Abstract
Chromosome duplications are found in about 2% of subjects with a typical chromosomal phenotype but their frequency is likely to be higher, as suggested by the first array-CGH data. According to the orientation of the duplicated segment, duplications may be in tandem or inverted. The latter are usually associated with a distal deletion. We studied a de novo 2.3 Mb inverted duplication of 8q24.3 without apparently associated deletion in a subject with profound psychomotor retardation, idiopathic epilepsy and growth delay. In spite of its small size, the presence of the rearrangement was suspected on standard karyotypes (approximately 400 bands) and later confirmed by Fluorescent in situ hybridization (FISH) analysis. We hypothesize that the GRINA gene, a glutamate binding subunit of NMDA receptor ion channel lying within the duplicated segment, may be responsible for the epilepsy. This paper confirms that small subtelomeric de novo duplications may be responsible for mental retardation, facial dysmorphisms and/or congenital malformations, although their presence may be overlooked by FISH analysis.
Similar content being viewed by others
Introduction
Chromosome duplications have an approximate frequency of 1:4000 in the general population1 and are found in about 2% of subjects with a typical chromosomal phenotype.2 Moreover, the first array-CGH screenings on these patients indicate that cryptic duplications have a much higher frequency, even considering that some of them might represent just polymorphisms.3, 4 According to the orientation of the duplicated segment, the duplications may be either in tandem or inverted.5 For both types of rearrangement segmental duplications play a primary role in causing nonallelic homologous recombination.6 Unlike direct duplications, inverted duplications are usually associated with the deletion of the distal region of the duplicated chromosome. Between the duplicated and the deleted region, a single copy region may be present flanked by homologous segmental duplications. The inv dup(8p) is the paradigmatic example of this type of rearrangement.7, 8 We describe here an inverted duplication of 8q with apparently no associated deletion in a child with profound psychomotor retardation, idiopathic epilepsy and growth delay. The rearrangement was rather obvious in 400 band karyotypes, although molecular techniques demonstrated that the duplication involves only the distal 2.3 Mb of 8q.
Materials and methods
Case report
MA, aged 6 years, is the second child of a healthy 37-year-old woman and a nonconsanguineous 39-year-old man, whose family history was unremarkable.
The pregnancy was uneventful, except for feeble foetal movements, which began at 6 months of gestation. Ultrasound findings at 5 and 8 months were referred as normal, with only a slight reduction of the biparietal diameter. Caesarean section was required at the 39th gestational week for breech presentation. Birth weight was 2850 g (between 10th and 25th centile) and birth length was 52 cm (between 50th and 75th centile). Head circumference was not recorded.
Owing to sucking and swallowing difficulties, poor growth and psychomotor retardation at the age of 3 months, the child was admitted for diagnosis to another hospital where a cerebral echo scan, ophthalmologic evaluation and visual-evoked potentials and EEG were performed, all with normal results. Cerebral MRI scan and a standard chromosomal analysis were also reported as normal.
Screening for aminoacidaemias, organic acidurias and fatty acid disorders were normal.
Cerebral MRI, repeated at the age of 1 year and 8 months, and brain stem auditory-evoked potentials at the age of 2 years and 4 months were normal.
At the age of 3.5 months, repeated episodes of generalized seizures were observed, controlled with barbiturate and vigabatrin. No seizures have been observed after the age of 11 months. Only the barbiturate therapy was maintained after the age of 2.5 years.
At the age of 5 years, generalized spike-wave discharges of typical absence seizures were detected without apparent clinical effect.
Psychomotor retardation was clearly evident from the first months of age. The child's language skills were below expectations for her developmental quotient (DQ). At the Brunet-Lezine Psychomotor Development Scale at the age of 2 years and 4 months and of 3 years and 8 months DQ was, respectively, 48 and 56 corresponding to a profound psychomotor retardation. Growth has been delayed with weight constantly around the 3rd centile, length between 3rd and 10th centile and head circumference between −0.5 and −1.0 s.d.
