Abstract
p53 is a potent inhibitor of cell growth and an inducer of apoptosis. During embryonic development, Mdm2 and Mdm4 inhibit the growth suppressive activities of p53. However, whether tight surveillance of p53 activity is required in quiescent cells is unknown. To test this, conditional inactivation of mdm2 and mdm4 was carried out in smooth muscle cells (SMCs). Upon SMC-specific inactivation of mdm2, and not of mdm4, mice rapidly became ill and died. Necropsy showed small intestinal dilation, and histological analyses indicated a severe reduction in the number of intestinal SMCs. Increased p53 levels and activity were detected in the remaining SMCs, and the phenotype was completely rescued on a p53-null background. Interestingly, intestinal SMCs are caspase-3-negative and therefore did not undergo caspase-3-dependent apoptotic cell death. Together, Mdm2, but not Mdm4, prevents accumulation of active p53 in quiescent SMCs and thereby the induction of p53-mediated caspase-3-independent cell death.
Similar content being viewed by others
Introduction
The p53 tumor suppressor protein plays a critical role in suppressing tumor formation by inducing two types of antiproliferative responses: cell cycle arrest and apoptotic cell death. Cell cycle arrest is mediated by transcriptional induction of genes whose products inhibit cell cycle progression, such as p21Waf1/Cip11 or Ptprv.2 The apoptotic function of p53 depends on both transcription-dependent and transcription-independent mechanisms.3
The importance of p53 in tumor suppression is highlighted by the observation that virtually all human cancers display an impaired p53 response. This is achieved either through direct inactivating mutations within the p53 gene, or through aberrant expression of proteins acting in the p53 pathway. For instance, Mdm2 and Mdm4 are overexpressed in human tumors of diverse origin.4, 5 The mdm2 gene was originally identified as an amplified and overexpressed gene in a spontaneously transformed mouse BALB/c cell line.6 Its transformation potential was later demonstrated and explained by the ability of Mdm2 to inactivate p53.7 Mdm4 was identified because of its ability to interact physically with p53.8 Mdm4 overexpression inhibits p53 transcriptional activity in transfection experiments.8 Accordingly, Mdm4 is overexpressed in human cancer cell lines and primary tumors harboring wild-type p53.9, 5
Lack of functional mdm2 or mdm4 is lethal in mouse embryos, which die before implantation and around mid-gestation, respectively. These dramatic phenotypes are completely rescued by concomitant deletion of p53.10, 11, 12, 13 In addition, mice with a hypomorphic mdm2 allele have lower body weight and higher rates of p53-dependent apoptosis in various tissues. Notably, this phenotype is only observed in a subset of actively dividing cells, such as lymphocytes, and in the crypts of the small intestine, which could indicate that, in adult mice, Mdm2 restraints p53 activity only in homeostatic tissues.14 Together, clear genetic evidence highlights the importance of the p53/Mdm2 and p53/Mdm4 interactions. However, a clear understanding of the physiological contributions of Mdm2 and Mdm4 to the regulation of p53 stability and activity is still lacking. Limitations of the existing mouse models, such as early embryonic lethality of the constitutive null mutation, indeed preclude analysis of the function of Mdm2 and Mdm4 in a spatial and temporal specific manner.
In addition, these models do not allow firm establishment of the role of Mdm2 in the regulation of p53 stability. It was indeed shown that, besides the ability of Mdm2 to bind p53 in its transactivation domain and to interfere with p53 transcriptional activity, Mdm2 acts as an E3 ubiquitin ligase responsible for the ubiquitination of p53 and itself.15, 16, 17 It was later proposed that Mdm2 mediates monomeric p53 ubiquitination on multiple lysine residues, rather than polyubiquitination, as previously thought.18 Because chains of multiple ubiquitin molecules are necessary for efficient protein degradation, the data suggested that the enzymatic activity of Mdm2 might not be sufficient for optimal degradation of p53, and that other proteins must aid in polyubiquitination and degradation of p53 in vivo. More recent data indicated that Mdm2 differentially catalyzes either monoubiquitination or polyubiquitination of p53 in a dosage-dependent manner.19 The authors proposed that Mdm2-mediated polyubiquitination and nuclear degradation occurs only in specific contexts, such as when Mdm2 is malignantly overexpressed. On the other hand, Mdm2-mediated monoubiquitination and subsequent cytoplasmic translocation of p53 may represent an important means of p53 regulation in unstressed cells, in which Mdm2 is maintained at low/physiological levels. In addition, in mice with the hypomorphic mdm2 allele, the level of p53 protein was not coordinately increased, suggesting that Mdm2 can inhibit p53 function in a manner independent of degradation.14 Moreover, other cellular ubiquitin ligases, such as Pirh2, Cop-1 or ARF-BP1, were reported to also promote p53 ubiquitination and degradation.20, 21, 22 Thus, whereas Mdm2 is a key regulator of p53 function in vivo, p53 degradation may be mediated through both Mdm2-dependent and Mdm2-independent pathways in vivo.
Here, we show that specific inactivation of mdm2, but not of mdm4, in terminally differentiated intestinal smooth muscle cells (SMCs) results, concomitantly with severe cell loss, in increased p53 protein levels and transcriptional activity. Interestingly, expression of caspase-3 could not be detected in intestinal SMCs and, therefore, SMCs undergoing cell death did not show evidence of caspase-3 activation. In conclusion, Mdm2, but not Mdm4, is critical for the regulation of p53 steady-state levels and activity in these quiescent cells. Moreover, this is the first report describing the ability of p53 to mediate caspase-3-independent cell death in vivo.
