Abstract
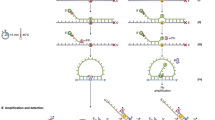
We describe the Multiple-Dye Cleavase Fragment Length Polymorphism (MD-CFLP) method set up for a sensitive and preliminary rapid screening of BRCA1 mutations. We analysed exons 11 and 16, which are known to cover slightly more than 70% of the whole coding region of the gene, subdivided into 4 amplicons and labelled with different fluorescent dUTPs. MD-CFLP was first utilised on a panel of 30 DNA samples in which the presence of single-base substitutions or small deletions/insertions had been previously identified by direct sequencing as gold standard, in order to define the optimal conditions in terms of PCR amplification and temperature of digestion. In a second step, we blindly analysed 21 DNA samples by MD-CFLP to verify its reliability. The sensitivity and specificity of MD-CFLP were both 100% in the first study, and 80% and 94%, respectively, in the blind sample assay. Our results demonstrate the capability of the MD-CFLP method to detect DNA sequence alterations in fragments of more than 1 kb. We conclude that CFLP is a powerful tool in mutational analysis, offering reliable results in a shorter time and at a lower cost than conventional methods, and its potential can be enhanced when internal fluorescent labelling and laser detection are used. © 2001 Cancer Research Campaign http://www.bjcancer.com
Similar content being viewed by others
Article PDF
Change history
16 November 2011
This paper was modified 12 months after initial publication to switch to Creative Commons licence terms, as noted at publication
References
Brow MAD, Oldenburg MC and Lyamichev V et al (1996) Differentiation of bacterial 16S rRNA genes and intergenic regions and Mycobacterium tuberculosis katG genes by structure-specific endonuclease cleavage. J Clin Microbiol 34: 3129–3137
Eisinger F, Jacquemier J, Charpin C and Stoppa-Lyonet D et al (1998) Mutations at BRCA1: the medullary breast carcinoma revisited. Cancer Res 58: 1588–1592
Ford D, Easton DF and Stratton M the Breast cancer Linkage Consortium (1998) Genetic heterogeneity and penetrance analysis of the BRCA1 and BRCA2 genes in breast cancer families. Am J Hum Genet 62: 676–689
Friedman LS, Ostermeyer EA and Szabo CI et al (1994) Confirmation of BRCA1 by analysis of germline mutations linked to breast and ovarian cancer in ten families. Nature Genet 8: 399–404
Hartge P, Struewing JP, Wacholder S, Brody LC and Tucker MA (1999) The prevalence of common BRCA1 and BRCA2 mutations among Ashkenazi Jews. Am J Hum Genet 64: 963–970
Hogervorst FB, Cornelis RS and Bout M et al (1995) Rapid detection of BRCA1 mutations by the protein truncation test. Nature Genet 10: 208–212
Maddox LO, Li P, Bennett A, Descartes M and Thompson JN (1997) Comparison of SSCP analysis and CFLP analysis for mutation detection in the human iduronate 2-sulfatase gene. Biochem Mol Biol Int 43: 1163–1171
Marshall DJ, Heisler LM and Lyamichev V et al (1997) Determination of hepatitis C virus genotypes in the United States by cleavase fragment length polymorphism analysis. J Clin Microbiol 35: 3156–3162
Miki Y, Swensen J and Shattuck-Eidens D et al (1994) A strong candidate for the breast and ovarian cancer susceptibility gene BRCA1. Science 266: 66–71
Ponder B (1997) Genetic testing for cancer risk. Science 278: 1050–1054
Rosetti S, Englisch S, Bresin E, Pignatti PF and Turco AE (1997) Detection of mutations in human genes by a new rapid method: cleavage fragment length polymorphism analysis (CFLPA). Mol Cell Probes 11: 155–160
Serova OM, Mazoyer S and Puget N et al (1997) Mutations in BRCA1 and BRCA2 in breast cancer families: are there more breast cancer-susceptibility genes?. Am J Hum Genet 60: 486–495
Shattuck-Eidens D, Oliphant A and McClure M et al (1997) BRCA1 sequence analysis in women at high risk for susceptibility mutations. JAMA 278: 1242–1250
Sidransky D (1997) Nucleic acid-based methods for the detection of cancer. Science 278: 1054–1058
Sreevatsan S, Bookout JB and Ringpis FM et al (1998) Algorithmic approach to high-throughput molecular screening for alpha interferon-resistant genotypes in hepatitis C patients. J Clin Microbiol 36: 1895–1901
Stoppa-Lyonnet D, Laurent-Puig P and Essioux L et al (1997) BRCA1 sequence variations in 160 individuals referred to a breast/ovarian family cancer clinic. Am J Hum Genet 60: 1021–1030
Tavtigian SV, Simard J and Rommens J et al (1996) The complete BRCA2 gene, mutations in chromosome 13q-linked kindred. Nature Genet 12: 333–337
Wooster R, Bignell G and Lancaster J et al (1995) Identification of the breast cancer susceptibility gene BRCA2. Nature 378: 789–792
Author information
Authors and Affiliations
Rights and permissions
From twelve months after its original publication, this work is licensed under the Creative Commons Attribution-NonCommercial-Share Alike 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/3.0/
About this article
Cite this article
Casadei, S., Cortesi, L., Pensotti, V. et al. Detection of germline BRCA1 mutations by Multiple-Dye Cleavase Fragment Length Polymorphism (MD-CFLP) method. Br J Cancer 85, 845–849 (2001). https://doi.org/10.1054/bjoc.2001.1988
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1054/bjoc.2001.1988
Keywords
This article is cited by
-
Vertical Transmission of Chlamydia trachomatis in Chongqing China
Current Microbiology (2009)
-
Diagnostic accuracy of methods for the detection of BRCA1 and BRCA2 mutations: a systematic review
European Journal of Human Genetics (2007)