Abstract
Autophagic programmed cell death occurs during the development of diverse animal groups, but the mechanisms that control this genetically regulated form of cell killing are poorly understood. Genetic studies of bulk protein degradation in yeast have provided important advances in our understanding of autophagy, and recent investigations of Drosophila autophagic cell death suggest that some of these mechanisms may be conserved. In Drosophila, several steroid-regulated genes that encode transcription regulators are required for autophagic cell death. These transcription regulators appear to activate a large number of genes that play a more direct role in cell killing, including genes that function in apoptosis such as caspases. While caspase function is required for autophagic cell death during Drosophila development, genes encoding proteins that are similar to the yeast autophagy regulators are also induced in dying salivary glands. Furthermore, numerous noncaspase proteases, cytoplasmic organizing factors, signaling molecules, and unknown factors are expressed in interesting patterns during autophagic cell death. This article reviews the current knowledge of the regulation of autophagic programmed cell death during development of Drosophila.
Similar content being viewed by others
Introduction
Schweichel and Merker1 recognized that at least three forms of cell death occur during development of animals based on the location and role of lysosomes. While nonlysosomal cell death has not been commonly observed, apoptosis and autophagic programmed cell death are prominent in developing animals of diverse taxa.2 Apoptotic cells exhibit condensation of the nucleus and cytoplasm, fragmentation, and they are engulfed by phagocytes where the lysosome completes degradation of the dying cell.3 Unlike apoptosis, cells that die by autophagic cell death are degraded with little or no help from phagocytes and utilize lysosomes within the dying cell for degradation. Defects in apoptosis have been associated with a variety of human disorders including cancer,4 and studies of the mechanisms underlying apoptosis have been the focus of cell death research.5,6 In contrast, little is known about the mechanisms that regulate autophagic cell death.
The degradation of cytoplasmic organelles and proteins is an important component of proper cell life and death. While the ubiquitin/proteosome pathway is used to degrade many short-lived regulatory proteins,7 long-lived proteins are targeted to the lysosome by five mechanisms including direct transport, vacuolar import and degradation, endocytosis, microautophagy, and macroautophagy.8 Macroautophagy (hereafter refered to as autophagy) is the best known mechanism for sequestration of long-lived proteins and cytoplasmic components into the lysosomal compartment of the cell for degradation. The first descriptions of lysosomal structures that contained cytoplasmic components such as mitochondria were observed in rat livers perfused with glucagon9 and rat hepatomas.10 These autophagic structures have since been described in organisms that are as different as yeast and humans. During autophagy, cytoplasm and organelles are sequestered in double-membrane vesicles known as autophagic vacuoles in organisms such as insects and mammals and as autophagosomes in yeast11 (Figure 1). Although the origin of autophagic vacuole membranes are a subject of debate, evidence indicates that they are derived from the rough endoplasmic reticulum.12 Autophagic vacuoles then dock against the lysosomal vacuole, and fusion of the outer autophagic vacuole membrane with the lysosomal membrane releases the inner membrane-bound cytoplasmic contents into the lysosome where degradation occurs.
Diagram of the steps in autophagy. Autophagy is induced by starvation in yeast and cell death signals such as steroids during autophagic cell death in Drosophila. Induction results in the formation of a double-membraned autophagic vacuole that encloses cytoplasmic components including organelles such as mitochondria. The autophagic vacuole then docks against the lysosome, and the fusion of the outer autophagic vacuole membrane with the lysosome releases the inner membrane-bound cytoplasmic components into the lysosome where degradation occurs. Redrawn after Klionsky and Emr11
Autophagic structures were first associated with developmental programmed cell death in insects13,14 and chickens,15 and it is now clear that autophagic cell death occurs during development of diverse organisms including humans.2,16,17,18 The recent association of autophagic cell death with models of tumorigenesis19,20,21,22 and neurodegenerative disorders23,24,25,26 has led to greater interest in the mechanisms that regulate this form of cell destruction.
Lessons from yeast
The molecular genetic basis of protein and organelle turnover by autophagy has been well characterized in the yeast Saccharomyces cerevisiae. In yeast, autophagy is induced under nutrient-limiting conditions. Genetic screens for yeast that either lacked accumulation of autophagic structures or exhibited defects in the degradation of cytoplasmic proteins under starvation conditions resulted in the identification of the apg27 and aut28 mutants. In addition, the isolation of altered cytoplasm to vacuole-targeting mutants (cvt) in screens of yeast for defects in proteolytic processing of aminopeptidase I29 led to the recognition of the genetic overlap between the apg, aut, and cvt pathways.30 In total, 19 apg, aut, and cvt genes have been identified that regulate the induction, formation, size, and breakdown steps in yeast autophagy (reviewed in Ohsumi8 and Kilonsky and Emr11).