Results
Strategy for conditional bi-allelic inactivation of mdm2 and mdm4 in quiescent SMCs in vivo
To test whether Mdm2 and Mdm4 are required for regulating p53 stability and activity in quiescent cells in vivo, we specifically inactivated mdm2 and mdm4 in G0 SMCs. To this end, conditional inactivation of both genes was carried out in mice harboring mdm2 or mdm4 floxed alleles and a tamoxifen (TMX)-inducible Cre-recombinase under the control of the SM22 promoter (SM-CreERT2(ki) mice).23 The mdm2 and the mdm4 floxed alleles (FM) had been previously described.24, 25 They carry loxP recombination sites in introns 4 and 6 of mdm2 and in introns 1 and 2 of mdm4 (Figure 1a). Cre-mediated recombination yields an mdm2 allele lacking exons 5 and 6, which encode for most of the p53-binding domain. Cre-mediated recombination yields an mdm4 allele lacking exon 2, the first coding exon. Mice homozygous for the FM alleles (both mdm2FM/FM and mdm4FM/FM) appear normal; however, ubiquitous deletion with Cre recombinase in vivo resulted in an embryonic lethality similar to the mdm2 or mdm4 null alleles.24, 25
(a) Schematic representation of part of the mdm2 and mdm4 floxed alleles (FM) and the Cre-mediated recombination events. The a, b and c, d arrows designate the position of the primers used to detect the Cre-mediated recombination by PCR. (b) PCR analysis showing increased detection of the Cre-mediated recombination event, using primers a and b in the intestine of CreERT2(ki);mdm2FM/FM mice (left panel) or c and d in the intestine of CreERT2(ki);mdm4FM/FM mice, treated with TMX for the indicated time. (c) Percentage of β-gal+ cells among the total number of SMCs present in sections of SM-CreERT2(ki);Rosa26 organs/tissues after 7 days of intraperitoneal TMX administration. No recombination was observed in TMX- or vehicle-treated control Rosa26 mice. (d) Detection of lacZ reporter gene expression in the stomach of SM-CreERT2(ki);Rosa26 mice and Rosa26;CreER-negative control mice after 7 days of TMX administration
We first examined by PCR the extent of recombination at both mdm loci in SM-CreERT2(ki);mdm2FM/FM and SM-CreERT2(ki);mdm4FM/FM mice (Figure 1b) and compared it with the Cre activity at the Rosa26 locus in SM-CreERT2(ki);Rosa26 mice at various sites containing SMCs (Figure 1c). SMCs were identified both morphologically and immunohistochemically by SM-α-actin staining (Figure 2c). In order to induce the latent CreER fusion protein, mice were injected daily with TMX for 7 days, and then analyzed. Upon TMX injection, we found that SMCs from the gastrointestinal (GI) tract, particularly the stomach (Figure 1d) and proximal ileum, were stably marked (β-galactosidase-positive (β-gal+)), whereas little reporter activity was found in cells of the cardiovascular system such as in the aorta. As expected, based on the use of an SMC-specific promoter to drive the expression of the Cre-ER fusion protein, no recombination was observed in the liver or spleen of TMX-injected SM-CreERT2(ki);Rosa26 mice (Figure 1c). As efficient recombination was observed in the SMCs of the small intestine (proximal ileum), we concentrated our studies at this site.
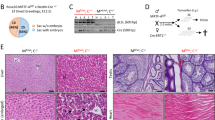
(a) Kaplan–Meier curves of age-matched CreERT2(ki);mdm2FM/FM mice (open circles, n=24) and control CreER-negative mice (filled squares, n=17). Mice were TMX-treated for 7 consecutive days. (b) Gross appearance of control (left panel) and CreERT2(ki);mdm2FM/FM mice (right panel) after 7 days of intraperitoneal TMX administration. (c) Hematoxylin–phloxin–saphron (HPS; left panels) and IHC for SM-α-actin (right panels) staining of the small intestine of CreERT2(ki);mdm2FM/FM mice 0, 5 and 7 days following TMX treatment (magnification × 200)
TMX-treated SM-CreERT2(ki);mdm2FM/FM mice exhibit severe lesions in the SMC-containing layers of the intestinal wall and eventually die
Following 7 days of TMX administrations, the body weight of SM-CreERT2(ki);mdm2FM/FM mice decreased as compared to TMX-treated control Cre-ER-negative mice. Moreover, SM-CreERT2(ki);mdm2FM/FM mice were not responsive to stimuli and were hunchbacked with ruffled coat. Strikingly, illness proceeded to death from day 8 on (Figure 2a). In contrast, TMX-treated mdm2FM/FM CreER-negative and, importantly, SM-CreERT2(ki);mdm4FM/FM mice (Figure 3a) appeared normal and did not differ from vehicle-treated SM-CreERT2(ki);mdm2FM/FM SM-CreERT2(ki);mdm4FM/FM or mdm2FM/FM CreER-negative mice.