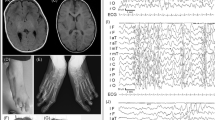
On physical examination at the age of 3 years and 9 months, she presented with short proportionate stature, prominence of the metopic suture with small but proportionate skull, mild ptosis of the left eyelid, large ears with prominent antitragus, large nose, malar hypoplasia, prominent and short philtrum and high palate. Bilateral clinodactyly and camptodactyly of the five fingers was observed. A cafè-au-lait spot with a diameter of 1 cm on the chest was observed. The remaining of the physical examination was normal.
From the age of 3 years till the last evaluation at the age of 6 years and 2 months isolated plasma transaminase (AST, ALT) and gamma glutamyl transpeptidase (GGT) levels have exceeded 2–3 times the normal values. Abdominal sonography and liver biopsy were normal.
Cytogenetic and molecular analysis
Routine cytogenetic analysis was performed on proposita and parents’ blood using standard G-bands with a resolution of approximately 400 bands.
Fluorescent in situ hybridization (FISH) with whole chromosome painting (wcp) (Multi-M kit, Cytocell Ltd, Adderbury, England) and with telomere-specific probes (Multi-T kit, Cytocell Ltd, Adderbury, England) for all chromosome was carried out on patient's metaphases.
Other FISH experiments were performed with bacterial artificial chromosome (BAC) and prokaryotic artificial chromosome (PAC) clones containing chromosome 8q24-specific sequences from several locations according to the publicly available genome resources (NCBI Map Viewer: http://www.ncbi.nlm.nih.gov; Santa Cruz Human Genome Browser: http://genome.ucsc.edu; http://genome.imb-jena.de/). DNA clones were labelled with biotin and/or digoxigenin using standard nick translation reactions. Double-colour FISH was performed with BAC clones RP11-520P18 (AC083982, ctg NT_037701) and RP11-1022M7 (AC083959 ctg NT_031818) labelled with biotin and digoxigenin, respectively. The probes were visualized with fluorescein isothiocyanate–avidin (Vector Laboratories, Burlingame, CA, USA) or rhodamine-labelled antidigoxigenin (Oncor Inc., Gaithersburg, MD, USA) and the chromosomes were counterstained with DAPI (Sigma-Aldrich, Milano, Italy). Hybridizations were analysed with an Olympus BX61 epifluorescence microscope and images were captured with the Power Gene FISH System (PSI, Newcastle upon Tyne, UK).
DNA was extracted from 3 ml of peripheral blood with standard protocols. Polymorphic loci were selected based on the NCBI Human July 2003 Assembly viewed on the UCSC browser; amplifications were performed with ABI fluorochrome-labelled primers and analysed on an ABI Prism 310 Genetic Analyzer (Applied Biosystems, Monza, Italy).
Results
Cytogenetic investigation revealed an alteration on one chromosome 8 consisting of a slight difference in size at the G-negative 8q24.3 band between the two homologues. This difference was evident even if the banding resolution was no higher than 400 bands (Figure 1). Wcp with libraries of all chromosomes revealed that the abnormal chromosome 8 was entirely composed by chromosome 8 material. FISH analysis with all subtelomeric regions was performed to exclude the presence of a cryptic translocation that could have been missed by the use of painting probes. This analysis demonstrated that the 8q subtelomeric clone was present at the end of the abnormal 8q, although the corresponding signal was much bigger than the one detected on the homologue. In metaphase, this 8q signal was compact and not interrupted by unstained regions, while in interphase two clearly distinguished and separated signals were evident (Figure 2a). FISH analysis with BAC and PAC clones from 8q (Table 1) demonstrated a duplication of about 2.3 Mb from RP11-520P18 clone (AC083982, ctg NT_023684, 144.1–144.3 Mb) to AF235103 (RP1-291P5, RP1-316H24, RP4-659B21, RP5-1047A19, RP5-1109M23, ctg NT_037704, 145.98–146.2 Mb).
