Abstract
Mutations of the myelin proteolipid protein gene (Plp) are associated with excessive programmed cell death (PCD) of oligodendrocytes. We show for the first time that PLP is a molecule ubiquitously expressed in non-neural tissues during normal development, and that the level of native PLP modulates the level of PCD. We analyze three non-neural tissues, and show that native PLP is expressed in trophoblasts, spermatogonia, and cells of interdigital webbing. The non-neural cells that express high levels of native PLP also undergo PCD. The level of PLP expression modulates the level of PCD because mice that overexpress native PLP have increased PCD and mice deficient in PLP have decreased PCD. We show that overexpression of native PLP causes a dramatic acidification of extracellular fluid that, in turn, causes increased PCD. These studies show that the level of native PLP modulates the amount of PCD during normal development via a pH-dependent mechanism.
Similar content being viewed by others
Introduction
Central nervous system (CNS) myelin proteins are perceived as a group of proteins unique to the nervous system that form and maintain myelin. Proteolipid protein (Plp) and DM20,1 an alternative splice product of the PLP gene, constitute roughly 50% of all CNS myelin proteins.2 PLP is the most extensively studied of all CNS myelin proteins, partly because animals and humans have more PLP mutations than all other CNS myelin genes combined. The hallmark of PLP mutations is abnormal myelin formation that ranges from nearly complete dysmyelination to demyelination.3, 4 Apoptosis of oligodendrocytes (Olgs) is extensive in many mutants that have point mutations or duplications of the native gene.5, 6, 7 It is postulated that Olg apoptosis in the PLP mutants is a primary defect that contributes to myelin deficits and, in turn, leads to seizures and death.8 However, Olg death in vivo might be a secondary phenomenon due to the cells' inability to assemble and maintain myelin sheaths. Tissue culture studies support the hypothesis that apoptosis of Olgs is a primary function of the PLP gene when it is misexpressed. Immortalized Olgs or non-Olg cells that overexpress native PLP or jimpy PLP (a mutant with a missense mutation) have dramatically shortened survival times compared to cells overexpressing other membrane proteins.9 In contrast, Olgs derived from PLP-deficient mice have lengthened survival times.10 These data show that the level of native as well as mutant PLP modulates Olg survival in culture but still does not resolve the question of whether PLP directly modulates apoptosis in vivo.
PLP also acts in a paracrine mode to regulate neuronal survival. Coculture of neurons with nonglial cell lines that overexpress native PLP but not DM20 shortens survival of neurons.11 This finding is mimicked in vivo where either modest overexpression of native PLP or its absence leads to axonal abnormalities and neuronal death.12, 13, 14 Thus, regulation of the level of native PLP in vivo is critical as deviation from baseline levels leads to both glial and neuronal abnormalities. We show for the first time that PLP is expressed in non-neural cells during normal development, and that its level of expression correlates with the amount of apoptosis.
Results
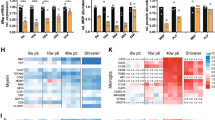
As PLP/DM20 is expressed at low levels in Schwann cells of peripheral nerves of many organs, we examined PLP expression in several tissues that lack nerves: blastocysts, interdigital webbing, and seminiferous tubules. A series of primer sets (Figure 1a) designed to amplify classical PLP/DM20 (Materials and Methods) confirm previous reports that Plp is more abundant than DM20 in the adult mouse brain. DM20 and PLP transcripts are expressed in embryonic day (E) 4.5 blastocysts and in a human trophoblast cell line15 (Figure 1b). Bands in preimplantation embryos are fainter than in the brain, but the embryonic cDNA loaded per lane is approximately three-fold less than the brain. PLP and DM20 bands cut from the blastocysts and brain were sequenced and sequences match published databases for classical PLP. Western blots using an antibody that recognizes PLP but not DM2016 detect PLP protein in the human trophoblast cell line, as well as the brain and myelin of wild-type mice but not in PLP-deficient transgenic mice (Figure 1c). Immunostaining of human trophoblasts in culture using a PLP-specific16 or PLP/DM20 affinity purified poly- and monoclonal antibodies17, 18 shows weak staining in flattened cells but stronger staining in cells bearing migratory processes (unpublished data).
(a) Scheme of PLP exons and primer sets used to detect classical PLP/DM20 transcripts. (b) 2 × 6 primers show 600 bp products for PLP and 500 bp products for DM20 in the adult brain, 4.5 day blastocysts, and human trophoblast line.15 Total RNA loaded for blastocysts is approximately that of brain.51, 52 (c) Homogenates prepared from the brain and myelin fractions of wild-type, PLP-deficient mice (−PLPtg), and human trophoblasts, cultured for 3 days probed with a polyclonal antibody that recognizes PLP but not DM20 isoform16 show the appropriate band in the brain and myelin of wild-type mice and human trophoblasts15 but not in −PLPtg mice
A nonradioactive in situ hybridization technique routinely employed in our laboratory to localize glial messages in the brain shows PLP/DM20 in cells on the blastocyst surface (Figure 2a and b). PLP/DM20 transcripts in single cells range from intensely positive to negative. Blastocysts incubated without probe or incubated with digoxigenin (DIG)-labeled plasmid DNA were negative (Figure 2c). Immunocytochemistry using characterized, affinity purified poly- and monoclonal antibodies that recognize PLP/DM20 show high levels of expression in subpopulations of trophoblasts (Figure 2d). Interestingly, trophoblasts ‘hatching’ from the zona pellucida and located at the abembryonic pole of the blastocyst are strongly labeled for PLP/DM20. Double labeling with 4′-6 diamidino-2-phenylindole (DAPI) shows many but not all nuclei in the ‘hatched’ blastocysts have a normal chromatin appearance, indicating their viability, whereas others appear pyknotic (Figure 2e). Both fluorescent and peroxidase immunostaining of blastocysts show plasma and nuclear membrane staining for PLP/DM20 (Figure 2g and h). Blastocysts immunostained for PLP/DM20 often have unstained spherical bodies within cells that appear to be nuclei, but they do not colocalize with DAPI stained nuclei (Figure 2g). These unstained structures are probably vacuoles and/or lipid droplets that are abundant in blastocyst cells.19 Confocal images show that some cells at the surface (trophoblast) and within the center of the blastocyst (inner cell mass) are unstained for PLP/DM20 (Figure 2i).