(a) Kaplan–Meier curves of age-matched CreERT2(ki);mdm2FM/FM mice (open circles, n=13), CreERT2(ki);mdm4FM/FM mice (open squares, n=6) and CreERT2(ki);mdm2FM/FMp53−/− mice (filled triangles, n=6) following 7 days of intraperitoneal TMX administration. (b) Hematoxylin and eosin (H&E) staining and p53 immunostaining of the small intestine of mice exposed to TMX for the indicated number of days (magnification × 40). (c) p53 and p21 protein levels determined by Western blotting analysis in extracts prepared from the small intestine of non-injected mdm2FM/FM CreER-negative (lane 1) and CreERT2(ki);mdm2FM/FMp53−/− (lane 2) mice, mdm2FM/FM CreER-negative (lane 3) and CreERT2(ki);mdm4FM/FM (lane 4) mice TMX-treated for 7 days, CreERT2(ki);mdm2FM/FM mice treated for 3 days (lane 5) and 7 days (lane 6). Extracts from UV-exposed MEFs were used as positive control (C+). α-Actin serves as a loading control. (d) Q-RT-PCR analysis shows induction of expression of p53 target genes in laser capture microdissected SMCs of mice with the indicated genotypes. Microdissected samples from three different mice were pooled before the analysis. The data represent therefore the mean expression levels in these three mice treated independently
SM-CreERT2(ki);mdm2FM/FM SM-CreERT2(ki);mdm4FM/FM and control mice were killed for gross necropsy and histopathological analysis at several time points. The stomach and small intestine of TMX-treated SM-CreERT2(ki);mdm2FM/FM mice adhered to spleen and liver, appeared vulnerable and friable, were filled with soft materials and were loose. Dilation of the small intestine was observed in TMX-treated SM-CreERT2(ki);mdm2FM/FM mice, and not in CreERT2(ki);mdm4FM/FM or control mice. This phenotype was variable between mice, but could be considerable (Figure 2b). This dilation was associated with a decreased length of the small intestine (from pylorus to ileo-cecal junction: 44.0±3.5 cm for control mice, 32.0±2.0 cm for SM-CreERT2(ki);mdm2FM/FM mice; P=0.007). Of note, histopathological examination revealed only very minor liver and spleen atrophy, in both TMX-injected SM-CreERT2(ki); mdm2FM/FM and mdm2FM/FM CreER-negative mice (data not shown). Moreover, serum levels of the liver enzyme ALAT, a marker of liver damage, were only slightly elevated but comparable in TMX-treated SM-CreERT2(ki); mdm2FM/FM and mdm2FM/FM CreER-negative mice (data not shown). As moderate atrophy of the liver and slightly elevated levels of ALAT were not observed in non-TMX-injected mice of the same genotypes, we concluded that these effects are the result of repetitive TMX injections.
The lesions in the dilated small intestine of SM-CreERT2(ki);mdm2FM/FM mice treated with TMX were limited to the lamina interna and externa of the muscularis (M.) propria of the intestinal wall, consistent with a specific activity of the Cre in the SMCs. The architecture of the lamina M. interna was disturbed owing to a decrease in cell layers and irregular alignment of the SMCs. Between 5 and 7 days after the first TMX injection, the number of SMC layers decreased from 6–8 to 3–5, with multifocal irregular increase of intercellular spaces, tapering of SMCs with wavy ends and irregular, often shortened hyperchromatic nuclei. At later time points (day 9), the SMCs were often simply missing (Figure 2c). In contrast, no obvious histological defects were observed in TMX-treated SM-CreERT2(ki);mdm4FM/FM mice and thus even 12 days after the first TMX-injection. Together, the data suggest that, Mdm2, but not Mdm4, is critical for the survival of the SMCs of the GI tract.
Alterations of the SMC-containing intestinal wall and lethality upon SMC-specific mdm2 inactivation are entirely p53-dependent
A large body of evidence suggests that Mdm2 can function both dependently and independently of p53. In agreement, Mdm2 binds several proteins involved in the regulation of cell cycle progression and survival other than p53, such as p19ARF p73, Rb and E2F-1/DP-1.26 In order to test whether the phenotype observed in the SM-CreERT2(ki);mdm2FM/FM mice is p53-dependent, they were crossed with p53-null mice (p53−/−)27 to create SM-CreERT2(ki);mdm2FM/FMp53−/− mice. Strikingly, as observed in control mice, TMX injection in SM-CreERT2(ki);mdm2FM/FMp53−/− mice did not cause death (Figure 3a). Gross necropsy did not reveal differences in liver and spleen weight and small intestinal length as compared to control mice. Histological examination did not reveal any obvious lesions, disorganization of intestinal cell alignment or loss of cell viability in SM-CreERT2(ki);mdm2FM/FMp53−/− (Figure 3b). We, therefore, concluded that loss of mdm2 in the SMCs of the GI tract causes loss of cell viability and acute lethality in a manner that is completely dependent on the presence of functional p53.
SMC-specific mdm2 inactivation leads to increased p53 levels and transcriptional activity
To determine whether specific deletion of mdm2 in the SMCs allows the level of p53 protein to increase, we performed immunostaining for p53. In sections of the proximal ileum from mdm2FM/FMCreER-negative mice, and importantly of SM-CreERT2(ki);mdm4FM/FM mice, injected with TMX, no p53 staining could be detected (Figure 3b). In contrast, nuclei of the SMCs of SM-CreERT2(ki);mdm2FM/FM mice showed marked p53 immunoreactivity as early as after 3 days of injection (Figure 3b). The p53 staining was consistently more pronounced after 5 and 7 days of injection. Importantly, no staining was observed in sections from SM-CreERT2(ki);mdm2FM/FMp53−/− mice, confirming the specificity of the p53 detection method (Figure 3b). The increase in p53 levels in SM-CreERT2(ki);mdm2FM/FM but not in SM-CreERT2(ki);mdm4FM/FM TMX-treated mice was further confirmed by Western blotting analysis (Figure 3c). These results suggest that p53 is maintained at low levels in a strict Mdm2-dependent manner in terminally differentiated SMCs. In addition, p53 was not only stabilized but it was also functionally active, as indicated by concomitant upregulation of several p53 target genes such as p21Waf1/Cip1 and the proapoptotic gene bax in laser capture microdissected SMCs in the proximal ileum of SM-CreERT2(ki);mdm2FM/FM mice, as determined by Q-RT-PCR (Figure 3d). Of note, mdm2 Δexons5/6 transcripts were also found upregulated in the SMCs of SM-CreERT2(ki);mdm2FM/FM mice. In agreement, upregulation of p21 could also be demonstrated at the protein level by Western blot analysis (Figure 3c).
