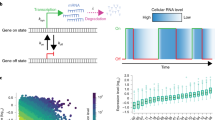
Abstract
Only one X chromosome functions in diploid human cells irrespective of the sex of the individual and the number of X chromosomes. Yet, as we show, more than one X is active in the majority of human triploid cells. Therefore, we suggest that (i) the active X is chosen by repression of its XIST locus, (ii) the repressor is encoded by an autosome and is dosage sensitive, and (iii) the extra dose of this key repressor enables the expression of more than one X in triploid cells. Because autosomal trisomies might help locate the putative dosage sensitive trans-acting factor, we looked for two active X chromosomes in such cells. Previously, we reported that females trisomic for 18 different human autosomes had only one active X and a normal inactive X chromosome. Now we report the effect of triplication of the four autosomes not studied previously; data about these rare trisomies – full or partial – were used to identify autosomal regions relevant to the choice of active X. We find that triplication of the entire chromosomes 5 and 11 and parts of chromosomes 1 and 19 is associated with normal patterns of X inactivation, excluding these as candidate regions. However, females with inherited triplications of 1p21.3–q25.3, 1p31 and 19p13.2–q13.33 were not ascertained. Thus, if a single key dose-sensitive gene induces XIST repression, it could reside in one of these locations. Alternatively, more than one dosage-sensitive autosomal locus is required to form the repressor complex.
Similar content being viewed by others
Introduction
Studies of the molecular mechanisms underlying X inactivation – the means of dosage compensation of X-linked genes in mammals – show that the non-coding XIST RNA is a key player.1 Transcription of XIST at the critical time in embryonic development is sufficient to initiate a cascade of events that silences all the X chromosomes in the cell except the one that remains active. How one X in diploid cells escapes inactivation and is chosen to be the future active X remains to be determined. However, it is likely that autosomes participate in this process. The two active X chromosomes in many 69, XXY and 69, XXX triploid cells2, 3, 4, 5, 6 suggest a role for the extra set of autosomes in protecting X chromosomes from inactivation in triploid cells and, by extension, a role for autosomes in choosing the future active X in diploid cells. The easiest way for an autosomal gene(s) to do this is by repressing the XIST locus on one X chromosome.7 We previously tried to identify candidate autosomes by examining X inactivation in spontaneously aborted females trisomic for individual autosomes. At that time, we determined the X inactivation status for 18 of the 22 possible autosomal trisomies and showed that, in each case, the specimen had one active and one inactive X.8 Assuming a single dosage sensitive gene is sufficient to repress XIST, these chromosomes could be excluded. However, it was impossible to study the rarest trisomies, 1, 5, 11 and 19, which are the subjects of the present study. We now have observations on four females, two with trisomy of chromosome 11, one with trisomy of chromosome 5 and another with tetrasomy of chromosome 5p; in all four cases there was a single active X and a normal inactive X. Because of their rarity, we could not examine the remaining two chromosomes directly. Therefore, to identify candidate regions for the critical autosomal transfactor we ascertained partial trisomies for these chromosomes among live-borns. In addition, we reconsidered the effect of triploidy on X inactivation using direct examination of XIST to look at the characteristics of X inactivation in triploid cells.
Materials and methods
Cells and tissues
Triploid and trisomic cell cultures were derived from spontaneous abortions. Most of the triploid specimens were reported previously by Jacobs et al5 or Migeon et al4 and have been maintained in liquid nitrogen. The others were obtained at the time of miscarriage, according to an IRB-approved protocol. Supplementary Table 1 shows the triploid specimens that are new or restudied by different methods for this paper.
Cytogenetic studies
Data about the sex chromosomes in triploids and non-mosaic trisomics were obtained from the laboratories of co-authors in Honolulu and Wessex (PAJ), Baltimore (GS), New York (CT), from reports in the literature as cited in Table 1, from the Coriell Institute, and the American Cytogenetics Forum List. Queries to cytogenetic laboratories of colleagues did not yield further cases.
X inactivation studies
The presence of an inactive X chromosome was detected by XIST expression and late replication; XIST transcription was analyzed by RTPCR.15 XIST and TSIX RNA were detected by FISH; unfixed metaphase cells were labeled by indirect immunofluorescence. XIST DNA was detected on the same specimens used for XIST RNA in sequential hybridizations.16 Late replicating X chromosomes were identified using the standard BrdU and acridine orange method. X inactivation studies of all the autosomal trisomies except 1, 5, 11, and 19 had been reported by us previously.8
Mapping candidate regions
For chromosomal trisomies that could not be examined directly we looked for partial trisomies among live-borns. Our strategy was based on the assumptions that(i) three copies of the region containing the putative transfactor would allow a second X to remain active and (ii) any fetus with two active X chromosomes would die before birth, as no diploid live-born with two intact active X chromosomes has ever been observed. Information was gathered mainly from two sources: the literature (PubMed) and unpublished data from The Wessex Regional Genetics Laboratory at Salisbury District Hospital, UK. Our inquiry to cytogenetics data banks and cytogeneticist colleagues was not productive, except for six partial trisomies from the Coriell Institute, included in Figure 2. As this approach was initiated prior to our ascertaining cells from the trisomy 5 specimens, we considered any chromosome anomaly that resulted in a partial trisomy of chromosomes 1, 5 and 19. In each case we determined if the evidence was sufficient to document a viable partial trisomy.