(a and b) PLP/DM20 in situ hybridization23 of 4.5 day blastocysts shows PLP/DM20 messages in the cytoplasm of cells located on the surface of blastocyst. Different focal planes of same blastocyst show two intensely stained cells (arrows); many cells have moderate amounts of staining; other cells show no staining (arrowheads). (c) In the same experiment, blastocysts incubated without probe show no staining. (d and g) A blastocyst harvested at 4.5 days and cultured for 2 days immunostained with an affinity purified polyclonal PLP antibody. The small sphere contains trophoblast cells ‘hatching’ through the zona pellucida, many of which are intensely stained. The embryonic half of the blastocyst, containing a blastocoele (arrow), has both brightly and moderately stained cells. (e) DAPI staining shows some cells in both parts of blastocyst have a normal chromatin pattern while others are condensed. (f) Blastocysts incubated without primary antibody lack specific staining. (g) Higher magnification of blastocysts immunostained for PLP and DAPI (h). The bar in (g) and (h) is the same length to show that the nuclear membrane (arrow) is stained as well as the plasma membrane (crossed arrow). (i) Confocal microscopy of a 4.5 day blastocyst immunostained for PLP/DM20 and counterstained for DAPI shows some cells on the surface as well as cells within the blastocyst are unstained (arrows). Other cells (crossed arrow) show strong cytoplasmic staining throughout the cytoplasm. (j) In this whole blastocyst, a cluster of brightly fluorescent cells is stained with an affinity purified antibody directed against the carboxy terminus of PLP. The cluster also contains many TUNEL-labeled cells (k) and cells with condensed nuclei (l). (m) Confocal microscopy shows two brightly fluorescent PLP/DM20+ cells (arrows) that are TUNEL labeled (n). (o) These PLP/TUNEL+ cells have condensed chromatin typical of dying cells (arrows). Bar=25 μm (a–f, j–n)=2.5 μm (g–i, o)
Cells intensely stained for PLP/DM20 in blastocysts are frequently TUNEL labeled and/or show a condensed chromatin pattern characteristic of dying cells (Figure 2j–o). Most TUNEL-labeled cells line the surface of the blastocyst while others are located in the inner cell mass. These TUNEL-labeled cells in the inner cell mass are usually not PLP/DM20+, indicating that not all apoptotic cells are necessarily PLP/DM20+.
We next asked whether the level of PLP/DM20 gene expression correlates with the level of apoptosis by examining postnatal testes of transgenics overexpressing native PLP (+PLPtg),20 transgenic PLP-deficient mice (−PLPtg),21 and jimpy mice (Figures 3, 4 and 5). The testes and brains of PLP overexpressing mice contain approximately 30% more PLP messages than wild-type mice based on quantitative RT-PCR of gels; more recently, using real-time PCR, we find 64 and 60% increases in testes and the cerebellum, respectively. These results are further confirmation that PLP is present in the testes and is transcriptionally upregulated (Figure 3a). PLP/DM20 message is localized in spermatogonia near the basal lamina of a few seminiferous tubules (Figure 3b and c). Incubation of the testes with plasmids containing DIG-labeled cDNAs such as pBR322 or no probe show no staining (Figure 3d). PLP/DM20 protein is similarly localized to spermatogonia located near the basal lamina (Figure 3e and f), a pattern that is strikingly similar to the distribution of apoptotic cells in mouse testes.22 This pattern suggests that spermatogonia expressing high levels of the Plp gene are apoptotic cells. To colocalize PLP/DM20 messages in apoptotic cells, 50 μm Vibratomed sections are first processed for TUNEL staining and then for PLP in situ hybridization.23 In at least 70% of the TUNEL-labeled cells, the PLP message is present in the cytoplasm and/or nuclear remnants (Figure 4a and b). The localization of the message and fragmented DNA in some double-labeled cells are in distinct cellular compartments: the message is aggregated in the cytoplasm near the plasma and nuclear membrane, whereas internucleosomal fragments are limited to the degenerating nucleus. Some cells show only blue reaction product indicating expression of PLP/DM20 messages (Figure 4c) or only brown reaction product indicating internucleosomal degradation (Figure 4d). The separate and appropriate location of the message and DNA fragments in the cytoplasm and nucleus demonstrates that the DIG-labeled probes do not artifactually bind to the DNA internucleosomal fragments.
(a) RT-PCR of 30-day wild-type and +PLPtg brains and testes from four different animals. DM20 transcripts are more abundant than PLP transcripts in testes of wild and +PLPtg mice. (b and c) DIG-labeled PLP/DM20 cDNA used for in situ hybridization of testes in 50 μm Vibratomed sections.23 (b) At low magnification, PLP/DM20 message is only found in certain seminiferous tubules (arrows). (c) PLP/DM20 gene expression is typically found in clusters of cells (arrows) located near the basal side of the seminiferous tubule. PLP/DM20 message is localized to the periphery in most cells but present throughout the cytoplasm of other cells. (d) Section of testes processed for in situ hybridization lacking DIG-labeled probe. (e) PLP/DM20 immunocytochemistry of testes using a monoclonal antibody directed against PLP/DM20 shows reaction product in cells (arrows) located adjacent to the basal region of the seminiferous tubules. Occasionally, cells in the apical region of the tubule are stained (arrowhead). (f) Using affinity purified polyclonal antibodies to PLP/DM20, immunostained cells are located adjacent to the basal lamina. (g) Specific staining is absent when primary antibody is eliminated from immunocytochemistry. Bar=250 μm (b) =25 μm (c–e)
PLP/DM20 in situ hybridization combined with TUNEL staining of testes.23 In situ hybridization and TUNEL staining are first performed on 50 μm sections, the tissue embedded in plastic, and resectioned at 2 μm because it is difficult to discriminate clearly the NBT/BCIP blue message from the DAB brown protein product when both colors are colocalized in thick sections. All pictures are from 2 μm plastic sections of the same tissue block. (a) Three TUNEL-labeled nuclei (N) visualized in brown are located in cells that express PLP/DM20 mRNA visualized in blue. PLP/DM20 message is concentrated in and around the nucleus (arrow), and also near the plasma membrane (crossed arrow). Many cells adjacent to labeled cells show no reaction product (asterisks). (b) A PLP/DM20 mRNA+ cell with strong reaction product in cytoplasm (arrow) surrounds a TUNEL+ nucleus (N). (c) A PLP/DM20+ mRNA cell exhibits strong cytoplasmic staining but nucleus (N) is TUNEL negative. A faint blue reaction product is often found at the periphery of spermatogonia (crossed arrow) but whether this color is background or low level of PLP/DM20 message is unknown. (d) A TUNEL+ cell (N) stains the spindle apparatus but shows no PLP/DM20 reaction product. Bar=2.5 μm
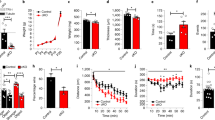
(a) Quantification of the number of cleaved caspase-3+ cells in testes (Tes) and spinal cord (SC) of jimpy (jp), PLP overexpressors (+PLPtg) and PLP-deficient mice (−PLPtg).16 Capase-3+ cells are increased in jimpy and +PLPtg mice but decreased in −PLPtg mice compared to control C57BL6 for jimpy, for +PLPtg, and C57BL6 for −PLPtg mice. *P<0.05, **P<0.01, ***P<0.0001, compared to wild-type, unpaired t-test. (b and c) All caspase-3+ cells are brightly stained in testes of jp (b) and WT mice (c) permitting accurate counting
To show that the level of PLP gene expression affects the number of apoptotic cells in non-neural cells, we quantified the number of cleaved caspase-3+ cells in the testes of normal and mutant mice. Jimpy and +PLP tg mice were sacrificed at 15 and 30 days, respectively, several days after each mutant developed tremors, but before they developed seizures. +PLPtg mice exhibit 633 and 200% increases in cleaved caspase-3+ cells in testes and the spinal cord white matter, respectively, and jimpy mice exhibit 52 and 1120% increases in the testes and cord (Figure 5a). In contrast, −PLPtg mice exhibit 67 and 17% decreases in cleaved caspase-3+ cells in the testes and spinal cord white matter, respectively (Figure 6a). The raw data show that the number of cleaved caspase-3+ cells in the testes decreases with age, whereas their numbers remain approximately the same in spinal cord white matter (Figure 5a). Antibody against the active form of caspase-3 brightly stains clusters of spermatogonia near the basal lamina, similar to the staining pattern for PLP message, protein, and TUNEL+ cells (Figure 5b and c). We also measured levels of total and cleaved poly(ADP-ribose) polymerase-1 (PARP), a downstream target of caspase-3 in the spinal cord and testes on Westerns.24 In +PLPtg mice, total PARP expression is significantly increased by 50% in the testes; in jimpy mice, total PARP is increased by 15% (unpublished data).
