were sampled without replacement prior to reproduction (Waples, 1989). A conservative number of 12 generations per year was used as in previous studies (Taylor et al, 1993; Lehmann et al, 1998). Calculations were made using programs written in the SAS language (SAS Institute, 1990).
Long-term Ne was calculated based on the expected heterozygosity at each microsatellite locus, for each locality. Estimates were made using equations described in Nei (1987) based on two mutation models, the stepwise mutation model (SMM) and the infinite alleles model (IAM). These are at the extreme ends of the spectrum of mutation models and therefore provide robust range estimates of long-term Ne (Lehmann et al, 1998). As in previous studies, average mutation rate (μ) was assumed to be 10−4 (Lehmann et al, 1998). For X-linked loci, Ne values were adjusted by a factor of 4/3 based on an assumption of a 1:1 sex ratio.
Two methods were used to assess deviations from MDE. Cornuet and Luikart's (1996) heterozygosity tests compare two estimates of expected heterozygosity, one based on allele frequencies (He) and another based on the number of alleles and sample size (Heq). In a population at MDE, both estimates should be equal (ie, He=Heq). If a population experiences a bottleneck, rare alleles will be lost and therefore Heq will decrease faster than He (ie, He>Heq). This apparent excess of heterozygosity is an indicator of a recent bottleneck event, whereas the converse (ie, He<Heq) may indicate an expansion event. Wilcoxon signed-ranks tests were used to determine if there was a significant number of loci in which He>Heq. Estimates of Heq were calculated under three mutation models, the SMM, the IAM and an intermediate two-phased model (TPM) with fractions of mutations greater than one repeat of 10, 20 and 30%. Tests were performed using BOTTLENECK 1.2.02 (Cornuet and Luikart, 1996).
Kimmel et al's (1998) imbalance index (β) is the quotient of two estimates of the genetic diversity parameter θ (for diploid loci: θ=4Neμ). One estimate is obtained from the variance of allele length (in repeat numbers) at each locus (θV) and the other from expected homozygosity (θP0). At MDE and under the SMM, θV=θP0; so β=θV/θP0=1. Simulations have shown that if β<1, this is an indication of recent population expansion from an original MDE (Kimmel et al, 1998). If β>1, expansion has followed a previous bottleneck event. Two estimators of mean β over loci, which yield normal-like distributions, were calculated as described in King et al (2000). The first is the log ratio of means, given by

and the second is based on the mean of log ratios,

where L is number of loci. Bootstrapped 95% confidence intervals were used to evaluate if the mean over loci β were significantly different from one.
Genetic differentiation was determined by the fixation index FST (Wright, 1978) and the analogous estimator for microsatellite data RST (Slatkin, 1995). Estimates of FST were calculated according to Weir and Cockerham (1984) using FSTAT v2.9.3 (Goudet, 1995). Estimates of RST were produced using RSTCALC software, according to Goodman (1997). The significance of FST and RST estimates was assessed by genotypic permutation tests among samples.
Isolation by distance was investigated by the regression of FST/(1−FST) and RST/(1−RST) on geographic distance (Rousset, 1997). Given the narrow distribution of the vector, a one-dimensional habitat was assumed and map-measured distances along coastal roads were used in the regression analyses (Rousset, 1997). Significance of the Spearman's rank correlation coefficient between variables (differentiation vs distance) was determined by Mantel tests. Calculations were carried out using GENEPOP v3.3 (Raymond and Rousset, 1995).
Whenever multiple tests were performed, the nominal significance level (P<0.05) was adjusted using the sequential Bonferroni procedure (Holm, 1979).
Results
Species identification and genetic diversity
All 448 specimens analysed belonged to the M molecular form of A. gambiae s.s. Microsatellite genetic diversity was moderate to high (Table 1). Overall, allele distribution was similar among localities (data available from authors on request). Numbers of alleles per locus varied from two to 11 (mean 6.3, SD 3.2) and heterozygosity varied from 0.389 to 0.805 (mean 0.623, SD 0.126). There were no significant differences in either variable across samples (Friedman test: mean number of alleles, P=0.300; heterozygosity, P=0.070). There were no significant pairwise differences in the mean number of alleles among samples. However, four of the five pairwise comparisons of mean heterozygosity that included the sample from the southernmost site, Porto Alegre, were significant prior to adjustment for multiple tests (Wilcoxon signed-ranks tests: 0.010<P<0.023). The only nonsignificant comparison was that between Porto Alegre and the next southernmost sample, Angolares (Figure 1).