Proband's FISH analysis: (a) Specific subtelomeric probes for chromosome 8p (green) and 8q (red). Abnormal chromosome 8 (arrowhead) shows on 8qtel (red) a signal bigger than the one detected on the homologue (arrow). In metaphase, this 8q signal was compact and not interrupted by unstained regions, while in interphase, two clearly distinguished and separated signals are evident. (b) BAC clones RP11-520P18 (green) and RP11-1022M7(red) demonstrating the inverted duplication (arrowhead).
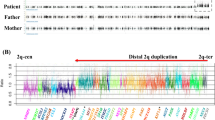
Dual-colour FISH (Figure 2b) revealed that the duplication was inverted. According to the whole of the data, the final cytogenetic interpretation was 46,XX,add(8)(q24.3).ish.inv dup (8)(q24.3) (wcp8+; WI-654+; SHGC-5263++; WI-12784++; RH40626++; RH74682++; 8QTEL25++) with the duplication encompassing the last 2.3 Mb of 8q. Cytogenetic and FISH analysis with Tel8q probe (Cytocell) on proposita's parents gave normal results. A PCR-based assay to establish the origin of the duplication was performed with markers 8QTEL11, 8QTEL25 and RH74682. RH74682 typing demonstrated that the duplication was of maternal origin, while 8QTEL25 showed that it involved only one of the two maternal alleles (Figure 3).
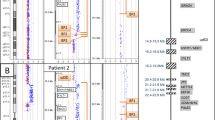
A schematic map of the 8q24.3 region showing the relative location of all probes is presented in Figure 4.
Map of the 8q24.3 region showing the relative position of contigs, sequence gaps, STS and FISH probes, and a selection of known genes; the drawing is based on the UCSC map, November 2003 release; FISH clones in the duplicated region, as well as the GRINA and GPT genes, are shown in a darker shade of grey.
Discussion
The rearrangement
Inverted duplications are probably more frequent than suspected until 5–10 years ago. In fact, some of them, originally interpreted as direct duplications on pure cytogenetic analysis, turned out to be complex rearrangements after proper molecular analysis showing not only a duplicated but also a deleted region. The studies on the inv dup(8p), a recurrent rearrangement associated with a rather characteristic syndrome,9 revealed that it derives by nonallelic homologous recombination between low-copy repeats.7, 8, 10 Other examples of inv dup are those concerning 1q,11, 12 2q,13 3p,14, 15 4p,16 5p,17, 18 9p,19 10p and 10q,20 18p,21 18q,22 21q23 and the X;X or the Y;Y rearrangements leading to duplications of parts of the short or the long arm with concomitant deletion,24 suggesting that the mechanism responsible for the inv dup(8p) can be generalized to all inverted duplications. In our inv dup(8q) case, we were not able to find any associated deletion distal to the last duplicated clone RP5-1109M23 nor any segmental duplication responsible for the occurrence of the inv dup(8q). In this regard, the inv dup(8q) seems to be atypical. However, in the duplicated 8q24.3 region two gaps are still present (UCSC, release July 2003). In silico analysis25 revealed that most of the genome sequence gaps are associated with segmental duplications and that telomeric regions are significantly enriched in segmental duplication content. We believe that the results we obtained are not compatible with the current sequence release of the distal 8q and it seems possible that a refinement of the sequence will support a mechanism similar to the classic inv dup rearrangements. Indeed, the finding that interphase FISH analysis with the subtelomeric probe (TEL8q Cytocell) (Figure 2a) showed two clearly separated signals suggests that a single copy region might be present distal to the duplicated one. Although we could not demonstrate the presence of the distal deletion associated to the duplication, we cannot exclude it and it seems likely that the original product of the abnormal maternal meiosis originating the rearrangement present in the proposita were a dicentric chromosome 8pter → 8q24.3∷8p24.3 → 8pter that, at meiosis II or during early embryogenesis,26 underwent a breakage leading to the present inv dup(8)(q24.3) characterized by a very distal 8q deletion and a 2.3 Mb duplication.
Genotype/phenotype correlation
The phenotype of our patient is characterized by severe psychomotor and mental retardation, idiopathic epilepsy (seizure without apparent clinical effect) and elevated blood levels (2–3 times higher then normal values) of plasma transaminase (AST, ALT) and GGT.