(a) RT-PCR of adult (A) and embryonic (E) (E-14.5) tissues using two different sets of primers specific for PLP/DM20. Embryonic tissue was pooled from fore and hind digits (Digits), from skin covering the digits and webbing (Dig/skin), brain, intestines, and liver. Adult tissue was isolated from the brain, lens, and bladder. DM20 transcripts are more abundant than PLP transcripts in the embryonic brain whereas PLP transcripts are more abundant in the adult brain. (b) PLP/DM20 DIG-labeled antisense riboprobe shows that only Olgs in the corpus callosum and septum of the adult brain are stained. (c, f and g) In situ hybridization with PLP/DM20 antisense riboprobe in an E-14.5 day mouse paw. Reaction product is localized to skin and interdigital webbing (arrows in c and f). (d) In situ hybridization with sense PLP riboprobe shows no specific staining. (e) In situ hybridization using a DIG-labeled full-length cDNA for PLP/DM2023 shows similar staining pattern to that of the antisense riboprobe. (g) Higher magnifications with PLP antisense riboprobe show the reaction product in the skin of digits and webbing (arrows). Cytoplasmic granules are strongly reactive but the nucleus is unstained. (h) The skin of digits and webbing are immunostained with a polyclonal antibody specific for PLP but not DM20 (arrows). (i) TUNEL staining of E-14.5 day mouse paw shows apoptotic cells located principally in the interdigital webbing. (j) TUNEL-labeled cells in webbing exhibit typical features of apoptotic cells. (k) Cleaved caspase-3 immunocytochemistry of E-14.5 day mouse paw shows labeling in skin, webbing, and between phalanges (arrows). (l) Double immunocytochemistry with cleaved caspase-3 (blue) and a polyclonal antibody specific for PLP (brown) shows double-labeled cells (arrow). Bar=250 μm (b–e, h, i, k); =25 μm (f, g and j,l)
To further test the hypothesis that PLP modulates apoptosis in non-neural cells, we examined interdigital webbing of embryonic mice as this transient structure lacks nerve fibers and undergoes massive cell death over several days.25 Embryonic brain expresses more DM20 than PLP message in keeping with previous observations (Figure 6a).26 Embryonic digits, intestines, and the liver also express more DM20 transcripts than PLP. The skin from digits and interdigital webbing was stripped away from the digits in these preparations, showing that PLP/DM20 messages are comparable to that of the E14 brain. Adult lens and bladder were used as negative controls on the assumption that they lack PLP/DM20 and serve as negative controls. Surprisingly, the lens expresses only DM20 transcripts and the bladder expresses both PLP and DM20 transcripts at levels comparable to the brain. Localization of PLP/DM20 message and protein in these two structures is unknown.
In situ hybridization of brain tissue with antisense riboprobes or full-length PLP cDNAs labeled with DIG stain only Olgs (Figure 6b), confirming the specificity of the probes. Neurons in the cerebrum and cerebellum are not labeled, even with extended colorimetric development, indicating that somal restricted PLP isoform is not labeled.27 DIG-labeled antisense riboprobes (Figure 6b, c, f and g) or full-length PLP/DM20 cDNAs (Figure 6e) reveal abundant transcript levels within interdigital webbing and skin. PLP/DM20 transcripts are localized to granular structures in the cytoplasm but not in the nuclei of skin cells (Figure 6g). A PLP antibody localizes to the interdigital webbing that generally matches the site of PLP/DM20 message (Figure 6h). TUNEL staining and caspase-3 immunocytochemistry confirm that the webbing undergoes cell death at the same time PLP/DM20 transcripts are present (Figure 6i–k). Many, but not all, PLP+ cells are caspase-3+ (Figure 6l) in the skin. Interestingly, a stripe of cells located between the phalanges also expresses PLP transcripts and are apoptotic (Figure 6e and k).
How does overexpression of native PLP cause increased programmed cell death (PCD)? In previous studies,11 we show that neurons cocultured with 293 cells stably expressing native PLP degenerate in a dose-dependent manner depending upon the number of 293 PLP expressing cells. The PLP but not DM20 expressing or control cells acidified the media, and culture of neurons with the acidified, conditioned media, or acidified media by itself was sufficient to cause increased neuronal death.11 To test whether overexpression of native PLP causes a similar acidification in vivo, we used a modification of a time lapse proton-flux assay,28, 29 which is used to measure the release of protons from synaptic and lipid vesicles and mitochondria.30, 31 The method is extremely reproducible and shows little variation from animal to animal. Cubes of brain tissue from wild-type and +PLPtg mice are placed into an artificial (a) cerebrospinal fluid (CSF) buffer32 that has low buffering capacity and is stable at room temperature for more than 15 min. A pH meter is coupled to a computer that samples pH every 0.5 s, and pH is expressed to 1 mpH (0.001) units. The details for the procedure, buffer, pH measurements, and controls are presented in the Materials and Methods.