Within-sample departures from Hardy–Weinberg expectations, as a result of heterozygote deficit, appeared to be clustered at loci AG2H147 and AG3H750 (Table 1). There was, however, no particular association of these two loci with any others in pairwise tests of linkage disequilibrium which were all insignificant after the sequential Bonferroni procedure was applied (16 tests out of 396 significant at the 0.05 level). The absence of genome-wide deviations from Hardy–Weinberg equilibrium and of linkage disequilibrium support that samples were drawn from homogenous and randomly mating populations.
Effective population size and MDE
Mean estimates (1-year) of current Ne in Riboque ranged from 1078 (349–∞), in 1998–1999, to infinity (1049–∞), in 1997–1998. For the 1997–1999 2-year estimate, a bound 95% CI was obtained (mean: 1457; 95% CI: 559–13 677). In single-locus estimates, the upper limit of 95% CIs was infinity in all but two cases (AG2H147: 86–2137; AG3H128: 43–5075) in 2-year estimates.
Estimates of long-term Ne were also in the order of thousands for both mutation models, values not consistent with recent bottleneck events (Table 2). No significant differences in Ne were found across localities (Friedman test: P=0.070, in both estimates). Depending on mutation model, two or three pairwise comparisons, which included Porto Alegre, were significant prior to adjustment for multiple tests (Wilcoxon signed-ranks tests: SMM: two out of five, 0.023<P<0.034; IAM: three out of five, 0.028<P<0.041). Furthermore, overlapping 95% confidence intervals were obtained among all estimates, indicating that long-term Ne is comparable among samples and that they appear to be subjected to similar amounts of genetic drift.
The results obtained from the heterozygosity tests (Cornuet and Luikart, 1996) are shown in Table 2. Significant numbers of loci with an apparent heterozygote excess were detected for the less suitable mutation model, IAM, in five out of the six samples, and for TPM-70% (two samples) and TPM-80% (one sample). Luikart and Cornuet (1998) point out that these tests under the IAM may wrongly detect heterozygosity excess in nonbottlenecked populations when microsatellite data are used. They suggest using the strict SMM or a TPM model with 5–10% multistep changes with such data. Under these models, no significant proportion of loci with heterozygote excess was found in any sample (Table 2).
Contrasting results were obtained by the two estimators of the imbalance index, as shown in Table 3. While β2 estimates, varying between 0.92 and 1.42, do not support population perturbation, β1 estimates were significantly greater than 1 for all samples, suggesting that A. gambiae is experiencing a demographic expansion following a bottleneck or founder effect (Kimmel et al, 1998). Notwithstanding overlapping confidence intervals, there was an apparent trend for estimates of β1 to increase towards the south of the island, varying between 2.42 and 2.61 in northern localities, to 3.09 and 4.32 in the southern Angolares and Porto Alegre.
Genetic differentiation
Pairwise over loci estimates of FST and RST showed comparable patterns of population differentiation (Table 4). Significant FST values were obtained for all comparisons involving Porto Alegre and between Bobo Forro and Angolares (Figure 1). Significant RST values were recorded in all comparisons with Porto Alegre, with the exception of the nearest site, Angolares. Patterns of population differentiation did not differ when loci AG2H147 and AG2H175, which exhibited highest heterozygote deficits, were excluded from the analysis, demonstrating that deviations from HWE had little effect on the FST/RST estimates (Table 4).
A significant positive correlation was found between both FST/(1−FST) and RST/(1−RST) and geographic distance (Mantel test; P=0.038 and 0.012, respectively) (Figure 2a). However, pairwise estimates of FST/(1−FST) with Porto Alegre were higher than expected by the regression with distance in four out of five cases and in three out of five cases for RST/(1−RST). When Porto Alegre was excluded from the analysis, no significant correlation was found for the remaining pairs (Mantel test; P=0.359 and 0.284, respectively) (Figure 2b).