Among patients with epilepsy and intellectual disability, about 6% have chromosomal abnormalities and this figure climbs to 50% in patients with seizures and multiple congenital anomalies.27, 28 To date no less than 400 different chromosomal imbalances have been associated with seizures or EEG abnormalities consisting in either deletions or duplications.29
The 8q24 region was previously implicated in different forms of epilepsy (MIM 601068; MIM 603210; MIM 606970; MIM 607876; MIM 600131; MIM 602232; MIM 121201; MIM 138251). The KCNQ3 locus, whose mutations are associated to BFNC (benign familial neonatal convulsions, MIM 121200), lies 10 Mb upstream from the 8q segment duplicated in our patient. Thus, any involvement of this gene in the aetiology of our patient's epilepsy is excluded. However, the GRINA gene (MIM 138251) (Figure 4), a glutamate binding subunit of NMDA receptor ion channel proposed as a candidate for the idiopathic generalized epilepsy mapping at 8q24 (MIM 600669), is located in the duplicated 8q region. We hypothesize that duplication of the GRINA gene is responsible for the epilepsy of our patient. The finding that ion channel genes are dosage sensitive, clearly demonstrated at least for haploinsufficiency,30 reinforces our hypothesis.
The human GPT gene (MIM 138200) was mapped to 8q24.3,31, 32 within 200 kb of the 8q telomere, thus within the duplicated region (Figure 4). This finding suggests that the increase of at least the GPT values is due to gene dosage effect. An increased GPT activity was similarly reported in a patient with 8q24.2-qter duplication.32
Final considerations
Subtelomeric FISH studies demonstrating the 2.3 Mb duplication have been performed to investigate the suspicion situation of 8q noticed on 400 G-band karyotypes and not because requested by the clinicians. In the last few years, a general consensus has been reached on the fact that the more closely chromosomes are studied, the lower is the chance of finding a rearrangement with the molecular test;33 in everyday practice, rearrangements of 5–10 Mb or even larger are easily missed.33, 34 Thus, it is rather astonishing that our 8q duplication was suspected on 400 G-band karyotypes. We might hypothesize that the chromatin status of the duplicated distal 2.3 Mb 8q is less compact than the rest of the metaphasic chromosome, as if the duplication had induced an epigenetic decondensation of the chromatin at least in lymphocytes. Whereas no example of chromatin decondensation induced by a chromosome rearrangement is known, the opposite phenomenon, that is, the silencing of the chromatin accompanied by chromatin condensation, is well known especially in unbalanced X/autosome translocations in which up to 45 Mb of the translocated autosome may be silenced and condensed.35 We shall see in the future if similar observations will confirm that chromatin decondensation induced by a chromosome rearrangement may render visible otherwise undetectable rearrangements.
Among the more than 2500 cases tested for subtelomeric rearrangements, only three de novo duplications have been detected36 out of 99 rearrangements. Obviously, duplications are more difficult to detect by FISH, since the operator should be on the look out for differences in the size of the signal and not simply discriminate between the presence or absence of a signal. Therefore, their frequency may have underestimated. This paper confirms that small subtelomeric de novo duplications may be responsible for mental retardation and congenital anomalies.
References
Phelan MC, Crawford EC, Bealer DM : Mental retardation in South Carolina III. Chromosome aberrations. Proc Greenwood Genet Center 1996; 15: 45–60.
Shaffer LG, Lupski JR : Molecular mechanisms for constitutional chromosomal rearrangements in humans. Annu Rev Genet 2000; 34: 297–329, (review).
Vissers LE, de Vries BB, Osoegawa K et al: Array-based comparative genomic hybridization for the genomewide detection of submicroscopic chromosomal abnormalities. Am J Hum Genet 2003; 73: 1261–1270.
Shaw-Smith C, Redon R, Rickman L et al: Microarray based comparative genomic hybridisation (array-CGH) detects submicroscopic chromosomal deletions and duplications in patients with learning disability/mental retardation and dysmorphic features. J Med Genet 2004; 41: 241–248.