The candidates for our region of interest were based on four criteria: (i) the subject must be female, as only females could have two active X chromosomes; (ii) the subject must be live-born, as the ability to survive gestation eliminates the possibility of having two intact active X chromosomes; (iii) the partial trisomy must involve three separate chromosomes, as is the case in unbalanced translocations; (iv) the translocation must be inherited, as de novo translocations may have arisen after the choice of the active X and would not affect the choice process. In addition, because the unbalanced translocations were associated with congenital malformations and were inherited from a phenotypically normal balanced translocation carrier, it is likely that the extra copy of the triplicated segment was expressed. Direct duplications in males and females were also mapped but were not used for exclusions, as both copies are on the same chromosome and might function as one copy rather than two and might be subject to position effects. Because a triple dose of the putative transfactor in males should not affect the activity status of their single X chromosome, we expected to see trisomic regions in males not seen in females. Therefore, we noted the occurrence of familial partial trisomies in males but did not use them for exclusions.
In the case of data reported at a resolution of less than the 700-band level, we interpreted breakpoints conservatively, so as to exclude the smallest region possible. For example, if the breakpoint was reported as 1p21, we interpreted it as 1p21.3.
Finally, for all the candidate regions on human autosomes we identified the orthologous regions in the mouse genome using the NCBI map viewer: comparative maps of Homo sapiens (Build 36.2), at the NCBI website.
Results and discussion
Evidence that triploid cells mainly have two active X chromosomes
The phenotype of human triploids is influenced by the parental origin of the extra haploid set of chromosomes. When it is paternal (diandric), the fetus tends to be more poorly developed and the placenta is abnormal. Zargosa et al9 noted that the majority of human triploids are diandric because dispermy is relatively common; the maternally derived ones result from less frequent errors in oocytes during meiosis II. Because the relative frequency of XYY, XXY and XXX dispermic triploids should be 1:2:1,17 one expects many triploids of the 69 XYY type. However, Table 1 shows that XYY triploids are extremely rare among recognizable triploid specimens, only 19 of 820 triploids (2.3%) having an XYY karyotype. It seems that XYY triploids are created at their expected frequency,18 but do not survive beyond implantation. Table 2 summarizes the results of our X inactivation analysis of XXX and XXY triploid specimens. See Supplementary Table 1 for details. Also included in Table 2 are the results of previous studies based on the presence of a late-replicating X,5 and/or heterodimers for G6PD4 and a recent study based on XIST expression.6 We re-analyzed some specimens and studied new ones using XIST and TSIX RNA as markers for an inactive X.16 In humans, unlike mice, TSIX is a marker for the inactive X chromosome as it is co-expressed with XIST from the inactive X and not from the active one16 (See Figure 1). In all cases, the FISH assay confirmed the findings based on chromosome replication. Of the 47 cases of XXY and XXX triploids studied, two active X chromosomes were seen in every cell in 24 cases and in some cells in 17 cases; they were not seen in only 6 cases. Therefore, 87% of these triploids had two active X's in at least some of their cells (Table 2 and Figure 1a–d).
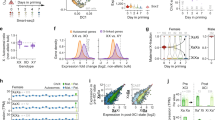
Analysis of X inactivation in triploid (a–d) and trisomic (e and f) cells using XIST and TSIX as markers for the inactive X. Red indicates the signals for XIST RNA and green for TSIX RNA. Yellow shows where the XIST and TSIX signals overlap, as both are expressed from the inactive X chromosome in humans, unlike the situation in the mouse.16 RNA FISH analysis of 69, XXY triploids (a and b) showing no TSIX or XIST signals in specimen 288 indicating two active Xs (a) and specimen 322 with one TSIX/XIST overlapping signal indicating a single active X chromosome (b). The same analysis of 69, XXX triploid specimen JWD (c and d) showing cells with one active X (c) and two active X chromosomes (d) respectively. XIST expression in cells from a 47, XX female with a trisomy for chromosome 11 showing XIST RNA FISH signal (e) and RT-PCR analysis (f).lane 1,marker; lanes 2 and 3, control female; lanes 6 and 7,control male and lanes 3 & 4,47, XX+11 . Even lanes show results in presence of the reverse transcriptase.
The mixed population of cells (some with two active X's and others with only one) seen in 17 of the 47 cases might reflect some instability of the process. Gartler et al6 suggest that the two active X chromosomes result either from reactivation of the inactive X or reduplication of the active one. However, such reactivation or reduplication events are inconsistent with most previous observations: spontaneous reactivation of the whole chromosome has not been seen in any human somatic cell, and duplication has been excluded in cases where the two X's can be distinguished by polymorphic variants.3, 4 Moreover, previous studies show that the ‘two active X phenotypes’ in triploid skin fibroblasts is highly stable4 and this is confirmed in the present study (see JWD clones in Supplementary Table 1). In fact, the mixed population of cells is consistent with expectations – if two copies of an autosomal trans-acting factor are needed for one active X chromosome, then three copies may not always be sufficient for two active X chromosomes. Thus, stochastic variation in transfactor abundance may result in a mixture of cells – some with one active X and others with two of them. In this case, the cells differing in the number of active X's may be subject to selection as they proliferate.6 Jacobs et al5 suggested that the advantage might be tissue specific, and hence reflect the predominant tissue present in the specimen.