Discussion
This study provided evidences that the vector control campaign of the 1980s did not drastically reduce the effective population size of A. gambiae in the island of São Tomé. Populations, however, are likely to be expanding, possibly reflecting a relatively recent founder effect associated with the human colonization of the island in the 15th century. The genetic differentiation observed between northern and southern localities may be a consequence of north–south environmental heterogeneity, coupled with the synanthropic nature of this vector. In the south, human settlements are scarce and relatively isolated from those in the north.
Estimates of current Ne calculated for Riboque were in the order of thousands. These were in the range of those obtained by microsatellite data for A. gambiae from Western and Eastern Kenya (2455–103 317 and 1671–24 203, respectively) or for A. arabiensis from Senegal (409–2768) (Lehmann et al, 1998; Simard et al, 2000). In Mali, estimates of Ne around 2000 for A. arabiensis were obtained by inversion chromosome frequencies and between 922 and 1922 for A. gambiae using mark-release-recapture methods (Taylor et al, 1993, 2001). Given the equatorial climatic conditions of São Tomé, 12 generations per year is probably a conservative number; hence presented values may be an underestimate of the actual current Ne. Such values are not consistent with strong reductions in effective size occurring between 1997 and 1999. It is therefore unlikely that, in this island, A. gambiae experiences seasonal episodes of drastic reduction of effective population size.
Although robust, inferences on population stability based on current Ne are limited to the period within sampling time points. Estimates of long-term Ne, based on expected heterozygosity assuming equilibrium, however, reflect the long-term history of a species or population (Waples, 1991; Lehmann et al, 1998). Regardless of mutation model, large estimates of long-term Ne were obtained in all locations analysed and are not consistent with a bottleneck having occurred during the 1980's DDT intervention. Estimates are within the range of those found in A. gambiae mainland populations. Long-term Ne values (IAM–SMM) around 6500–21 000 (Senegambia), 7500–23 000 (Western Kenya), 8500–24 000 (Gabon) and 11 500–49 000 (Cameroon) are derived by reanalysing microsatellite data in previous publications (Lehmann et al, 1996, 1998; Pinto et al, 2002; Wondji et al, 2002). In addition, present estimates of long-term Ne may also be underestimated. This could be due to: (i) the conservatively high mutation rate used in calculations, since microsatellite mutation rates in some insect species may be lower than expected (Schug et al, 1997), and (ii) size constraints at microsatellite loci (Garza et al, 1995; Estoup and Cornuet, 1999), that could disrupt the correlation between allele size variation/heterozygosity and Ne.
Should a recent bottleneck have occurred, it would have been detected by heterozygosity tests which are sensitive to such events for a period of less than 4Ne generations but that can be as short as 0.2Ne generations (Cornuet and Luikart, 1996; Luikart and Cornuet, 1998). In the 17 years from the start of the DDT intervention, no more than 400 generations will have passed, even assuming 23 generations/year, an estimate for A. gambiae under optimal laboratory conditions (Lehmann et al, 1998). Heterozygosity tests under the most appropriate mutation models for microsatellite data (SMM and TPM 10%), however, showed no evidence of departures from MDE.
While values of β2 also provided no evidence to reject MDE, values of β1 suggest that populations are expanding. Estimator β2 tends to be seriously downward biased, being less sensitive to situations where expansion is preceded by a bottleneck or founder effect (Kimmel et al, 1998). This is more likely to have been the case for A. gambiae in São Tomé. Since the power of β1 and β2 increases to values above 0.5 after more than 625 generations following expansion (King et al, 2000), it is unlikely that the signature of expansion captured by β1 is associated with a bottleneck occurring in the 1980s. Rather, it may be the consequence of a founder effect associated with colonization of the island in the 15th century. The increasing β1 values towards south may also reflect more recent founder events or range expansions, since increased establishment of farms in the south occurred only in the 19th century (Tenreiro, 1961). In continental A. gambiae populations, β values significantly different from 1 were consistent with a population expansion scenario, possibly associated with the agrarian revolution that occurred in Africa ca. 10 000 years ago (Coluzzi, 1999; Donnelly et al, 2001).