Van Dyke DL, Miller MJ, Weiss L : The origin of inverted tandem duplications, and phenotypic effects of tandem duplication of the X chromosome long arm. Am J Med Genet 1983; 15: 441–450.
Shaw CJ, Lupski JR : Implications of human genome architecture for rearrangement-based disorders: the genomic basis of disease. Hum Mol Genet 2004; 13 (Spec No. 1): R57–R64 : [Epub 2004 Feb 05].
Floridia G, Piantanida M, Minelli A et al: The same molecular mechanism at the maternal meiosis I produces mono- and dicentric 8p duplications. Am J Hum Genet 1996; 58: 785–796.
Giglio S, Broman KW, Matsumoto N et al: Olfactory receptor–gene clusters, genomic-inversion polymorphisms, and common chromosome rearrangements. Am J Hum Genet 2001; 68: 874–883.
de Die-Smulders CE, Engelen JJ, Schrander-Stumpel CT et al: Inversion duplication of the short arm of chromosome 8: clinical data on seven patients and review of the literature. Am J Med Genet 1995; 59: 369–374.
Samonte RV, Eichler EE : Segmental duplications and the evolution of the primate genome. Nat Rev Genet 2002; 3: 65–72, (review).
Mewar R, Harrison W, Weaver DD, Palmer C, Davee MA, Overhauser J : Molecular cytogenetic determination of a deletion/duplication of 1q that results in a trisomy 18 syndrome-like phenotype. Am J Med Genet 1994; 52: 178–183.
De Brasi D, Rossi E, Giglio S et al: Inv dup del (1) (pter → q44∷q44 → q42:) with the classical phenotype of trisomy 1q42-qter. Am J Med Genet 2001; 104: 127–130.
Bonaglia MC, Giorda R, Poggi G et al: Inverted duplications are recurrent rearrangements always associated with a distal deletion: description of a new case involving 2q. Eur J Hum Genet 2000; 8: 597–603.
Jenderny J, Poetsch M, Hoeltzenbein M et al: Detection of a concomitant distal deletion in an inverted duplication of chromosome 3. Is there an overall mechanism for the origin of such duplications/deficiencies? Eur J Hum Genet 1998; 6: 439–444.
Kennedy D, Silver MM, Winsor EJ et al: Inverted duplication of the distal short arm of chromosome 3 associated with lobar holoprosencephaly and lumbosacral meningomyelocele. Am J Med Genet 2000; 91: 167–170.
Cotter PD, Kaffe S, Li L, Gershin IF, Hirschhorn K : Loss of subtelomeric sequence associated with a terminal inversion duplication of the short arm of chromosome 4. Am J Med Genet 2001; 102: 76–80.
Sreekantaiah C, Kronn D, Marinescu RC, Goldin B, Overhauser J : Characterization of a complex chromosomal rearrangement in a patient with a typical catlike cry and no other clinical findings of cri-du-chat syndrome. Am J Med Genet 1999; 86: 264–268.
Perfumo C, Cerruti Mainardi P, Mainardi P et al: The first three mosaic cri du chat syndrome patients with two rearranged cell lines. J Med Genet 2000; 37: 967–972.
Teebi AS, Gibson L, McGrath J, Meyn MS, Breg WR, Yang-Feng TL : Molecular and cytogenetic characterization of 9p− abnormalities. Am J Med Genet 1993; 46: 288–292.
Hoo JJ, Chao M, Szego K, Rauer M, Echiverri SC, Harris C : Four new cases of inverted terminal duplication: a modified hypothesis of mechanism of origin. Am J Med Genet 1995; 58: 299–304.
Moog U, Engelen JJ, de Die-Smulders CE et al: Partial trisomy of the short arm of chromosome 18 due to inversion duplication and direct duplication. Clin Genet 1994; 46: 423–429.
Courtens W, Grossman D, Van Roy N et al: Noonan-like phenotype in monozygotic twins with a duplication-deficiency of the long arm of chromosome 18 resulting from a maternal paracentric inversion. Hum Genet 1998; 103: 497–505.