In any case, one must conclude that (i) irrespective of their parental origin or sex, most triploid specimens have two active X chromosomes (Xa) in at least some of their cells and often in all of them, and (ii) the one Xa/two Xa mixed cultures originate at the time when the active X is chosen. Therefore, studies of triploids tell us that the factor(s) enabling the activity of more than one X chromosome is not present in sufficient amounts to ensure two active X chromosomes in every cell. On the other hand, because there are two active X's in most triploid specimens, the additional set of chromosomes must in some way enable this to happen, presumably because of extra doses of an autosomal gene(s).
Hypothesis: the human active X is chosen by XIST repression
One way that the extra set of autosomes in triploids could enable the activity of more than one X is if they provide an extra dose of trans-acting factor, and if the transfactor is an XIST repressor. Because an XIST-expressing chromosome cannot be the active X, the XIST locus on that chromosome needs to be repressed. The concept of a blocking factor that protects the active X from inactivation was suggested by Rastan19 and Lyon20 long before the Xist locus was discovered. We suggest that this blocking factor is an XIST repressor, and that in diploid cells there is only enough XIST repressor for a single active X, but in triploid cells there is more. Therefore, based on the observations of triploids our working hypothesis is that (i) the single active X is chosen by XIST repression; (ii) there is an autosomal gene or element that is the XIST repressor or that initiates the events that silence XIST on the future active X; (iii) This XIST repressor is a dosage sensitive gene product: two doses of it can repress only one XIST gene, but three doses can repress more than one XIST gene. Although the transfactor might bind directly to cis elements within the XIST locus, it could work indirectly to repress XIST by binding to other elements within the X inactivation center.
The search for trans-acting factors, using autosomal trisomies
To test our hypothesis, we embarked upon a search with the long-term goal of identifying factors that when expressed in triple dose could enable the activity of two X chromosomes. If triploids can, and usually do, have two active X's and if the XIST repressor was encoded by a single autosomal locus, then trisomies, with three copies of the relevant gene, might also have two active X's. We realized that not all cells trisomic for the putative trans-acting factor might have two active X chromosomes. Nevertheless, while the search for such a transfactor using trisomies cannot prove that it is absent, it can show that it is present.
Initially, we studied female trisomic specimens obtained from miscarriages and could show that for all of them the trisomy did not alter the inactivation process.8 Recently, we ascertained two specimens of trisomy 11, and in each case XIST was expressed from their normal inactive X chromosome (Figure 1d and e).
Being aware that having two active X chromosomes might be lethal prior to the time when a pregnancy is recognized, we examined the sex ratio for the various autosomal trisomies. Conceivably, the trisomy we sought might be more lethal to females than to males as, irrespective of the amount of trans-acting factor, only females could have more than one active X. Table 3 shows the small excess of males among these specimens in general – a good deal of it attributable to the excess of males with trisomy 21 and 22. Chromosomes 1 and 5 were also more prevalent in males, but the number of specimens is very small.
From the literature we learned of three instances of trisomy 1 in recognizable pregnancies (two males and one female),21, 22, 23 and another male with just eight cells.24 The sole female conceptus, an empty sac with no sign of fetal development, was miscarried at 39 days post conception22 relatively close to the time of X inactivation in human fetal tissues (∼20 days). We also studied a specimen trisomic for the entire long arm of chromosome 1 that expressed XIST. However, as there were two normal 46, XX cells in the culture that could not be definitively attributed to maternal contamination, we could not exclude the possibility that the trisomy occurred after X inactivation.
There are no reports of complete trisomy of chromosome 5 in the literature, but recently we studied two relevant female specimens. The first has a complete trisomy 5 and a gestational age of 8 weeks. In all cells with three chromosome 5, only one of the X chromosomes was active, whereas the other was late replicating. The second fetal specimen had four copies of the short arm of chromosome 5, two on the normal chromosome 5s and the other two as components of an isochromosome of the entire short arm; in each of the 30 cells suitable for study, one X was active and the other late replicating.
Mapping of trisomic regions in partial trisomies



For the chromosomes that could not be studied directly (1 and 19), we looked for partial trisomies among live births. This permitted us to identify triplicated regions in individuals who had a single active X. We are aware that such females might have had mosaic X inactivation patterns, surviving only because selection favored cells with a single active X. However, if they were not mosaic initially, then their very existence could exclude these regions as candidates. Partial trisomies were ascertained from the laboratory of one of us (PAJ), from the Corielle Institute and from the literature. We established strict criteria for exclusion (see Materials and Methods). Triplication needed to involve three separate chromosomes and the translocation or insertion had to be familial to assure that it was present at the time that the active X was chosen. Although the locations of tandem duplications were noted, they were not used to eliminate a candidate region. See Figure 2.