Population substructure was found between northern and the southernmost sites, particularly Porto Alegre. Both FST and RST estimators showed significant differentiation between this locality and the remaining ones, suggesting a degree of genetic isolation between them. Little genetic differentiation was observed among northern localities of the island. The genetic uniformity in northern samples suggests that they were drawn from the same panmictic unit, implying a continuous distribution of A. gambiae along the northern coast. By itself, geographic distance may not be sufficient to explain the observed patterns of differentiation. The significant correlation observed under an isolation by distance model was lost when Porto Alegre was excluded from the analysis. Although removing data points reduces the power of the tests, the observed genetic differentiation of Porto Alegre was consistently higher than expected. In the southern parts of the island, the scarcity of human settlements coupled with the mountainous terrain and dense forest may act as a barrier to gene flow.
The observed patterns of population structure of A. gambiae in the island of São Tomé beg a number of questions related to its control. Conventional vector control measures directed solely to certain aspects of its lifecycle, such as indoor insecticide spraying, may have little long-term effect. The apparently low impact of such a control strategy on the genetic structure of A. gambiae may be related to the vector's exophagic and exophilic preferences in this island (Sousa et al, 2001). The campaign was probably successful in reducing human–vector contact or vector longevity and thus transmission. However, once the programme was disrupted, increased human–vector contact from a still large vector population, combined with loss of acquired immunity and appearance of chloroquine-resistant Plasmodium falciparum strains, led to the subsequent epidemic (Ceita, 1986).
High estimates of current Ne, suggesting little seasonal fluctuations in effective size, may pose difficulties in assessing the timing of transgenic insect releases. In such scenarios, the use of conventional methods aimed at reducing vector populations may be required prior to releases. Restrictions to gene flow may also pose obstacles to malaria control with transgenic insects. Releases undertaken at a limited number of sites might be sufficient to transform northern vector populations, but these might not affect those in the south. On the other hand, an isolated population such as Porto Alegre, located at the extreme end of the vector distribution on the island, may be regarded as a potential site for experimental transgenic releases, a crucial first step prior to the implementation of any large-scale releases.
References
Ballinger-Crabtree ME, Black IV WC, Miller BR (1992). Use of genetic polymorphisms by the random-amplified polymorphic DNA polymerase chain reaction (RAPD-PCR) for differentiation and identification of Aedes aegypti subspecies and populations. Am J Trop Med Hyg 47: 893–901.
Baptista JL (1996). Subsídio para a história do paludismo em S. Tomé. Acta Med Port 9: 259–265.
Ceita JGV (1986). Malaria in São Tomé and Príncipe. In: Buck AA (ed) Proceedings of the Conference on Malaria in Africa. American Institute of Biological Sciences/USAID: Washington, DC. pp 142–155.
Collins FH, Kamau L, Ranson HA, Vulule JM (2000). Molecular entomology and prospects for malaria control. Bull World Health Org 78: 1412–1423.
Coluzzi M (1999). The clay feet of the malaria giant and its African roots: hypothesis and inferences about origin, spread and control of Plasmodium falciparum. Parassitologia 41: 277–283.
Coluzzi M, Petrarca V, Di Deco MA (1985). Chromosomal inversion intergradation and incipient speciation in Anopheles gambiae. Boll Zool 52: 45–63.
Cornuet J-M, Luikart G (1996). Description and power analysis of two tests for detecting recent population bottlenecks from allele frequency data. Genetics 144: 2001–2014.
Curtis CF, Pates HV, Takken W, Maxwell CA, Myamba J, Priestman A et al (1999). Biological problems with the replacement of a vector population by Plasmodium-refractory mosquitoes. Parassitologia 41: 479–481.
Davies JC, Hall T, Chee EM, Majala A, Minjas J, Shiff CJ (1995). Comparison of sampling anopheline mosquitoes by light-trap and human-bait collections indoors at Bagamoyo, Tanzania. Med Vet Entomol 9: 249–255.