Pangalos C, Theophile D, Sinet PM et al: No significant effect of monosomy for distal 21q22.3 on the Down syndrome phenotype in ‘mirror’ duplications of chromosome 21. Am J Hum Genet 1992; 51: 1240–1250.
Schinzel A : Catalogue of Unbalanced Chromosome Aberrations in Man, 2nd edn. Berlin, New York: de Gruyter, 2001.
Eichler EE, Clark RA, She X : An assessment of the sequence gaps: unfinished business in a finished human genome. Nat Rev Genet 2004; 5: 345–354, (review).
Pramparo T, Giglio S, Gregato G et al: Inverted duplications: how many of them are mosaic? Eur J Hum Genet 2004; 12: 713–717.
Ieshima A, Takeshita K : Chromosome abnormalities and epileptic seizure. Jpn J Hum Genet 1988; 33: 49–60.
Aicardi J : Disease of the Nervous System in Childhood, Clinics in Developmental Medicine No. 115/118. London: Mac Keith Press, 1992, pp 240–267.
Singh R, Gardner RJ, Crossland KM, Scheffer IE, Berkovic SF : Chromosomal abnormalities and epilepsy: a review for clinicians and gene hunters. Epilepsia 2002; 43: 127–140, (review).
Biervert C, Schroeder BC, Kubisch C et al: Potassium channel mutation in neonatal human epilepsy. Science 1998; 279: 403–406.
Sohocki MM, Sullivan LS, Harrison WR et al: Human glutamate pyruvate transaminase (GPT): localization to 8q24.3, cDNA and genomic sequences, and polymorphic sites. Genomics 1997; 40: 247–252.
Rocha J, Amorim A, Almeida VM et al: Gene dosage evidence for the regional assignment of GPT (glutamate-pyruvate transaminase; E.C. 2.6.1.2) locus to 8q24.2-8qter. Hum Genet 1988; 80: 299–300.
Flint J, Knight S : The use of telomere probes to investigate submicroscopic rearrangements associated with mental retardation. Curr Opin Genet Dev 2003; 13: 310–316.
Biesecker LG : The end of the beginning of chromosome ends. Am J Med Genet 2002; 107: 263–266.
Sharp AJ, Spotswood HT, Robinson DO, Turner BM, Jacobs PA : Molecular and cytogenetic analysis of the spreading of X inactivation in X;autosome translocations. Hum Mol Genet 2002; 11: 3145–3156.
Riegel M, Baumer A, Jamar M et al: Submicroscopic terminal deletions and duplications in retarded patients with unclassified malformation syndromes. Hum Genet 2001; 109: 286–294.
Acknowledgements
We are grateful to the YAC Screening Centre of San Raffaele Biomedical Science Park (Milan) and to Gaiping Wen at Institute of Molecular Biotechnology (Jena, Germany) for providing BAC and PAC clones. This work was supported by cofin02- and cofin03-MIUR (to OZ), the Italian Telethon Foundation (GP0247Y01 to OZ) and the Cariplo Foundation (to OZ).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Bonaglia, M., Giorda, R., Tenconi, R. et al. A 2.3 Mb duplication of chromosome 8q24.3 associated with severe mental retardation and epilepsy detected by standard karyotype. Eur J Hum Genet 13, 586–591 (2005). https://doi.org/10.1038/sj.ejhg.5201369
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.ejhg.5201369
Keywords
This article is cited by
-
Cytogenomic epileptology
Molecular Cytogenetics (2023)
-
A gain-of-function screen to identify genes that reduce lifespan in the adult of Drosophila melanogaster
BMC Genetics (2014)
-
Scratch regulates neuronal migration onset via an epithelial-mesenchymal transition–like mechanism
Nature Neuroscience (2013)
-
TMBIM3/GRINA is a novel unfolded protein response (UPR) target gene that controls apoptosis through the modulation of ER calcium homeostasis
Cell Death & Differentiation (2012)