Map locations of partial trisomies for chromosomes 1, 5 and 19, for which no trisomic specimens were available. A full trisomy 5 was ascertained only after these studies were initiated (see text). Ideograms of metaphase chromosomes (700 bands, ISCN, 2005)25 showing the location of trisomic regions. Males are shown on the left and females on the right. The numbers (26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, 40, 41, 42, 43, 44, 45, 46, 47, 48, 49, 50, 51, 52, 53, 54, 55, 56, 57, 58, 59, 60, 61, 62, 63, 64, 65, 66, 67, 68, 69, 70, 71, 72, 73, 74, 75, 76, 77, 78, 79, 80, 81, 82, 83, 84, 85, 86) refer to literature citations; the letters P and B refer to cases studied by PAJ and BRM respectively and C refers to specimens from the Coriell collection. *Samples shown to have an inactive X, trisomic regions due to inherited unbalanced translocations. trisomic regions due to de novo or possible de novo duplications, candidate Regions (CR) for autosomal transfactor.
Chromosome 1
A large part of the short arm of chromosome 1 can be eliminated by two case reports. Halal et al46 described a female infant with trisomy 1p31 → 1pter, resulting from a translocation between chromosomes 1 and 2. It is likely that one of the parents (who refused testing) carries a balanced translocation, as there were seven previous miscarriages. The second case is a 30-year-old female with a trisomy resulting from a direct insertion of 1p21 → p31 into the long arm of the chromosome 13 inherited from her mother.49 Based on phenotypic similarities, two other relatives (one male and one female) were also affected. In addition, because of the conservative interpretation of breakpoints, a small region from 1p31.1–p31.3 cannot be excluded. Therefore, the candidate region on the short arm of chromosome 1 is reduced to 1p10 → p21.3 and the small interval in 1p31 (Figure 2).
The distal long arm of chromosome 1 can be eliminated because of two females with a duplication of 1q25 → qter resulting from a familial balanced translocation (Rosenthal et al70 and PAJ). A girl with a trisomy for 1q24-qter caused by a non-familial X/1 translocation had an inactive X chromosome with spreading of inactivation into the translocated 1q segment ameliorating her phenotype;86 this suggests that 1q24.3 → qter could be excluded (Figure 2, asterisk). The previously noted female fetus with full trisomy 1q and XIST transcripts was not used to eliminate candidate regions because of uncertainty as to when the trisomy occurred. Using strict criteria, the major candidate region for the entire chromosome is 1p21.3 → q25.3, but, based on the spreading of X inactivation into the translocation chromosome, it is likely that more of the long arm could be excluded. The orthologous regions in the mouse genome are on mouse chromosomes 3 and 1.
Chromosome 19
Based on viable trisomies, ∼one-third of chromosome 19 can be excluded. The distal segment of the short arm of chromosome 19 is excluded by a female infant with a partial trisomy of 19p13.2 → pter derived from a paternal balanced 3/19 translocation.72 Therefore, the candidate region for the short arm of chromosome 19 is 19p10 → p13.2 (Figure 2).
The distal long arm of chromosome 19 is represented by several females with familial trisomies 19q13.33 → qter.55, 37, 73 Therefore, the candidate region is 19q10 → q13.33 for the q arm, and 19p13.2–q13.33 for the entire chromosome. Two trisomies in males overlap this candidate region (Figure 2). The orthologous regions have been fragmented among several mouse chromosomes, predominantly chromosomes 7 and 8 with some sequences on 10 and 17.
Chromosome 5
Our search for partial trisomies of chromosome 5 among live-borns was initiated before we ascertained the complete trisomy 5 and tetrasomy 5p specimens. We found that the short arm of chromosome 5 could be excluded because of a female with a trisomy of the entire short arm of chromosome 5 inherited from a maternal balanced translocation.29 This finding was confirmed by our studies of the fetal specimen with 5p tetrasomy. Also, most of the long arm of chromosome 5 is not a candidate region based on evidence from live-borns.45, 52, 69, 83, 77, 35 Although several males have been reported,56, 57, 38 there are no females with trisomy spanning any part of 5q22–q31.1. The trisomy of the entire long arm of chromosome 5 reported as due to unproven gonadal mosaicism71 is shown as a dotted line in Figure 2. The only regions of chromosome 5 not represented by partial trisomies (5q22–q31.1 and 5q10–q11.2) can now be excluded by our X inactivation studies of the female specimen with complete trisomy 5.
Conclusions based on observations of X inactivation in trisomies
The results show that for the 20 trisomies (one to nine female specimens of each kind) that were studied directly, there was only a single active X, so X inactivation had occurred in each of the 51 fetal specimens analyzed. Therefore, three copies of these autosomes neither induced the activity of more than one X nor interfered with the ability to inactivate an X. Unlike the triploids, not one of the 51 specimens had cells with two active Xs. Although we observed no mosaicism among any of the 51 trisomic specimens – many of them early first trimester embryos – it is conceivable, but unlikely, that the extra dose of transfactor invariably leads to a mosaic fetus, and the cells with a single active X have a major advantage so that we ascertain only those cells. The time when specimens with two active X chromosomes die is not known, but if all trisomic specimens with two active X chromosomes were lost prior to implantation, then one might expect significantly more male specimens with the relevant trisomy among recognized abortions. Although the numbers are relatively small, there is no obvious selective loss of females for any trisomy (Table 3).