The number of protons released into the buffer is first measured in nanomoles, and this value may be converted to the total number of protons (Figure 7a). The increase in protons released into the buffer at 15 min between +PLPtgs and wild-type mice is approximately 4 × 1013 (40 000 000 000 000). However, the increase between the +PLP overexpressors and controls occurs very quickly, and even at 2 min a pronounced difference is observed. A typical trace (Figure 7b) for one of these measurements shows virtually complete overlap between the two overexpressors. Since the volume of the extracellular space (ECS) and the brain is known,33 we can calculate the difference in the total number of protons in solution (see Materials and Methods). We estimate the pH of the ECS in the +PLPtg brain to be 5.6 compared to 7.2–7.3 for the wild-type brain.
(a) Bar graph shows the total number of protons released into the buffer from +PLPtg and control mice. N=5, 6 +PLPtg and 4, 5 control mice at 2, 5 min, and 10, 15 min, respectively. *P<0.05, unpaired t-test. (b) A scan from an experiment that shows early and sustained acidification of buffer by two PLPtg. The scans from the two overexpressors virtually overlie each other
Discussion
Detection of PLP/DM20 transcripts outside the nervous system has been shown in other studies,34, 35, 36, 37 but the existing belief of glial biologists is that Schwann cells or neural crest cells are the cell types that express these PLP transcripts. An exception to this opinion is the thymus where T cells and macrophages have been shown to express myelin genes.36, 37 We eliminated the possibility that myelin genes are expressed by neural crest or Schwann cells in non-neural tissues by examining blastocysts, interdigital webbing, and seminiferous tubules which are structures that lack nerves and display programmed cell death. We show, using high-resolution in situ hybridization and immunocytochemistry of perfused material, that PLP/DM20 is present in trophoblasts, cells of interdigital webbing, and spermatagonia.
Based on the proposed function of Plp as an adhesive molecule in myelin, the phenotypes exhibited by animals and humans deficient in Plp was unpredicted. The lifespan of Plp-deficient mice is normal;21, 38 likewise, myelin sheath formation proceeds normally with subtle abnormalities in the periodicity of the myelin sheaths.21, 38 Similarly in Pelizaeus–Merzbacher patients lacking PLP, CNS myelination is fairly normal and death does not begin until men are in their 30s.14, 38 In contrast, modest overexpression of the native proteolipid protein gene (PLP/Plp) in man and in transgenic mice is almost always lethal, with death in some males by their second year.39 Nearly a hundred Plp mutations have been described in animals almost all of which are lethal (http://www.med.wayne.edu/Neurology/plp.html). These phenotypes show that the absence of PLP/DM20 is preferable to its overexpression, prompting investigators to view Plp as a dominant-negative gene.
Importantly, although both PLP and DM20 isoforms are membrane proteins, the PLP isoform mainly confers toxicity to cells. In nonglial cell lines stably transfected to express PLP or DM20 in roughly equal amounts, only PLP was toxic to neurons.11 This observation is consistent with animal studies that show knockin mice, which overexpress DM20 but lack PLP, as having nearly normal lifespans.40 In transgenic mice containing two to as many as 17 copies of DM20 along with native PLP, mice do not die until 8–10 months and neuronal viability was not affected.41 In our study, the PLP isoform is detected in non-CNS apoptotic cells, suggesting PLP functions to modulate apoptosis.
The sequence of biological and behavioral events leading to death in PLP mutants is unclear, but mutant animals and humans develop tremors postnatally or during infancy that are usually followed by fatal seizures. The paucity of myelin is certainly a major factor contributing to behavioral deficits but is an incomplete explanation because certain rodents that have PLP mutations and virtually no CNS myelin have longer lifespans than some myelin-deficient rodents with reasonable numbers of myelinated fibers.42 Oligodendrocyte death is a common finding in animals that overexpress native Plp or have point mutations, but the extent to which Olg death contributes to the animals' death is still unclear.7, 8 Induction of the unfolded protein response (UPR) occurs in COS-7 cells engineered to express PLP point mutations,43 and accumulation of misfolded protein in the endoplasmic reticulum generally correlates with the decreased lifespan of these mutants.43 However, caspase-12, which is associated with a UPR is not upregulated in PLP overexpressing mice, suggesting that UPR is not involved in +PLPtg apoptosis.5 These studies leave unresolved the question of whether Olg apoptosis in vivo is a secondary effect due to improper myelination. The question can be resolved by studying whether non-neural cells in vivo express PLP, and if levels of native or mutant PLP affect levels of apoptosis. Our findings indicate that levels of native or expression of mutant PLP in non-neural cells correlates with levels of apoptosis. Taken together, previous in vitro and these in vivo analyses indicate that abnormally expressed PLP directly functions as a toxic molecule in Olgs.
In the CNS, PLP also functions in a paracrine manner as a toxic molecule to neurons. In vivo, modest overexpression of native PLP leads to axonal abnormalities and neuronal death in rodents and humans.12, 13 Coculture of neurons with non-neural cells stably transfected to express PLP but not DM20 increases the number of degenerating axons and neurons.11 Conditioned media derived from these PLP-expressing cells also induces apoptosis, indicating factor(s) in the media contribute to the neurodegeneration. We noticed that the PLP-expressing cells either cultured alone or cocultured with neurons acidified culture media, regardless of the type of media composition.11 Neurons cultured at pH 6.7 degenerated much more quickly than those cultured at pH 7.2, indicating acidic media is sufficient to accelerate neuronal death. To investigate whether overexpression of native PLP causes acidification of the ECS, we used an ex vivo proton-flux assay to measure the release of protons into a buffer.28, 29, 30 The assay is strikingly consistent in measuring the nanomolar concentration of H+ released from each +PLPtg and control mouse, and the data are statistically significant. Importantly, the increased acidification of the buffer from the +PLPtg brains was detectable within 2 min of adding the brain tissue to the buffer. This indicates that the vast majority of the increased protons in the buffer comes from diffusion of protons from the ECS. The pH differential between the overexpressors and controls continues at the same rate for the duration of the experiment, indicating release of protons from the ECS and/or from cytosol into the buffer. Assuming all the protons are derived from the ECS, we calculate the pH of the +PLPtg's ECS to be 5.67. While this pH value may be lower than that which occurs in vivo, the difference between the overexpressors and controls is so dramatic that it is certain to have biological significance. The acidification of media by the PLP overexpressors is directly relevant to understanding the cause of neural apoptosis because low extracellualar pH44 and/or cytosolic acidification facilitates apoptosis.45 The first step in cytosolic acidification is mitochondrial alkalinization and loss of the H+ gradient of the mitochondrial membrane that is followed by cytochrome c release, cytosolic acidification, and caspase activation.45 Activation of the apoptotic cascade, including several caspases, is much more efficient in the pH 6 range than in the pH 7 range.45 Since we are performing proton-flux analyses of brain tissues from PLP overexpressors, the acidification of these brain tissues is presumably present for days, weeks, and even until their death 2–3 months later. This persistent acidification of PLP transgenic brains must have deleterious effects upon neural respiration. Whether non-neural tissues are also acidified by overexpression of PLP in PLP transgenics and normal development is under investigation.