Denny LM, Ray DI (1989). São Tomé and Príncipe. Economic, Politics and Society. Printer Publishers: London.
Donnelly MJ, Cuamba N, Charlwood JD, Collins FH, Townson H (1999). Population structure in the malaria vector, Anopheles arabiensis Patton, in East Africa. Heredity 83: 408–417.
Donnelly MJ, Licht M, Lehmann T (2001). Evidence for recent population expansion in the evolutionary history of the malaria vectors Anopheles arabiensis and Anopheles gambiae. Mol Biol Evol 18: 1353–1364.
Estoup A, Cornuet J-M (1999). Microsatellite evolution: inferences from population data. In: Goldstein DB, Schlötterer C (eds) Microsatellites. Evolution and Applications, Oxford University Press: New York. pp 49–65.
Favia G, Lanfrancotti A, Spanos L, Sidén-Kiamos I, Louis C (2001). Molecular characterisation of ribosomal DNA (rDNA) polymorphisms discriminating chromosomal forms of Anopheles gambiae s.s. Insect Mol Biol 10: 19–23.
Garza JC, Slatkin M, Freimer NB (1995). Microsatellite allele frequencies in humans and chimpanzees, with implications for constraints on allele size. Mol Biol Evol 12: 594–603.
Goodman SJ (1997). RST Calc: a collection of computer programs for calculating estimates of genetic differentiation from microsatellite data and determining their significance. Mol Ecol 6: 881–885.
Goudet J (1995). FSTAT (version 1.2): a computer software to calculate F-statistics. J Hered 86: 485–486.
Holm S (1979). A simple sequentially rejective multiple test procedure. Scand J Stat 6: 65–70.
Kimmel M, Chakraborty R, King JP, Bamshad M, Watkins WS, Jorde LB (1998). Signatures of population expansion in microsatellite repeat data. Genetics 148: 1921–1930.
King JP, Kimmel M, Chakraborty R (2000). A power analysis of microsatellite-based statistics for inferring past population growth. Mol Biol Evol 17: 1859–1868.
Lehmann T, Hawley WA, Grebert H, Collins FH (1998). The effective population size of Anopheles gambiae in Kenya: implications for population structure. Mol Biol Evol 15: 264–276.
Lehmann T, Hawley WA, Kamau L, Fontenille D, Simard F, Collins FH (1996). Genetic differentiation of Anopheles gambiae populations from East and West Africa: comparison of microsatellite and allozyme loci. Heredity 77: 192–208.
Luikart G, Cornuet J-M (1998). Empirical evaluation of a test for identifying recently bottlenecked populations from allele frequency data. Cons Biol 12: 228–237.
Nei M (1987). Molecular Evolutionary Genetics. Columbia University Press: New York.
Pinto J, Donnelly MJ, Sousa CA, Gil V, Ferreira C, Elissa N et al (2002). Genetic structure of Anopheles gambiae (Diptera: Culicidae) in São Tomé and Príncipe (West Africa): implications for malaria control. Mol Ecol 11: 2183–2187.
Pinto J, Sousa CA, Gil V, Ferreira C, Gonçalves L, Lopes D et al (2000). Malaria in São Tomé and Príncipe: parasite prevalences and vector densities. Acta Trop 76: 185–193.
Pollak E (1983). A new method for estimating the effective population size from allele frequency changes. Genetics 104: 531–548.
Raymond M, Rousset F (1995). GENEPOP (version 1.2): population genetics software for exact tests and ecumenicism. J Hered 86: 248–249.
Rousset F (1997). Genetic differentiation and estimation of gene flow from F-statistics under isolation by distance. Genetics 145: 1219–1228.
SAS Institute (1990). SAS Language: References. Version. SAS: Cary, NC.
Schug MD, Mackay TF, Aquadro CF (1997). Low mutation rates of microsatellite loci in Drosophila melanogaster. Nat Genet 15: 99–102.
Scott JA, Brogdon WG, Collins FH (1993). Identification of single specimens of the Anopheles gambiae complex by the polymerase chain reaction. Am J Trop Med Hyg 49: 520–529.