In lieu of the fetal specimens unavailable for study, we ascertained live-born females with partial trisomies of chromosomes 1, 5 and 19 who came to attention because of congenital abnormalities. Not all of the partial trisomies had been studied specifically for their X inactivation pattern, but as no live-born with two intact active X chromosomes has ever been identified, it seemed reasonable to expect that they had an inactive X. We found that some chromosomal regions were not represented in the females with partial trisomies; some of these regions were seen in males. The regions not observed may merely reflect the relatively small number of samples. This seems to be the case for the missing segment of chromosome 5, as we found that complete trisomy for chromosome 5 was associated with an inactive X chromosome. With regard to chromosomes 1 and 19, the alternative explanation for regions that are not represented in females is that they are the ones in which autosomal transfactors lie, and their triplication is incompatible with intrauterine survival.
Using ENU mutagenesis, Percec et al87 looked for autosomal dominant mutations affecting X-inactivation choice in mice and reported some evidence supporting the existence of candidate regions on mouse chromosomes 5, 10 and 15. However, till now no autosomal genes have been definitively identified. We have identified regions of the mouse genome that correspond to our human candidate regions, so as to facilitate experimental attempts to find the relevant genes or elements in mouse models. Searching for physical interactions between X and an autosome in embryos prior to the onset of X inactivation might identify relevant autosomes, as this has been able to identify interacting chromosomes involved in the choice of olfactory alleles.88
We have presented evidence that the two active X phenotype is characteristic of the majority of triploid embryos. We have also proposed that three copies of a single dosage-sensitive transfactor are sufficient for the activity of two active X's, at least in some cells. As a preliminary test of the hypothesis, we have identified potential candidate regions of human autosomes and their corresponding location in the mouse. We are aware that our studies cannot eliminate the possibility of more than one dosage-sensitive locus; in this case, trisomy for a single autosome might not be sufficient to repress more than one XIST locus. In any case, further studies of these candidate regions may identify a key dosage-dependent autosomal locus involved in choosing the active X chromosome. We hope that publication of these results (i) encourages the search for autosome/X interactions within the X inactivation center of mouse embryos during the appropriate window of development, and (ii) elicits clinical and cytogenetic reports of regions of the human chromosome that have not yet been ascertained. If triplication of all of the genome fails to identify a candidate for the putative transfactor, then other possibilities need to be considered. Future studies will either support or refute our hypothesis.
References
Brown CJ, Hendrich BD, Rupert JL et al: The human XIST gene: analysis of a 17 kb inactive X-specific RNA that contains conserved repeats and is highly localized within the nucleus. Cell 1992; 71: 527–542.
Harnden DG : Nuclear sex in triploid XXY human cells. Lancet 1961; ii: 488.
Weaver D, Gartler S, Boue A, Boue JG : Evidence for two active X chromosomes in a human XXY triploid. Hum Genet 1975; 28: 39–42.
Migeon BR, Sprenkle JA, Do TT : Stability of the two active X phenotype in triploid cells. Cell 1979; 18: 637–641.
Jacobs PA, Masuyama AM, Buchanan IM, Wilson C : Late replicating X chromosomes in human triploidy. Am J Hum Genet 1979; 31: 446–457.
Gartler SM, Varadarajan KR, Luo P, Norwood TH, Canfield TK, Hansen RS : Abnormal X:autosome ratio, but normal X chromosome inactivation in human triploid cultures. BMC Genet 2006; 7: 41.
Migeon BR : X chromosome inactivation: theme and variations. Cytogenet Genome Res 2002; 99: 8–16.
Jacobs PA, Migeon BR : Studies of X-chromosome inactivation in trisomies. Cytogenet Cell Genet 1989; 50: 75–77.
Zaragoza M, Surti U, Redline RW, Millie E, Chakravarti A, Hassold TJ : Parental origin and phenotype of triploidy in spontaneous abortions. Am J Hum Genet 2000; 66: 1807–1820.
McFadden DE, Langlois S : Parental and meiotic origin of triploidy in the embryonic and fetal periods. Clin Genet 2000; 58: 192–200.
McFadden DE, Robinson WP : Phenotype of triploid embryos. J Med Genet 2006; 43: 609–612.
Baumer A, Balmer D, Blinkert F, Schinzel A : Parental origin and mechanisms of formation of triploidy: a study of 25 cases. Eur J Hum Genet 2000; 8: 911–917.
Daniel A, Wu Z, Bennetts B et al: Karyotype, phenotype and parental origin in 19 cases of triploidy. Prena Diagn 2001; 21: 1034–1048.
Kajii T, Niikawa N : Origin of triploidy and tetraploidy in man: 11 cases with chromosome markers. Cytogenet Cell Genet 1977; 18: 109–125.
Migeon BR, Chowdhury AK, Dunston JA, McIntosh I : Identification of TSIX encoding an RNA antisense to human XIST, reveals differences from its murine counterpart: implications for X inactivation. Am J Hum Genet 2001; 69: 951–960.
Migeon BR, Lee CH, Chowdhury AK, Carpenter H : Species differences in TSIX/Tsix reveal the roles of these genes in X chromosome inactivation. Am J Hum Genet 2002; 71: 286–293.
Jacobs PA, Morton NE : Origin of human trisomics and polyploids. Hum Hered 1977; 27: 59–72.
Kaufman MH, Speirs S, Lee KK : The sex-chromosome constitution and early postimplantation development of diandric triploid mouse embryos. Cytogenet Cell Genet 1989; 50: 98–101.
Rastan SJ : Non-random X-chromosome inactivation in mouse X-autosome translocation embryos – location of the inactivation centre. J Embryol Exp Morph 1983; 78: 1–22.