The results of many in vitro and in vivo studies show overexpression of native PLP is toxic to Olgs, neurons and, in the present study, to certain non-neural cells.5, 9, 11 If this is so, an intriguing question is how do Olgs in wild-type mammals synthesize large amounts of PLP and survive? Between birth and the peak of myelination, levels of PLP transcripts increase 20-fold but sharply decline in adults.46, 47, 48 At the same time as PLP is elevated in development, immature Olgs undergo massive cell death, estimated to be as high as 50%.49 Recently, we found all apoptotic Olgs have much higher levels of PLP than non-apoptotic Olgs during development. While PLP is traditionally viewed as an adhesion molecule involved in stabilization of myelin sheaths, our and other recent studies show that Plp has multiple functions, including signaling via its interaction with integrins.50 Our demonstration that native PLP modulates apoptosis in nonmyelinating cells shows that this gene has a role in normal development that is independent of myelination. In the CNS, native PLP modulates pH by causing acidification of the ECS. Acidification of ECS and cells facilitates apoptosis in cells. Whether native PLP acts by a similar mechanism in non-neural tissues remains to be determined but it is a likely candidate when abundantly expressed.
Materials and Methods
RT-PCR
Different primer sets designed to amplify PLP and DM20 gene products were tested on adult mouse brain cDNA reverse transcribed from total RNA extracts. The following two sets of primers produced the strongest bands: sense primer-6 (5′AGCAATAAACAGGTGGAAGGTC3′) in exon 6 and antisense primer-2 (5′GCTAATTGAGACCTATTTCTCC3′) in exon 2; and sense primer-1 (5′TTTCCGCGGATACATTCTGGCATC3′) in exon 4 and antisense primer-3 (5′ CGGGATCCTCAGAGTGCCAAAGACATG 3′) in exon 1, the last two primers (1 and 3) include 5′ restriction site sequences used for cloning. Recently, a new exon (1.1) in the Plp gene located between exons 1 and 2 has been identified. This protein is restricted to the cell bodies of neurons and Olgs.27 When this exon is spliced, it produces alternate proteins somal-restricted proteolipid protein1 (srPLP) and srDM20. The 2 × 6 primer set will not detect the ‘sr’ PLPs as the primers are located downstream of the new 1.1 exon. The 1 × 3 primer set could potentially detect ‘sr’ PLPs as well as the classical products. However, the srPLP and srDM20 products should be 109 bp's larger than the classical products.27 We detect a band around 650 as predicted for classical PLP but not a higher band around 760 for srPLP. The absence of higher bands in our RT-PCRs may be due to the low abundance of these isoforms or the 1 × 3 primer set does not amplify the srPLPs under the PCR conditions used.
Total RNA from C57Bl/6J mouse brains was isolated using Trizol reagent (Invitrogen Life Technologies, Carlsbad, CA, USA) according to the manufacturer's instructions. The concentration of RNA from the brain was determined by absorption at 260 nm. Contaminating DNA was removed by incubation at 37°C for 30 min with I U amplification grade DNase I (Life Technologies) in 10 mM Tris-HCl (pH 8.3), 50 mM KCl, 2.5 mM MgCl2, and 40 U of RNasin (5′–3′ Inc., Boulder, CO, USA). Total RNA was reverse transcribed as described.51, 52 cDNA equivalent to 100 ng total RNA from the adult brain was subjected to PCR amplification in 50 μl reaction mixture containing 20 mM Tris-HCl (pH 8.4), 50 mM KCl, 2.5 mM MgCl2, and 2.5 U Taq Polymerase (Life Technologies). After the addition of 0.25 mM sense and antisense primers, PCR was started with a hot start at 95°C for 2 min, and continued for 40 cycles, each consisting of denaturation at 95°C for 30 s, annealing at 57°C for 1 min and extension at 72°C for 1 min. A final extension was conducted at 72°C for 7 min, and the PCR product electrophoresed. DNA ladder (100 bp) (Life Technologies) was also run in parallel lanes. DNA bands were visualized with ethidium bromide staining. The bands were cut from the gel, and DNA purified using QIAEX II gel extraction kit (QIAGEN Inc., Valencia, CA, USA). Nucleotide sequences were obtained using Wayne State University Nucleic Acid Core facility and compared against known sequences to confirm their identity. The bands in the embryos are not amplified genomic DNA because the primers span several introns that would produce very large bands, and because RNA is DNAse digested.
Animals
+PLPtg mice (line 66 homozygous mice)20 that have at least three extra copies of the native PLP gene3 were used in these studies. Real-time PCR performed in our lab confirmed that four to 12 copies of PLP gene were present in these mice.−PLPtg mice21 were obtained by mating homozygous −PLPtg mice, and confirmed by the absence of PLP signal in the brain after in situ hybridization and PLP immunocytochemistry or by genotyping tail-clipped DNA using PLP primers. Jimpy Tabby mice (jpTa/Y), or crossovers were (jp+/Y) obtained by mating jpTa and Ta−/Y (Charles River Laboratories Inc., Wilmington, MA, USA).
Immunocytochemistry
For immunocytochemistry, blastocysts were rinsed in 0.1 M phosphate-buffered saline (PBS) containing 2 mg/ml bovine serum albumin (BSA) (PBS/BSA) for 5 min. They were fixed with 4% paraformaldehyde in 0.1 M PBS at 4°C for 30 min, rinsed twice in PBS/BSA for 5 min, treated with 0.25% Triton X-100 in PBS/BSA at room temperature (RT) for 30 min, rinsed in PBS/BSA for 5 min, then treated with 10% goat or 10% mouse serum in PBS/BSA at RT for 30 min, and rinsed in PBS/BSA for 5 min. They were incubated with rabbit polyclonal antibodies directed against the PLP-specific peptides 109–128,16 as well as affinity purified rabbit polyclonal anti-PLP/DM20 directed against peptides 269–27617 diluted 1 : 50 or 1 : 100. A mouse monoclonal anti-PLP/DM20 AA3 supernatant that recognizes the carboxy-terminus18 was used 1 : 2 in a solution containing 0.1% Triton X-100 at 4°C for 16–20 h on blastocysts, or blastocysts containing 0.1% Triton X-100 in PBS/BSA at 4°C for 16–20 h were used as controls. The following morning the blastocysts were rinsed 2 × in PBS/BSA for 5 min, incubated in goat anti-rabbit IgG rhodamine (Boehringer Mannheim, Indianapolis, IN, USA) at a dilution of 1 : 200 or 1 : 400, goat anti-mouse IgG rhodamine (Boehringer Mannheim) at a dilution of 1 : 200 and DAPI (1 ng/ml) for 1 h at RT, 2 × rinsed in PBS/BSA for 5 min and mounted with Aquamount (Polysciences Inc., Warrington, PA, USA) mounting media. Blastocysts were photographed on a Leica fluorescent inverted microscope (Bannockburn, IL, USA) or a Leica Laborlux 12 fluorescent microscope equipped with a Nikon camera system. Slices (1 μm) of whole blastocysts were imaged using a Zeiss LSM310 Confocal laser scanning microscope. Both antibody-treated and control preparations were photomicrographed, imaged, and printed using identical parameters.