Slatkin M (1995). A measure of population subdivision based on microsatellite allele frequencies. Genetics 139: 457–462.
Simard F, Lehmann T, Lemasson J-J, Diatta M, Fontenille D (2000). Persistence of Anopheles arabiensis during the severe dry season conditions in Senegal: an indirect approach using microsatellite loci. Insect Mol Biol 9: 467–479.
Sousa CA, Pinto J, Almeida APG, Ferreira C, do Rosário VE, Charlwood JD (2001). Dogs as favoured hosts of Anopheles gambiae sensu stricto (Diptera Culicidae) of São Tomé, West Africa. J Med Entomol 38: 122–125.
Taylor CE, Touré YT, Carnham J, Norris DE, Diolo G, Traoré SF et al (2001). Gene flow among populations of the malaria vector Anopheles gambiae, in Mali, West Africa. Genetics 157: 743–750.
Taylor CE, Touré YT, Coluzzi M, Petrarca V (1993). Effective population size of Anopheles arabiensis during the dry season in West Africa. Med Vet Entomol 7: 351–357.
Tenreiro F (1961). A ilha de São Tomé. Memórias da Junta de Investigação do Ultramar 2nd Ser. No. 4: Lisbon.
Waples RS (1989). A generalized approach for estimating effective population size from temporal changes in alleles frequency. Genetics 121: 379–391.
Waples RS (1991). Genetic methods for estimating effective population size of cetacean populations. In: Hoelzel AR (ed) Genetic Ecology of Whales and Dolphins. International Whaling Commission No. 13: Cambridge. pp 279–300.
Weir BS, Cockerham CC (1984). Estimating F-statistics for the analysis of population structure. Evolution 38: 1358–1370.
Wondji C, Simard F, Fontenille D (2002). Evidence for genetic differentiation between the molecular forms M and S within the forest chromosomal form of Anopheles gambiae in an area of sympatry. Insect Mol Biol 11: 11–19.
Wright S (1978). Evolution and the Genetics of Populations. Vol.1: Variability Among and Within Populations, 2nd edn. University of Chicago Press: Chicago.
Zheng L, Benedict MQ, Cornel AJ, Collins FH, Kafatos FC (1996). An integrated genetic map of the African human malaria vector mosquito, Anopheles gambiae. Genetics 143: 941–952.
Acknowledgements
We thank the people of São Tomé and Príncipe for embracing the study, the entomology team of CNE for support in the field collections and the Ministry of Health (RD STP) for logistics. We are indebted to H Townson (LSTM, Liverpool, UK) for technical training and to two anonymous referees for useful comments on an earlier version of the manuscript. This work was financed by the INCO-DC/European Union projects (IC18CT960030 and IC18CT970244). J Pinto and JD Charlwood were funded by the PRAXIS XXI/FCT, Portugal (BD/15754/98 and BCC/7328/96). V Petrarca was partly supported by MIUR/COFIN funds.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Pinto, J., Donnelly, M., Sousa, C. et al. An island within an island: genetic differentiation of Anopheles gambiae in São Tomé, West Africa, and its relevance to malaria vector control. Heredity 91, 407–414 (2003). https://doi.org/10.1038/sj.hdy.6800348
Received:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.hdy.6800348
Keywords
This article is cited by
-
Is the São Francisco River a historical barrier to gene flow for populations of Melipona mandacaia Smith, 1863 (Hymenoptera: Apidae)?
Journal of Insect Conservation (2023)
-
Effects of indoor residual spraying and outdoor larval control on Anopheles coluzzii from São Tomé and Príncipe, two islands with pre-eliminated malaria
Malaria Journal (2019)
-
Effective population size of Anopheles funestus chromosomal forms in Burkina Faso
Malaria Journal (2006)
-
Genetic structure of Anopheles gambiae populations on islands in northwestern Lake Victoria, Uganda
Malaria Journal (2005)
-
Revisiting the role of introgression vs shared ancestral polymorphisms as key processes shaping genetic diversity in the recently separated sibling species of the Anopheles gambiae complex
Heredity (2004)