Lyon MF, Rastan S : Paternal source of chromosome imprinting and its relevance for X chromosome inactivation. Differentiation 1984; 26: 63–67.
Dunn TM, Grunfeld L, Kardon NB : Trisomy 1 in a clinically recognized IVF pregnancy. Am J Med Genet 2001; 99: 152–153.
Banzai M, Sato S, Matsuda H, Kanasugi H : Trisomy 1 in a case of a missed abortion. J Hum Genet 2004; 49: 396–397.
Hanna JS, Shires P, Matile G : Trisomy 1 in a clinically recognized pregnancy. Am J Med Genet 1997; 68: 98.
Watt JL, Templeton AA, Messinis I, Bell L, Cunningham P, Duncan RO : Trisomy 1 in an eight cell human pre-embryo. J Med Genet 1987; 24: 60–64.
Shaffer LG, Tommerup N : ISCN (2005): An International System for Human Cytogenetic Nomenclature. Basel, Switzerland: S Karger, 2005.
Baialardo EM, del V Torrado M, Barreiro C, Gallego MS : Partial distal 5p trisomy resulting from paternal translocation (5;15)(p15.1;013) in a boy with no mental retardation. Clin Dysmorphol 2003; 12: 257–259.
Bhat M, Morrison PJ, Getty A, McManus D, Tubman R, Nevin NC : First clinical case of small de novo duplication of 19q (13.3 → 13.4) confirmed by FISH. Am J Med Genet 2000; 91: 201–203.
Boyd E, Grass FS, Parke JC, Knutson K, Stevenson R : Duplication of distal 19q: clinical report and review. Am J Med Genet 1992; 42: 326–330.
Brimblecombe FS, Lewis FJ, Vowles M : ‘Complete 5p’ trisomy: 1 case and 19 translocation carriers in 6 generations. J Med Genet 1977; 14: 271–275.
Brown J, Horsley SW, Jung C et al: Identification of a subtle t (16;19)(p13.3;p13.3) in an infant with multiple congenital abnormalities using a 12-colour multiplex FISH telomere assay, M-TEL. Eur J Hum Genet 2000; 8: 903–910.
Byrne JLB, Korn GA, Dev VG et al: Partial trisomy. Am J Hum Genet 1990; 6A: 32.
Cervera M, Sanches S, Molina B et al: Trisomy of the short arm of chromosome 5 due to a de novo inversion and duplication (5)(p15.3p13.3). Am J Med Gen 2005; 136A: 381–385.
Chen H, Kusyk CJ, Tuck-Muller CM, Martinez JE, Dorand RD, Wertelecki W : Confirmation of proximal 1q duplication using fluorescence in situ hybridization. Am J Med Genet 1994; 50: 28–31.
Cousineau AJ, Higgins JV, Hackel E et al: Cytogenetic recognition of chromosomal duplication [dup (1)(p31.4 → p22.1)] and the detection of three different alleles at the PGM locus. Ann Hum Genet 1981; 45: 337–340.
Curry CJR, Loughman WD, Francke U et al: Partial trisomy for the distal long arm of chromosome 5 (region q34 → qter). A new clinically recognizable syndrome. Clin Genet 1979; 15: 454–461.
Dhellemes C, Choiset A, Narbouton R et al: Interstitial dup (1p) and severe intrauterine growth retardation. Ann Genet 1988; 31: 129–131.
Dorn T, Riegel M, Schinzel A, Siegel AM, Kramer G : Epilepsy and trisomy 19q – different seizure patterns in a brother and a sister. Epilepsy Res 2001; 47: 119–126.
Douyard J, Hawley P, Shaham M, Kimois V : Duplication of 5q15-q23.2: case report and literature review. Birth Defects Res A Clin Mol Teratol 2006; 76: 272–276.
Yatsenko SA, Sahoo T, Rosenkranz M, Mendoza-Londono R, Naeem R, Scaglia F : Attenuated phenotype in a child with trisomy for 1q due to unbalanced X;1 translocation [46,X,der(X),t(X;1)(q28;q32.1)]. Am J Med Genet A 2004; 128: 72–77.
Elejalde BR, Opitz JM, de Elejalde MM et al: Tandem dup (1p) within the short arm of chromosome 1 in a child with ambiguous genitalia and multiple congenital anomalies. J Med Genet 1984; 17: 723–730.
Elias-Jones AC, Habibl P, Larcher VF, Spencer T, Butler LJ : The trisomy (5)(q31 → qter) syndrome: study of a family with a t(5;14) translocation. ArcDis Child 1988; 63: 427–431.
Fryns JP, Kleczkowska A, Borghgraef M, Raveschot J, Van den Berghe H : Distinct dysmorphic syndrome in a child with inverted distal 5q duplications. Ann Genet 1987; 30: 186–188.
Furforo L, Rittler M, Slavutsky I : Proximal trisomy 1q in a girl with developmental delay and minor abnormalities. Am J Med Genet 1996; 64: 551–555.
Garcia-Heras J, Corley N, Garcia MF, Kukolich MK, Smith KG, Day DW : De novo partial duplications 1p: report of two new cases and review. Am J Med Genet 1999; 82: 261–264.