A cleaved caspase-3 antibody (Cell Signaling Technology Inc., Beverly, MA, USA) was used to localize apoptotic cells with routine laboratory procedures5 Jimpy Tabby mice (jpTa/Y) or crossovers (jp+/Y) obtained by mating jpTa and Ta−/Y (Charles River) were compared to age-matched +Ta/Y or ++/Y littermates; +PLPtg are compared to age-matched B6CBA wild-type mice; −PLPtg are compared to age-matched B6CBA or B6129 wild-type mice. Cleaved caspase-3+ cells were counted in cross-sections of testes and numbers converted to number per sq/mm. Cleaved caspase-3+ cells were counted in transverse sections of white matter spinal cord. For each animal, three to four sections were counted, mean determined, and mean and S.E.M. obtained from at least six control and six mutant animals at each time point.
In situ hybridization
In situ hybridization for PLP/DM20 was performed using routine laboratory methods as described23 with the following modifications. Blastocysts were harvested as described above, but rinsed in PBS/BSA, zona pellucida removed, fixed in 4% paraformaldehyde at RT for 10 min, rinsed in Tris-HCl (pH 7.5) with BSA (1 mg/ml), placed in 70% ethanol (ETOH) at −20°C for 10 min and rinsed in 2 × sodium chloride, sodium citrate buffer (SSC) with BSA twice for 5 min.52 The blastocysts were then placed in prehybridization solution with BSA at 52°C for 10 min and then in hybridization solution containing a DIG-labeled PLP/DM20 full-length cDNA (courtesy of AT Campagnoni, UCLA) with BSA at 52°C for 14–16 h. The following day, blastocysts were washed in 2 × SSC/50% formamide at 37°C twice for 30 min, 1 × SSC/50% formamide at 37°C twice for 30 min, 1 × SSC at RT for 30 min, and 0.5 × SSC at RT for 30 min. The blastocysts were treated with S1 nuclease (Invitrogen Life Technologies) at 37°C for 15 min, rinsed in 0.1 M glycine, rinsed in dH2O, rinsed in Tris-HCl (pH 7.5) at RT twice for 30 min, incubated in anti-DIG conjugated to alkaline phosphatase 1 h 15 min, rinsed in Tris-HCl (pH 7.5) four times for 15 min and rinsed in Tris-HCl (pH 9.4). The detection of label was carried out using nitroblue tetrazolium/5-bromo-4-chloro-3-indolyl phosphate (NBT/BCIP) (Boehringer Mannheim) for 2 to 24 h. Blastocysts used for controls (B6CBAF1/J) were processed exactly as described above except that DIG-labeled probe was not added to hybridization buffer. Imaging of control blastocyst (Figure 2c) is identical to blastocyst treated with probe.
Antisense and sense PLP riboprobes were made from a partial cDNA BAS1013, courtesy of AT Campagnoni, UCLA, linearized with HindIII and transcribed from the vector's T7 promoter for the sense strand and linearized with BamHI and transcribed from T3 for the antisense strand. Sense and antisense strands were transcribed using DIG RNA labeling mix (Boehringer Mannheim), precipitated, dried, resuspended in hybridization buffer, and dot blotted to show DIG incorporation using NBT/BCIP detection.
Proton-flux assay
A modification of a time lapse proton-flux assay28, 29, 30, 31, 32 was used to measure H+ in the buffer from brains of male +PLPtg and male B6129 mice. Brains from wild-type and +PLPtgs are quickly dissected, cut into transverse slices, further sliced into 2 mm cubes, weighed (wt. for mutants and wt.=0.14 g), and placed into a vial containing 3 ml artificial cerebrospinal fluid (aCSF) at 26°C. The buffer is a modification of an aCSF33 that has a low buffering capacity and is stable at RT for more than 15 min. A pH meter is coupled to a computer that samples pH every 0.5 s. pH is measured to 1 mpH (0.001 pH) units.
The pH of the ECS may be calculated as follows. The volume of brain used in the aCSF is about 130 mm3, approximately of the total brain volume that is estimated to be 450 mm3, and the volume of ECS is 20% of the total brain volume or approximately 90 mm3.34 Accordingly, 26 mm3 of ECS was placed in the aCSF and the change in pH of the buffer was measured over a 15 min period. The difference in pH measured after 15 min between B6129 and +PLPtg was used to calculate the difference in total number of H+ atoms in solution by using the fact that pH=log (1/[H+]), where [H+] is the concentration in moles as there are 6.022 × 1023 atoms in a mole. Assuming that the pH of the 90 mm3 of ECS in a normal brain is 7.3,34 the total number of H+ in the ECS would therefore be 2.716 × 1012 atoms. Assuming the buffering capacity of the ECS is the same for both sets of mice, is at maximum buffering capacity, and adjusting for the fact that we have approximately of total ECS, we calculate the excess number of H+ in the +PLPtg compared to the B6129 animal to be 1.253 × 1014 atoms. Addition of these values yields a total number of H+ atoms in the ECS of a +PLPtg brain of 1.28 × 1014 atoms. Using this value, the pH of the ECS in a +PLPtg animal is calculated to be 5.627 as follows: the [H+] in the brain ECS=1/10pH=1/107.3=5.012 × 10−8 mol=50.12 nmol; the number of H+ in total ECS=5.012 × 10−8 mol/l × 6.022 × 1023 atoms/mol × 90 mm3 × cm3/1000 mm3 × l/1000 cm3=2.716 × 1012 H+. This is the difference in total number of H+ after 15 min in 3 ml of buffer between B6129 and +PLPtg as calculated from the average pH of the 3 ml of buffer 15 min after the brain pieces were added. ΔH+=25.81 × 1013 (+PLPtg)−22.19 × 1013 (B6129)=3.62 × 1013 H+ (in 26 mm3 of ECS). In 90 mm3, we have 3.62 × 1013 × (90/26)=1.253 × 1014 H+ (for entire ECS). Total number of H+ in +PLPtg brain= 2.716 × 1012+1.253 × 1014=1.28 × 1014 H+. Therefore, [H+] is: 1.28 × 1014 atoms × mol/6.022 × 1023 atoms × 1/90 mm3 × 1000 mm3/cm3 × 1000 cm3/l=2.362 × 10−6 mol and pH=log(1/[H+])=log (1/2.362 × 10−6)=5.6267.