Caspase-3 is not expressed in intestinal SMCs
As one of the main p53 antiproliferative activities is induction of apoptotic cell death, we hypothesized that the hypocellularity observed following mdm2 inactivation was a consequence of increased p53-mediated apoptosis. However, absence of the non-activated form of caspase-3 protein (pro-caspase-3), the main effector caspase, was previously reported in rat and mouse adult skeletal muscles, despite the abundant presence of its mRNA.28 Similarly, pro-caspase-3 was not significantly detected in the intestinal SMCs of both control (CreER-negative) and SM-CreERT2(ki);mdm2FM/FM TMX-treated mice (Figure 4a). This observation suggests that p53-mediated cell death in intestinal SMCs is caspase-3-independent. To investigate this possibility further, we performed staining using an antibody specifically recognizing the activated form of caspase-3. As expected, cleaved caspase-3 could not be detected at all time points analyzed, both in control and SM-CreERT2(ki);mdm2FM/FM TMX-treated mice (Figure 4a and b). To verify that our detection method could detect both forms of caspase-3, sections of E16.5 embryos expressing p53 specifically in post-mitotic neurons deficient for mdm2 were analyzed.29 Clear and specific staining for pro-caspase-3 and activated form of caspase-3 was detected in these embryos (Figure 4), confirming the sensitivity and the specificity of the method.
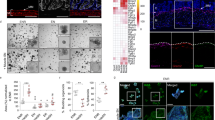
Immunostaining for pro-caspase-3 (non-cleaved caspase-3) (a) and activated form of caspase-3 (casp-3*) (a and b) of the small intestine of mice with the indicated genotypes and in the lateral ventricle region of the cerebral cortex of E16.5 embryos expressing p53 specifically in post-mitotic neurons deficient for mdm229 (control) (magnification × 40, except when indicated otherwise)
Absence of apoptotic signatures in intestinal SMCs undergoing p53-mediated cell death
As other effector caspases might compensate for the lack of caspase-3 in intestinal SMCs, we explored the presence of other apoptotic features. Apoptosis is morphologically defined by several hallmarks including nuclear shrinkage (pyknosis), chromatin condensation, DNA degradation and nuclear fragmentation (karyorrhexis) with formation of apoptotic bodies. Examination of H&E or 4′-6-diamidino-2-phenylindole (DAPI) staining did not reveal the presence of apoptotic bodies or pyknotic nuclei. Moreover, in situ end labeling (ISEL)30 (Figure 5a) and TUNEL assays (data not shown) did not detect DNA fragmentation at any time points. In contrast, DNA fragmentation was evident in mdm2-deficient post-mitotic neurons expressing p5329 (Figure 5a).
(a) ISEL staining of the small intestine of mice with the indicated genotypes and in the lateral ventricle region of the cerebral cortex of E16.5 embryos expressing p53 specifically in post-mitotic neurons deficient for mdm229 (control) (magnification × 40). (b) Immunostaining for the mitochondrial protein endonuclease G (EndoG) in the small intestine of mice exposed to TMX for the indicated number of days (magnification × 40). (c) Electron microscopy of the small intestine of control (creER-negative; top panels) and CreERT2(ki);mdm2FM/FM (low panels) TMX-treated mice. Nuclei in control tissues are sparse, equally distributed and surrounded by a plasma membrane. Nuclei in tissues from CreERT2(ki);mdm2FM/FM mice are clustered and not surrounded by a plasma membrane. Evidence of plasma membrane ruptures is shown in the middle right panel. Evidence of dilation of cytoplasmic organelles such as endoplasmic reticulum (bottom left panel) or mitochondria (bottom right panel) is presented. Magnifications are indicated
Mitochondria permeability change, occurring during apoptosis, may also release factors that trigger caspase-independent apoptotic-like cell death, such as EndoG31 and AIF.32 To further investigate whether some of the cytotoxic effect was due to caspase-independent cell death pathways, we examined the cellular localization of EndoG in TMX-treated mice at various time points. As expected, immunostaining located EndoG to the mitochondria in control animals (Figure 5b). Furthermore, even if the level of EndoG appears slightly increased in SM-CreERT2(ki);mdm2FM/FM TMX-treated mice, there was no significant translocation into the nuclei of the SMCs (Figure 5b).
Electron microscopy confirmed absence of pyknosis and karyorrhexis with formation of apoptotic bodies. This analysis revealed ‘mottled’ nuclei caused by clumped, but not marginalized, and only loosely packed chromatin (Figure 5c). Most of the nuclei appeared largely intact; however, in some cases, nuclear membrane detachment and rupture were apparent. The affected cells also showed dilated cytoplasmic organelles such as endoplasmic reticulum, Golgi and mitochondria (Figure 5c). External membrane rupture was also evident (Figure 5c). In agreement, clusters of roughly intact nuclei, which are no longer surrounded by cytoplasmic membranes, were also observed. Various intermediate aspects could be observed, which ranged from apparently intact cells that had a mottled nucleus, to cells that had a mottled nucleus and gross membrane alterations. Finally, whereas apoptosis includes phagocytosis, even heavily altered cells appeared not to be within phagocytes. Surprisingly, absence of phagocytosis did not lead in this case to local signs of inflammation, but this absence might be related to local anatomical constraints. Together, the observed phenotype is not suggested to coincide with an increase in apoptotic cell death.