Gilgenkrantz S, Dulucq P, Bresson JL, Gouget A, Pernot C, Gregoire MJ : Partial proximal trisomy of the long arm of chromosome 5 (q13 → q22) resulting from maternal insertion der ins (10;5). J Med Genet 1981; 18: 465–480.
Halal F, Vekemans M, der Kaloustian VM : A presumptive translocation 1p;2q resulting in duplication 1p and deletion 2q. J Med Genet 1989; 32: 376–379.
Heilstedt HA, Shapira SK, Gregg AR, Shaffer LG : Molecular and clinical characterization of a patient with duplication of 1p36.3 and metopic synostosis. Clin Genet 1999; 56: 123–128.
Hiraki Y, Fujita H, Yamamori S et al: Mild craniosynostosis with 1p36.3 trisomy and 1p36.3 deletion syndrome caused by familial translocation t(Y;1). Am J Med Genet A 2006; 140: 1773–1777.
Hoechstetter L, Soukup S, Schorry EK : Familial partial duplication (1) (p21p31). J Med Genet 1995; 59: 291–294.
Hustinx T, Nabben F, Scheres J : Partial trisomy of chromosome 1 resulting from a complex maternal rearrangement of chromosomes 1, 5 and 6. Am J Med Genet 1979; 3: 353–358.
Jalbert P, Jalbert H, Sele B et al: Partial trisomy for the long arms of chromosome No. 5 due to insertion and further aneusomie de recombinaison. JMed Genet 1975; 12: 418–423.
Jones LA, Jordan DK, Taysi K, Strauss AW, Toth JK : Partial duplication of the long arm of chromosome 5: a case due to balanced paternal translocation and review of the literature. Hum Genet 1979; 51: 37–42.
Lo IFM, Cheung LYK, Ng AYY, Lam STS : Interstitial dup (1p) with findings of Kabuki make-up syndrome. J Med Genet 1998; 78: 55–57.
Lorda-Sanchez I, Urioste M, Villa A et al: Proximal partial 5p trisomy resulting from a maternal (19;5) insertion. Am J Med Genet 1997; 68: 476–480.
Madokoro H, Ohdo S, Sonoda T, Kawaguchi K, Ohba K : Partial trisomy for 19q due to paternal 18/19 reciprocal translocation. Jpn J Hum Genet 1988; 33: 61–65.
Martin NJ, Cartwright DW, Harvey PJ : Duplication 5q (5q22 → 5q33): from an intrachromosomal insertion. J Med Genet 1985; 20: 57–62.
Martin DM, Mindell MH, Kwierant CA, Glover TW, Gorski JL : Interrupted aortic arch in a child with trisomy 5q31.1q35.1 due to a maternal (20;5) balanced insertion. J Med Genet 2003; 116A: 268–271.
Michels VV, Berseth CL, O'Brien JF, Dewald G : Duplication of part of chromosome 1q: clinical report and review of literature. Am J Med Genet 1984; 18: 125–134.
Mohammed FMA, Farag TI, Gunawardana SS et al: Direct duplication of chromosome 1, dir dup(1)(p21.2 → p32) in a bedouin boy with multiple congenital anomalies. J Med Genet 1998; 32: 353–355.
Mowat D, Jauch A, Robson L, Smith A : Duplication within chromosome 5q characterized by fluorescence in situ hybridization. Am J Med Genet 1999; 83: 361–364.
Mutchinick OM, Shaffer LG, Kashork CD, Cervantes EL : Miller–Dieker syndrome and trisomy 5p in a child carrying a derivative chromosome with a microdeletion in 17p13.3 telomeric to the LIS1 and D17S379 loci. Am J Med Genet 1999; 85: 99–104.
Pettenati M, Berry M, Shashi V, Hartley Bowen J, Harper M : Prenatal diagnosis of complete sole trisomy 1q. Prenat Diagn 2001; 21: 435–440.
Polityko A, Starke H, Rumyantseva N, Claussen U, Liehr T, Raskin S : Three cases with rare interstitial rearrangements of chromosome 1 characterized by multicolor banding. Cytogenet Genome Res 2005; 111: 171–174.
Qorri M, Oei P, Dockery H, McGraghran J : A rare case of a de novo dup (19q) associated with a mild phenotype. J Med Genet 2003; 39: 61.
Quack B, Van Roy N, Verschraegen-Spae MR, Klein F : Interstitial deletion and ring chromosome derived from 19q. Ann Genet 1992; 35: 241–244.
Quigley DI, Kaiser-Rogers K, Aylsworth AS, Rao KW : Submicroscopic deletion 9(q34.2) and duplication 19(p13.3): identified by subtelomer specific FISH probes. Am J Med Genet A 2004; 125: 67–72.
Ramussen SA, Frias JL, Lager CZ, Eunpu DL, Zacaki EH : Partial duplication 1q: report of four patients and review of the literature. J Med Genet 1990; 36: 137–143.
Reichenbach H, Holland H, Dalitz E et al: De novo complete trisomy 5p: clinical report and FISH studies. Am J Med Genet 1999; 85: 447–451.
Rodewald A, Zankl M, Gley EO, Zang KD : Partial trisomie 5q: three different phenotypes depending on different duplication segments. Hum Genet 1980; 55: 191–198.
Rosenthal J, Abeloliovich D, Carmi R : Clinical variability of partial duplication 1q: a clinical report and literature review. J Med Genet 1987; 27: 787–792.