Abbreviations
- aCSF:
-
artificial cerebrospinal fluid
- BSA:
-
bovine serum albumin
- CNS:
-
central nervous system
- CSF:
-
cerebrospinal fluid
- DAPI:
-
4′-6 Diamidino-2-phenylindole
- DIG:
-
digoxigenin
- E:
-
embryonic
- ECS:
-
extracellular space
- ETOH:
-
ethanol
- NBT/BCIP:
-
nitroblue tetrazolium/5-bromo-4-chloro-3-indolyl phosphate
- Olg:
-
oligodendrocyte
- PARP:
-
poly(ADP-ribose) polymerase-1
- PBS:
-
phosphate-buffered saline
- PCD:
-
programmed cell death
- PLP/Plp:
-
proteolipid protein
- S.E.M.:
-
standard error of the mean
- srPLP:
-
somal-restricted proteolipid protein
- SSC, sodium chloride:
-
sodium citrate buffer
- UPR:
-
unfolded protein response
- +PLPtg:
-
transgenic proteolipid protein overexpresser mouse
- −PLPtg:
-
transgenic proteolipid protein null mouse
References
The approved symbol for the classical myelin proteolipid protein gene is now Plp (modified 02.04) to distinguish it from the non-myelin proteolipid proteins. The RGD ID is 3354 and the MGI ID is 97623
Greenfield S, Norton WT and Morell P (1971) Quaking mouse: isolation and characterization of myelin protein. J. Neurochem. 18: 2119–2128
Anderson TJ, Klugmann M, Thomson CE, Schneider A, Readhead C, Nave KA and Griffiths IR (1999) Distinct phenotypes associated with increasing dosage of the PLP gene: implications for CMT1A due to PMP22 gene duplication. Ann. NY. Acad. Sci. 883: 234–246
Griffiths IR, Schneider A, Anderson J and Nave KA (1995) Transgenic and natural mouse models of proteolipid protein (PLP)-related dysmyelination and demyelination. Brain Pathol. 5: 275–281
Cerghet M, Bessert DA, Nave KA and Skoff RP (2001) Differential expression of apoptotic markers in jimpy and in Plp overexpressors: evidence for different apoptotic pathways. J. Neurocytol. 30: 841–855
Grinspan JB, Coulalaglou M, Beesley JS, Carpio DF and Scherer SS (1998) Maturation-dependent apoptotic cell death of oligodendrocytes in myelin-deficient rats. J. Neurosci. Res. 54: 623–634
Skoff RP (1995) Programmed cell death in the dysmyelinating mutants. Brain Pathol. 5: 283–288
Knapp PE, Skoff RP and Redstone DW (1986) Oligodendroglial cell death in jimpy mice: an explanation for the myelin deficit. J. Neurosci. 6: 2813–2822
Bongarzone ER, Jacobs E, Schonmann V, Kampf K, Campagnoni CW and Campagnoni AT (2001) Differential sensitivity in the survival of oligodendrocyte cell lines to overexpression of myelin proteolipid protein gene products. J. Neurosci. Res. 65: 485–492
Yang X and Skoff RP (1997) Proteolipid protein regulates the survival and differentiation of oligodendrocytes. J. Neurosci. 17: 2056–2070
Boucher SE, Cypher MA, Carlock LR and Skoff RP (2002) Proteolipid protein gene modulates viability and phenotype of neurons. J. Neurosci. 22: 1772–1783
Anderson TJ, Schneider A, Barrie JA, Klugmann M, McCulloch MC, Kirkham D, Kyriakides E, Nave KA and Griffiths IR (1998) Late-onset neurodegeneration in mice with increased dosage of the proteolipid protein gene. J. Comp. Neurol. 394: 506–519
Griffiths I, Klugmann M, Anderson T, Yool D, Thomson C, Schwab MH, Schneider A, Zimmermann F, McCulloch M, Nadon N and Nave KA (1998) Axonal swellings and degeneration in mice lacking the major proteolipid of myelin. Science 280: 1610–1613
Garbern JY, Yool DA, Moore GJ, Wilds IB, Faulk MW, Klugmann M, Nave KA, Sistermans EA, van der Knapp MS, Bird TD, Shy ME, Kamholz JA and Griffiths IR (2002) Patients lacking the major CNS myelin protein, proteolipid protein 1, develop length-dependent axonal degeneration in the absence of demyelination and inflammation. Brain 125: 551–561
Graham CH, Hawley TS, Hawley RG, MacDougall JR, Kerbel RS, Khoo N and Lala PK (1993) Establishment and characterization of first trimester human trophoblast cells with extended lifespan. Exp. Cell Res. 206: 204–211
Trifilieff E, Luu B, Nussbaum JL, Roussel G, Espinosa de los Monteros A, Sabatier JM and Van Rietschoten J (1986) A specific immunological probe for the major myelin proteolipid. Confirmation of a deletion in DM-20: FEBS Lett. 198: 235–239
Benjamins JA, Studzinski DM and Skoff RP (1994) Analysis of myelin proteolipid protein and F0 ATPase subunit 9 in normal and jimpy CNS. Neurochem. Res. 19: 1013–1022
Yamamura T, Konola JT, Wekerle H and Lees MB (1991) Monoclonal antibodies against myelin proteolipid protein: identification and characterization of two major determinants. J. Neurochem. 57: 1671–1680
Abe H, Otoi T, Tachikawa S, Yamashita S, Satoh T and Hoshi H (1999) Fine structure of bovine morulae and blastocysts in vivo and in vitro. Anat. Embryol. (Berl). 199: 519–527
Readhead C, Schneider A, Griffiths I and Nave KA (1994) Premature arrest of myelin formation in transgenic mice with increased proteolipid protein gene dosage. Neuron 12: 583–595
Klugmann M, Schwab MH, Puhlhofer A, Schneider A, Zimmermann F, Griffiths IR and Nave KA (1997) Assembly of CNS myelin in the absence of proteolipid protein. Neuron 18: 59–70
Kon Y, Horikoshi H and Endoh D (1999) Metaphase-specific cell death in meiotic spermatocytes in mice. Cell Tissue Res. 296: 359–369
Bessert DA and Skoff RP (1999) High-resolution in situ hybridization and TUNEL staining with free-floating brain sections. J. Histochem. Cytochem. 47: 693–702
D'Amours D, Germain M, Orth K, Dixit VM and Poirier GG (1998) Proteolysis of poly(ADP-ribose) polymerase by caspase 3: kinetics of cleavage of mono(ADP-ribosyl)ated and DNA-bound substrates. Radiat. Res. 150: 3–10
Dunker N, Schmitt K and Krieglstein K (2002) TGF-beta is required for programmed cell death in interdigital webs of the developing mouse limb. Mech. Dev. 113: 111–120