Discussion
It is well documented that the p53 protein is maintained at low levels in embryonic and adult tissues; however, it remained to be established whether p53 degradation occurs in a strict Mdm2-dependent manner. Here, we provide genetic evidence that Mdm2 plays a key role in maintaining p53 at low levels in the SMCs of the GI tract. This observation is entirely consistent with very recent additional reports describing conditional inactivation of mdm2 in other cell types. Loss of mdm2 in cardiomyocytes and proliferating and post-mitotic neuronal cells also leads to dramatic accumulation of the p53 protein.25, 29, 33 Thus, even if other E3 protein ligases have been reported to induce p53 ubiquitination and degradation,20, 21, 22 none of these proteins can fully compensate for loss of Mdm2 function in vivo. Therefore, even if these data do not exclude the possibility that these proteins might aid in p53 degradation, they strongly suggest that p53 degradation in vivo occurs through Mdm2-dependent pathways.
While downregulation of Mdm2 is sufficient to activate a p53 response in homeostatic tissues,14 its relevance in non-proliferating, terminally differentiated cells remained unexplored. Herein, we show that Mdm2 is critical to maintain p53 activity in check in quiescent, terminally differentiated SMCs. This observation is also in line with a recent report describing the importance of Mdm2 for the downregulation of p53 activity in post-mitotic neurons.29 These observations have clinically relevant implications. Mdm2 ubiquitination activity and the physical interaction between Mdm2 and p53 have become the targets of adjuvant chemotherapies designed to sensitize human tumors to cancer therapies.34, 35 There is indeed clear evidence that cancer cells are more sensitive to activation of p53 apoptotic function than the resting host cells. However, as Mdm2 is critical for maintaining p53 activity at low levels also in quiescent, terminally differentiated cells, these data would predict that complete ablation of Mdm2 function in vivo would be detrimental not only to cancer cells but also to resting host cells. Thus, it seems that controlled, limited inhibition of Mdm2 is necessary to reach specific killing of the cancer cells.
In contrast to Mdm2, Mdm4 appears dispensable for controlling p53 levels and activity in the SMCs of the GI tract. Loss of mdm4 in these cells indeed did not lead to any obvious phenotype or histological abnormalities within the timeframe of the study. Similarly, mice lacking mdm4 in the heart were born at the correct ratio and appeared normal.25 In contrast, loss of mdm4 in neuronal progenitors and in post-mitotic neurons leads to activation of p53 and cell cycle arrest or apoptosis, respectively.29, 33 Together, inhibition of p53 by Mdm4 seems to be required only in a restricted number of cell types. In the same line, although very high levels of recombination were found in the bladder of SM-CreERT2(ki); Rosa26 mice, TMX injection in SM-CreERT2(ki); mdm2FM/FM mice did not show increased p53 levels in bladder cells nor obvious bladder abnormalities (data not shown). This observation, therefore, suggests that, similar to Mdm4, Mdm2 might be dispensable in limited cell types.
All previous studies converge to the crucial role of both the mitochondrial pathway (cytochrome c release, ROS production and/or Δψm drop) and activation of caspase-9 and its downstream caspases36 in p53-induced apoptosis. In agreement with this view, Apaf-137 and several genes encoding for the BH3-containing proapoptotic proteins such as Bax, Noxa and Puma are among the many reported p53-regulated genes and key mediators of p53-induced apoptosis in vivo.38, 39 Caspase-3, the main effector caspase, acting downstream of caspase-9, is also of fundamental importance for many forms of apoptosis and required for p53-induced apoptosis. In agreement, inactivation of the mdm2 gene led to increased p53 activity and caspase-3-dependent cell death in all cellular settings analyzed thus far.25, 29, 33 In contrast, the data presented herein provide the first compiling evidence that p53 is able to activate a caspase-3-independent cell death program in vivo.
Additional experiments are required to firmly establish the mechanism of cell death associated with the progressive clearance of the mdm2-deficient SMCs. One possibility is that p53 once activated provokes, through some of its mediators, mitochondrial damages that are sufficient to induce cell death in a caspase-independent manner. We could indeed detect in mdm2-deficient SMCs high levels of expression of Bax, a protein particularly important for the triggering of mitochondrial outer membrane permeabilization (MOMP) and cytochrome c release. MOMP frequently marks the ‘point of no return’ of the lethal process. It is clear today that artificial or genetic inhibition of caspases is often not sufficient to avoid physiological cell death, and in this case this often leads to a shift in the morphology of cell death, from classical apoptosis to ‘apoptosis-like cell death’, autophagic cell death or necrosis. For instance, the loss of interdigital cells in the mouse embryo, a prototype of mitochondrial apoptotic cell death, still occurs by necrosis in mice deficient for Apaf-1. Similarly, because they lack caspase-3, the mdm2-deficient SMCs could undergo necrotic cell death following activation of MOMP by p53. In agreement with this possibility, many of the remaining SMCs had a morphotype reminiscent of that seen in necrosis. Alternatively, p53 might be able to activate directly a genetic program leading to MOMP-independent cell death in vivo in a tissue-specific manner. Interestingly, p53-mediated MOMP-independent (and Bcl-2-insensitive) cell death was recently described in cultured cells.40
Regardless of the molecular mechanism, this study opens new perspectives for cancer therapies. For instance, inactivation of Apaf-1, which is essential for p53-induced caspase-3-dependent apoptosis, may contribute to the low frequency of p53 mutations observed in therapy-resistant melanomas.36 The ability of p53 to induce caspase-3-independent cell death may be the basis for new therapies killing cells in which p53 is wild type but have acquired defects in the signaling pathways that are downstream p53.