Rovescalli A, Ghidoni A : An unusually large 5q duplication in an adult female subject: spreading of inactivation and in vitro instability of the derivative Xp/5q chromosome. Ann Genet 1989; 32: 235–240.
Salbert BA, Solomon M, Spence JE, Jackson-Cook C, Brown J, Bodurtha J : Partial trisomy 19p: case report and natural history. Clin Genet 1992; 41: 143–146.
Schmid W : Trisomy for the distal third of the long arm of chromosome 19 in brother and sister. Hum Genet 1979; 46: 263–270.
Schorry EK, Dietrich KN, Saal HM et al: Partial trisomy 1q with growth hormone deficiency and normal intelligence. Am J Med Genet 1998; 77: 257–260.
Stratton RF, DuPont BR, Olsen AS, Fertitta A, Hoyer M, Moore CM : Interstitial duplication 19p. Am J Med Genet 1995; 57: 562–564.
Su P-H, Kuo P-L, Chen S-J, Huang S-C, Chen J-Y, Hung H-N : Six cases of deletion 9p24 and trisomy 19q13.4 inherited from a familial balanced translocation. J Formos Med Assoc 2005; 104: 525–530.
Uchiyama A, Haneda N, Saito K et al: A girl with partial trisomy 5q35 → qter and partial trisomy 13pter → q31 derived via a maternal balanced translocation. Eur J Pediatr 2001; 161: 360–361.
Utkus A, Sorokina I, Kucinskas V et al: Duplication of segment 1p21 following paternal insertional translocation, ins (6;1)(q25;p13.3p22.1). J Med Genet 1999; 36: 73–76.
Valerio D, Lavorgna F, Scalona M, Conte A : A new case of partial trisomy 19q (q13.2 → qter) owing to an unusual maternal translocation. J Med Genet 1993; 30: 697–699.
Velagaleti GVN, Morgan DL, Tonk VS : Trisomy 5p. A case report and review. Anne Genet 2000; 43: 143–145.
Warden CR, Pillers DM, Rice MJ et al: Interstitial duplication of the short arm of chromosome 1 in a newborn with congenital heart disease and multiple malformations. J Med Genet 2001; 101: 100–105.
Wieacker P, Missbach D, Jakubiczka S, Borgmann S, Albers N : Sex reversal in a child with the karyotype 46, XY, dup (1) (p22.3p32.3). Clin Genet 1996; 49: 271–273.
Wysocka B, Brozek I, Wierzba J et al: Partial trisomy of distal 5q and partial monosomy of Xp as a result of mating between two translocation carriers: a female with a balanced translocation t (X:5)(p11;q31) and a male with a der (13;14)(q10;q10) – a case report and a family study. Ann Genet 2002; 45: 143–146.
Zabel B, Baumann W, Gehler J, Conrad G : Partial trisomy for short and long arm of chromosome no. 5. J Med Genet 1978; 15: 143–147.
Zonana J, Brown MG, Magenis RE : Distal 19q duplication. Hum Genet 1982; 60: 267–270.
Zuffardi O, Tiepolo L, Scappaticci S et al: Reduced phenotypic effect on partial trisomy 1q in a X/1. Translocation. Ann Genet 1977; 20: 191–194.
Percec I, Thorvaldsen JL, Plenge RM et al: An N-ethyl-N-nitrosourea mutagenesis screen for epigenetic mutations in the mouse. Genetics 2003; 164: 1481–1494.
Lomvardas S, Barnea G, Pisapia DJ, Mendelsohn M, Kirkland J, Axel R : Interchromosomal interactions and olfactory receptor choice. Cell 2006; 126: 403–413.
Acknowledgements
We gratefully acknowledge Cathy Lee, Ashis Chowdhury and Conor Steuer for their contributions to the FISH analysis, RTPCR study and literature search, respectively, and Daniel Saul and Sarah Dowey for DNA replication studies. This work was supported in part by Grant HD 05465 from the National Institute of Child Health and Development to BRM and from The Wellcome Trust to PAJ.
Author information
Authors and Affiliations
Corresponding author
Additional information
Web Resources
The URLs for data presented herein are as follows:
NCBI, URL: http://www.ncbi.nlm.nih.gov/Homology, accessed November 15, 2006.
Supplementary Information accompanies the paper on European Journal of Human Genetics website (http://www.nature.com/ejhg)
Supplementary information
Rights and permissions
About this article
Cite this article
Migeon, B., Pappas, K., Stetten, G. et al. X inactivation in triploidy and trisomy: the search for autosomal transfactors that choose the active X. Eur J Hum Genet 16, 153–162 (2008). https://doi.org/10.1038/sj.ejhg.5201944
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.ejhg.5201944
Keywords
This article is cited by
-
Silencing XIST on the future active X: Searching human and bovine preimplantation embryos for the repressor
European Journal of Human Genetics (2022)
-
Mouse models of aneuploidy to understand chromosome disorders
Mammalian Genome (2022)
-
Thoughts about SLC16A2, TSIX and XIST gene like sites in the human genome and a potential role in cellular chromosome counting
Molecular Cytogenetics (2016)
-
X-chromosome inactivation: molecular mechanisms from the human perspective
Human Genetics (2011)
-
The single active X in human cells: evolutionary tinkering personified
Human Genetics (2011)