LeVine SM, Wong D and Macklin WB (1990) Developmental expression of proteolipid protein and DM20 mRNAs and proteins in the rat brain. Dev. Neurosci. 12: 235–250
Bongarzone ER, Campagnoni CW, Kampf K, Jacobs EC, Handley VW, Schonmann V and Campagnoni AT (1999) Identification of a new exon in the myelin proteolipid protein gene encoding novel protein isoforms that are restricted to the somata of oligodendrocytes and neurons. J. Neurosci. 19: 8349–8357
Schneider E and Altendorf K (1985) All three subunits are required for the reconstitution of an active proton channel (F0) of the Escherichia coli ATP synthase (F1F0). EMBO. J. 4: 515–518
Cao NJ, Brusilow WS, Tomashek JJ and Woodbury DJ (2001) Characterization of reconstituted F0 from wild-type Escherichia coli and identification of two other fluxes co-purifying with F0 . Cell Biochem. Biophys. 34: 305–320
Kelly ML and Woodbury DJ (1996) Ion channels from synaptic vesicle membrane fragments reconstituted into lipid bilayers. Biophys. J. 70: 2593–2599
Cuello LG, Romero JG, Cortes DM and Perozo E (1998) pH-dependent gating in the Streptomyces lividans K+ channel. Biochemistry 37: 3229–3236
Tong CK, Cammer W and Chesler M (2000) Activity-dependent pH shifts in hippocampal slices from normal and carbonic anhydrase II-deficient mice. Glia 31: 125–130
Nicholson C (1995) Extracellular space as the pathway for neuron–glial cell interaction. In Neuroglia, Kettenman H and Ransom BR (eds) (New York: Oxford University Press) pp. 387–397
Nadon NL, Miller S, Draeger K and Salvaggio M (1997) Myelin proteolipid DM20: evidence for function independent of myelination. Int. J. Dev. Neurosci. 15: 285–293
Campagnoni CW, Garbay B, Micevych P, Pribyl T, Kampf K, Handley VW and Campagnoni AT (1992) DM20 mRNA splice product of the myelin proteolipid protein gene is expressed in the murine heart. J. Neurosci. Res. 33: 148–155
Pribyl TM, Campagnoni C, Kampf K, Handley VW and Campagnoni AT (1996) The major myelin protein genes are expressed in the human thymus. J. Neurosci. Res. 45: 812–819
Pribyl TM, Campagnoni CW, Kampf K, Kashima T, Handley VW, McMahon J and Campagnoni AT (1996) Expression of the myelin proteolipid protein gene in the human fetal thymus. J. Neuroimmunol. 67: 125–130
Boison D and Stoffel W (1994) Disruption of the compacted myelin sheath of axons of the central nervous system in proteolipid protein-deficient mice. Proc. Natl. Acad. Sci. USA 91: 11709–11713
Woodward K and Malcolm S (1999) Proteolipid protein gene: Pelizaeus–Merzbacher disease in humans and neurodegeneration in mice. Trends Genet. 15: 125–128
Stecca B, Southwood CM, Gragerov A, Kelley KA, Friedrich Jr VL and Gow A (2000) The evolution of lipophilin genes from invertebrates to tetrapods: DM-20 cannot replace proteolipid protein in CNS myelin. J. Neurosci. 20: 4002–4010
Johnson RS, Roder JC and Riordan JR (1995) Over-expression of the DM-20 myelin proteolipid causes central nervous system demyelination in transgenic mice. J. Neurochem. 64: 967–976
Wolf MK, Nunnari JN and Billings-Gagliardi S (1999) Quaking*shiverer double-mutant mice survive for at least 100 days with no CNS myelin. Dev. Neurosci. 21: 483–490
Southwood CM, Garbern J, Jiang W and Gow A (2002) The unfolded protein response modulates disease severity in Pelizaeus–Merzbacher disease. Neuron 36: 585–596
Lee JL, Song JJ, Kim JH, Kim H-RC and Song YK (2004) Low extracellular pH augments TRAIL-induced apoptotic death through the mitochondria-mediated caspase signaling pathway. Exp. Cell Res. 293: 129–143
Matsuyama S, Liopis J, Deveraux QL, Tsien RY and Reed JC (2000) Changes in mitochondrial and cytosolic pH: early events that modulate caspase activiation during apoptosis. Nature Cell Biol. 2: 318–325
Gardinier MV and Macklin WB (1988) Myelin proteolipid protein gene expression in jimpy and jimpy(msd) mice. J. Neurochem. 51: 360–369
Kanfer J, Parenty M, Goujet-Zalc C, Monge M, Bernier L, Campagnoni AT, Dautigny A and Zalc B (1989) Developmental expression of myelin proteolipid, basic protein, and 2′,3′-cyclic nucleotide 3′-phosphodiesterase transcripts in different rat brain regions. J. Mol. Neurosci. 1: 39–46
Scherer SS, Vogelbacker HH and Kamholz J (1992) Axons modulate the expression of proteolipid protein in the CNS. J. Neurosci. Res. 32: 138–148
Barres BA, Hart IK, Coles HS, Burne JF, Voyvodic JT, Richardson WD and Raff MC (1992) Cell death and control of cell survival in the oligodendrocyte lineage. Cell 70: 31–46
Gudz TI, Schneider TE, Haas TA and Macklin WB (2002) Myelin proteolipid protein forms a complex with integrins and may participate in integrin receptor signaling in oligodendrocytes. J. Neurosci. 22: 7398–7407
Rout UK, Krawetz SA and Armant DR (1997) Ethanol-induced intracellular calcium mobilization rapidly alters gene expression in the mouse blastocyst. Cell Calcium 22: 463–474
Wang J, Rout UK, Bagchi IC and Armant DR (1998) Expression of calcitonin receptors in mouse preimplantation embryos and their function in the regulation of blastocyst differentiation by calcitonin. Development 125: 4293–4302
Acknowledgements
This research was supported by grants from the NIH and NMSS.
Author information
Authors and Affiliations
Corresponding author
Additional information
Edited by T Ferguson
Rights and permissions
About this article
Cite this article
Skoff, R., Bessert, D., Cerghet, M. et al. The myelin proteolipid protein gene modulates apoptosis in neural and non-neural tissues. Cell Death Differ 11, 1247–1257 (2004). https://doi.org/10.1038/sj.cdd.4401498
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.cdd.4401498
Keywords
This article is cited by
-
Neuronal loss in Pelizaeus–Merzbacher disease differs in various mutations of the proteolipid protein 1
Acta Neuropathologica (2009)