Materials and Methods
Transgenic mice
To achieve SMC-specific mdm2 and mdm4 deletion, we combined mice that carry a TMX-inducible Cre-recombinase under the control of the SMC-specific SM22 promoter (SM-CreERT2(ki) mice)23 and mice carrying mdm2 or mdm4 floxed alleles24, 25 to create SM-CreERT2(ki);mdm2FM/FM and CreERT2(ki);mdm4FM/FM mice. To determine p53-dependent effects of mdm2 deletion, SM-CreERT2(ki);mdm2FM/FM mice were crossed with p53 knockout (p53−/−) mice27 resulting in SM-CreERT2(ki);mdm2FM/FMp53−/− mice. In addition, to facilitate the monitoring of Cre activity in vivo, we combined the SM-CreERT2(ki) mice and the Rosa26 reporter mouse line to generate SM-CreERT2(ki); Rosa26 mice. The SM-CreERT2(ki);mdm2FM/FM SM-CreERT2(ki);mdm2FM/FMp53−/−, SM-CreERT2(ki);Rosa26 and their control littermates mice were born at the expected Mendelian frequency, developed normally and were genotyped by PCR, as described previously.24, 25, 27
Conditional deletion of mdm2 and quantification of recombination
Mice, aged 8–10 weeks, were injected intraperitoneally with 100 μl of 20 mg/ml TMX (Sigma) or vehicle (peanut oil) for 0, 2, 5 and 7 continuous days. Intraperitoneal TMX injections did not result in liver toxicity as measured by serum amino-L-acetyl transferase levels (40.6±12.7 for mdm2FM/FM mice versus 36.3±16.0 U/l for SM-CreERT2(ki);mdm2FM/FM mice; P=0.754, ALT, Roche). Recombination of both mdm2 and mdm4 floxed alleles was assessed by PCR.24, 25 Recombination in SM-CreERT2(ki);Rosa26 mice was quantified by counting β-gal+ cells and was expressed as a percentage of the total number of SMCs present (Figure 1c).
Tissue preparation and histology
Mice were killed and a complete gross necropsy was performed. Organs and selected tissues, including esophagus, stomach, jejunum, proximal and distal ileum, colon, aorta, urinary bladder and liver, were sampled for further investigation. Sampled organs were either directly snap frozen in sterile eppendorf tubes and stored at −80°C or fixed in phosphate-buffered formalin pH 7.4 and embedded in paraffin. Of the tissues sampled and fixed in formalin, microscopical analysis was performed of 5 μm sections routinely stained with HPS, HE or DAPI. In addition, slides were stained with antibodies against SM-α-actin (clone 1A4, dilution 1 : 1500, DAKO), pro-caspase-3 (1 : 1000, Cell Signaling), cleaved form of caspase-3 (1 : 1000, Cell Signaling), p53 (CM5, 1 : 1000, Novocastra Lab Ltd.) and EndoG (1 : 500, Chemicon International). DNA fragmentation (ISEL staining) was assessed with the FragEL kit (Oncogene Research Products) according to the manufacturer's directions and sections were counterstained with methyl green (Vector Laboratories).
Laser capture microdissection and Q-PCR
Laser capture microdissection samples were prepared from frozen sections of three control and three SM-CreERT2(ki);mdm2FM/FM mice and pooled. Total RNA was extracted using the PicoPure RNA isolation kit and amplified using the RiboAmp RNA Amplification Kit according to the manufacturer's instructions (Acturus Bioscience). A 1 μg portion of total RNA from each pool was reverse-transcribed using a SuperScript kit (Invitrogen). These assays were performed following the manufacturer's specifications (PE Applied Biosystems). Primer pairs and TaqMan probes were designed by Applied Biosystems (Assays on demand).
Western blot analysis
Western blot analysis was performed as previously described.13 The primary antibodies used were as follows: anti-α-actin mouse monoclonal, clone 1A4 (1 : 400, Sigma), anti-p53 sheep polyclonal Ab-7 (1 : 300, Calbiochem) and anti-p21 mouse monoclonal F5 (Santa Cruz Biotechnology).
Electron microscopy
Tissue samples were immersed in a fixative solution of 2% paraformaldehyde and 2.5% glutaraldehyde and post-fixed in 1% OsO4 with 1.5% K3Fe(CN)6 in 0.1 M NaCacodylate buffer, pH 7.2. Samples were dehydrated through a graded ethanol series, including a bulk staining with 2% uranyl acetate at the 50% ethanol step followed by embedding in Spurr's resin. Ultrathin sections, made on an Ultracut E microtome (Reichert-Jung), were post-stained in an ultrastainer (Leica, Herburgg, Switzerland) with uranyl acetate and lead citrate. Sections were viewed with a transmission electron microscope 1010 (JEOL, Tokyo, Japan).
Statistical analysis
All data are represented as mean±S.D. Data were analyzed using the non-parametric Mann–Whitney rank sum test. P-values less than 0.05 were regarded as statistically significant.
Abbreviations
- β-gal:
-
β-galactosidase
- Mdm2:
-
murine double minute 2
- SMC:
-
smooth muscle cell
- TMX:
-
tamoxifen
References
el-Deiry WS, Tokino T, Velculescu VE, Levy DB, Parsons R, Trent JM, Lin D, Mercer WE, Kinzler KW and Vogelstein B (1993) WAF1, a potential mediator of p53 tumor suppression. Cell 75: 817–825.
Doumont G, Martoriati A, Beekman C, Bogaerts S, Mee PJ, Bureau F, Colombo E, Alcalay M, Bellefroid E, Marchesi F, Scanziani E, Pelicci PG and Marine JC (2005) G1 checkpoint failure and increased tumor susceptibility in mice lacking the novel p53 target Ptprv. EMBO J. 24: 3093–3103.
Chipuk JE and Green DR (2003) p53's believe it or not: lessons on transcription-independent death. J. Clin. Immunol. 23: 355–361.
Momand J, Jung D, Wilczynski S and Niland J (1998) The MDM2 gene amplification database. Nucleic Acids Res. 26: 3453–3459.
Danovi D, Meulmeester E, Pasini D, Migliorini D, Capra M, Frenk R, de Graaf P, Francoz S, Gasparini P, Gobbi A, Helin K, Pelicci PG, Jochemsen AG and Marine JC (2004) Amplification of Mdmx (or Mdm4) directly contributes to tumor formation by inhibiting p53 tumor suppressor activity. Mol. Cell. Biol. 24: 5835–5843.
Fakharzadeh SS, Trusko SP and George DL (1991) Tumorigenic potential associated with enhanced expression of a gene that is amplified in a mouse tumor cell line. EMBO J. 10: 1565–1569.
Momand J, Zambetti GP, Olson DC, George D and Levine AJ (1992) The mdm2 oncogene product forms a complex with the p53 protein and inhibits p53-mediated transactivation. Cell 69: 1237–1245.
Shvarts A, Steegenga WT, Riteco N, Van Laar T, Dekker P, Bazuine M, Van Ham RC, Van der Houven van Oordt W, Hateboer G, Van der Eb AJ and Jochemsen AG (1996) MDMX: a novel p53-binding protein with some functional properties of MDM2. EMBO J. 15: 5349–5357.
Ramos YF, Stad R, Attema J, Peltenburg LT, van der Eb AJ and Jochemsen AG (2001) Aberrant expression of Hdmx proteins in tumor cells correlates with wild-type p53. Cancer Res. 61: 1839–1842.
Jones SN, Roe AE, Donehower LA and Bradley A (1995) Rescue of embryonic lethality in Mdm2-deficient mice by absence of p53. Nature 378: 206–208.
Montes de Oca Luna R, Wagner DS and Lozano G (1995) Rescue of early embryonic lethality in mdm2-deficient mice by deletion of p53. Nature 378: 203–206.
Parant J, Chavez-Reyes A, Little NA, Yan W, Reinke V, Jochemsen AG and Lozano G (2001) Rescue of embryonic lethality in Mdm4-null mice by loss of Trp53 suggests a nonoverlapping pathway with MDM2 to regulate p53. Nat. Genet. 29: 92–95.
Migliorini D, Lazzerini-Denchi E, Danovi D, Jochemsen A, Capillo M, Gobbi A, Helin K, Pelicci PG and Marine JC (2002) Mdm4 (Mdmx) regulates p53-induced growth arrest and neuronal cell death during early embryonic mouse development. Mol. Cell. Biol. 22: 5527–5538.
Mendrysa SM, McElwee MK, Michalowski J, O'Leary KA, Young KM and Perry ME (2003) mdm2 is critical for inhibition of p53 during lymphopoiesis and the response to ionizing irradiation. Mol. Cell. Biol. 23: 462–472.
Haupt Y, Maya R, Kazaz A and Oren M (1997) Mdm2 promotes the rapid degradation of p53. Nature 387: 296–299.
Kubbutat MH, Jones SN and Vousden KH (1997) Regulation of p53 stability by Mdm2. Nature 387: 299–303.
Honda R and Yasuda H (2000) Activity of MDM2, a ubiquitin ligase, toward p53 or itself is dependent on the RING finger domain of the ligase. Oncogene 19: 1473–1476.
Lai Z, Ferry KV, Diamond MA, Wee KE, Kim YB, Ma J, Yang T, Benfield PA, Copeland RA and Auger KR (2001) Human mdm2 mediates multiple mono-ubiquitination of p53 by a mechanism requiring enzyme isomerization. J. Biol. Chem. 276: 31357–31367.
Li M, Brooks CL, Wu-Baer F, Chen D, Baer R and Gu W (2003) Mono- versus polyubiquitination: differential control of p53 fate by Mdm2. Science 302: 1972–1975.
Leng RP, Lin Y, Ma W, Wu H, Lemmers B, Chung S, Parant JM, Lozano G, Hakem R and Benchimol S (2003) Pirh2, a p53-induced ubiquitin-protein ligase, promotes p53 degradation. Cell 112: 779–791.
Dornan D, Wertz I, Shimizu H, Arnott D, Frantz GD, Dowd P, O'Rourke K, Koeppen H and Dixit VM (2004) The ubiquitin ligase COP1 is a critical negative regulator of p53. Nature 429: 86–92.
Chen D, Kon N, Li M, Zhang W, Qin J and Gu W (2005) ARF-BP1/Mule is a critical mediator of the ARF tumor suppressor. Cell 121: 1071–1083.
Kuhbandner S, Brummer S, Metzger D, Chambon P, Hofmann F and Feil R (2000) Temporally controlled somatic mutagenesis in smooth muscle. Genesis 28: 15–22.
Grier JD, Yan W and Lozano G (2002) Conditional allele of mdm2 which encodes a p53 inhibitor. Genesis 32: 145–147.
Grier JD, Xiong S, Elizondo-Fraire AC, Parant JM and Lozano G (2006) Tissue-specific differences of p53 inhibition by Mdm2 and Mdm4. Mol. Cell. Biol. 26: 192–198.
Iwakuma T and Lozano G (2003) MDM2, an introduction. Mol. Cancer Res. 1: 993–1000.
Jacks T, Remington L, Williams BO, Schmitt EM, Halachmi S, Bronson RT and Weinberg RA (1994) Tumor spectrum analysis in p53-mutant mice. Curr. Biol. 4: 1–7.
Ruest LB, Khalyfa A and Wang E (2002) Development-dependent disappearance of caspase 3 in skeletal muscle is post-transcriptionally regulated. J. Cell. Biochem. 86: 21–28.
Francoz S, Froment P, Bogaerts S, De Clercq S, Maetens M, Doumont G, Bellefroid E and Marine J-C (2006) Mdm4 and Mdm2 cooperate to inhibit p53 activity in proliferating and quiescent cells in vivo. Proc. Natl. Acad. Sci. USA 103: 3232–3237.
Gavrieli Y, Sherman Y and Ben Sasson SA (1992) Identification of programmed cell death in situ via specific labeling of nuclear DNA fragmentation. J. Cell Biol. 119: 493–501.
Li LY, Luo X and Wang X (2001) Endonuclease G is an apoptotic Dnase when released from mitochondria. Nature 412: 95–99.
Cande C, Cecconi F, Dessen P and Kroemer G (2002) Apoptosis-inducing factor (AIF): key to the conserved caspase-independent pathways of cell death? J. Cell Sci. 115: 4727–4734.
Xiong S, Van Pelt CS, Elizondo-Fraire AC, Liu G and Lozano G (2006) Synergistic roles of Mdm2 and Mdm4 for p53 inhibition in central nervous system development. Proc. Natl. Acad. Sci. USA 103: 3226–3231.
Vassilev LT (2004) Small-molecule antagonists of p53–MDM2 binding: research tools and potential therapeutics. Cell Cycle 3: 419–421.
Yang Y, Ludwig RL, Jensen JP, Pierre SA, Medaglia MV, Davydov IV, Safiran YJ, Oberoi P, Kenten JH, Phillips AC, Weissman AM and Vousden KH (2005) Small molecule inhibitors of HDM2 ubiquitin ligase activity stabilize and activate p53 in cells. Cancer Cell 7: 547–559.
Soengas MS, Alarcon RM, Yoshida H, Giaccia AJ, Hakem R, Mak TW and Lowe SW (1999) Apaf-1 and caspase 9 in p53-dependent apoptosis and tumor inhibition. Science 284: 156–159.
Moroni MC, Hickman ES, Lazzerini Denchi E, Caprara G, Colli E, Cocconi F, Muller H and Helin K (2001) Apaf-1 is a transcriptional target for E2F and p53. Nat. Cell Biol. 3: 552–558.
Jeffers JR, Parganas E, Lee Y, Yang C, Wang J, Brennan J, MacLean KH, Han J, Chittenden T, Ihle JN, McKinnon PJ, Cleveland JL and Zambetti GP (2003) Puma is an essential mediator of p53 dependent and independent apoptotic pathways. Cancer Cell 4: 321–328.
Michalak E, Villunger A, Erlacher M and Strasser A (2005) Death squads enlisted by the tumor suppressor p53. Biochem. Biophys. Res. Commun. 331: 786–798.
Godefroy N, Lemaire C, Renaud F, Rincheval V, Perez S, Parvu-Ferecatu I, Mignotte B and Vayssière JL (2004) p53 can promote mitochondria and caspase independent apoptosis. Cell Death Differ. 11: 785–787.
Acknowledgements
We thank the technical staffs of TNO-Biomedical Research, Dieter Defever, Ines Bonk and Riet De Rycke for their excellent technical assistance. We thank Guillermina Lozano for the mdm2 and mdm4 conditional knockout mice. This work was supported by Public Health Service Grant 2000B051 from the Netherlands Heart Foundation and by Grants 902-26-242 and 912-02-03 from NOW/ZonMw. The research of BJM van Vlijmen has been made possible by a fellowship of the Royal Netherlands Academy of Arts and Sciences. S Francoz was supported by grants from ‘Télévie (FNRS)’. This work was supported in part by grants from Association for International Cancer Research (J-C Marine), Belgium Federation against Cancer (non-profit organization; J-C Marine) and by EC FP6 funding (AG Jochemsen and J-C Marine). This publication reflects only author's views. The commission is not liable for any use that may be made of the information herein.
Author information
Authors and Affiliations
Corresponding author
Additional information
Edited by A Villunger
Rights and permissions
About this article
Cite this article
Boesten, L., Zadelaar, S., De Clercq, S. et al. Mdm2, but not Mdm4, protects terminally differentiated smooth muscle cells from p53-mediated caspase-3-independent cell death. Cell Death Differ 13, 2089–2098 (2006). https://doi.org/10.1038/sj.cdd.4401973
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.cdd.4401973
Keywords
This article is cited by
-
Tau protein binds to the P53 E3 ubiquitin ligase MDM2
Scientific Reports (2023)
-
Relevance of the p53–MDM2 axis to aging
Cell Death & Differentiation (2018)
-
Indispensable role of Mdm2/p53 interaction during the embryonic and postnatal inner ear development
Scientific Reports (2017)
-
Apoptosis of Sertoli cells after conditional ablation of murine double minute 2 (Mdm2) gene is p53-dependent and results in male sterility
Cell Death & Differentiation (2016)
-
β-Catenin C-terminal signals suppress p53 and are essential for artery formation
Nature Communications (2016)