Genetic and biochemical studies of yeast indicate that the apg, aut, and cvt genes have distinct and related functions. While the role of some of these genes remains mysterious, two related ubiquitin-like systems are required for yeast autophagy. APG12 and AUT7/APG8/CVT5 encode proteins that show no apparent homology to ubiquitin, but both of these proteins appear to function in ubiquitin-like conjugation systems.31,32 APG12 and AUT7/APG8/CVT5 are both activated by the E1-like protein APG7/CVT2.33,34 APG12 is then conjugated to the E2-like protein APG10,35 while AUT7/APG8/CVT5 becomes conjugated to the E2-like protein AUT1/APG3.31 APG12 is then conjugated to APG532, and AUT7/APG8/CVT5 is attached to the lipid phosphatidylethanolamine.31 Mutations in either of these pathways prevent the proper formation of autophagic vacuoles,27 and indicate that ubiquitin-like systems are involved in the bulk degradation of long-lived cytoplasmic proteins. While it is not clear if the genes that are required for yeast autophagy are also required for the formation of autophagic structures during programmed cell death, recent studies in Drosophila suggest similarity in the mechanisms of cytoplasmic degradation.36
Activation of autophagic cell death in Drosophila
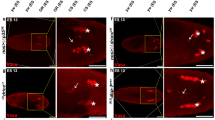
The steroid hormone 20-hydroxyecdysone (ecdysone) titer begins to rise 10 h after puparium formation, and reaches its peak level 12 h after puparium formation when it triggers the synchronous death of larval salivary gland cells.37,38 This cell death is preceded by markers of apoptosis including DNA fragmentation and nuclear acridine orange staining.39 While these data suggest that salivary glands die by apoptosis, their morphology indicates that they die by autophagy (Figure 2). At 1 h after this peak in ecdysone titer (13 h after puparium formation), salivary gland cells become round in shape, large cytoplasmic vacuoles appear to fragment, and a second class of vacuoles appears that is associated with the plasma membrane.40 By 14 h after puparium formation, salivary glands possess autophagic vacuoles that contain cytoplasmic structures including mitochondria.40 The cellular changes that occur following the rise in ecdysone titer are accompanied by changes in the tubulin and actin cytoskeleton, and accumulation of acid phosphatase activity.41 Salivary glands are destroyed 16-h following puparium formation. These changes in cell morphology indicate that the production and movement of vacuoles, reorganization of the cytoplasm and cell shape, and cytoplasmic degradation by proteases may all be involved in the autophagic destruction of salivary glands.
Steroid regulation of autophagic programmed cell death in Drosophila salivary glands. At 10 h after puparium formation, the ecdysone titer starts to rise peaking 12 h after puparium formation. This pulse of steroid triggers the autophagic destruction of salivary glands, and this process is complete 16 h after puparium formation. Following the peak in ecdyone, cells become round in shape, large eosin-positive vacuoles (V) appear to fragment, and a new class of eosin-negative vacuoles (*) appear along the plasma membrane, and the nuclei (N) remains intact even though DNA fragmentation has taken place. At 14 hours after puparium formation, autophagic vacuoles containing mitochondria are present in the cytoplasm of salivary gland cells (arrows). Data reproduced from Lee and Baehrecke40
Drosophila salivary glands have served as a long-standing model for studies of steroid signaling.42,43,44 Ecdysone activates salivary gland cell death through a transcriptional regulatory hierarchy (Figure 3). The ecdysone receptor is encoded by the nuclear receptor genes EcR and usp.45,46,47,48,49,50 The ecdysone receptor complex, and the nuclear receptor competence factor βFTZ-F1, activate transcription of the early genes BR-C, E74A, and E93.51,52 Salivary glands are not destroyed in animals with mutations in βFTZ-F1, BR-C, E74A, and E93.52,53,54,55 While βFTZ-F1 and E93 mutants exhibit defects in vacuolar changes during salivary gland autophagic cell death, BR-C and E74A mutant salivary glands progress to a very late stage in cell death even though complete destruction is prevented.40 In addition, E93 mutants exhibit similar defects in the formation of autophagic vacuoles in dying midgut cells,56 indicating that E93 is required for autophagy in dying Drosophila cells. BR-C, E74A, and E93 all encode proteins that regulate transcription of target late genes,54,55,57,58 and these downstream genes appear to function more directly in programmed cell death.
Model for genetic regulation of autophagic programmed cell death in Drosophila salivary glands. At 10 h after puparium formation, the ecdysone titer rises and this steroid is bound by the receptor complex that is encoded by EcR and usp. The ecdysone receptor complex and βFTZ-F1 activate the BR-C, E74A, and E93 early genes. BR-C, E74A, and E93 regulate genes involved in signaling, cytoplasmic reorganization, proteolysis, autophagy (apg-related genes), and apoptosis
Apoptosis genes function in Drosophila autophagic cell death
Several lines of evidence indicate that the core apoptosis machinery functions in autophagic cell death during Drosophila development. Autophagic cell death of Drosophila salivary glands and midguts is foreshadowed by markers of apoptosis including DNA fragmentation.39,40,56 In addition, destruction of salivary glands and midguts is prevented by expression of the caspase inhibitor p35.39,40,56 Significantly, the proapoptotic genes rpr and hid (W) increase in transcription following the rise in ecdysone that triggers salivary autophagic cell death, and this change is concurrent with a decrease in the inhibitor of apoptosis diap2.39 At the same time, the fly Apaf-1 homologue ark (dark, dapaf1, hac1), the caspases dronc (Nc), drice (Ice), and dream (strica), the Bcl-2 family member buffy (dborg-2), the CD36 realtive crq, and the DNase rep4, all exhibit dramatic increases in transcription just before salivary gland autophagic cell death.36,59,60
Animals with mutations in the transcription regulators BR-C, E74A, and E93 prevent proper transcription of apoptosis genes (Figure 3). Both BR-C and E93 mutant salivary glands have dramatically reduced levels of rpr, hid, dronc, dream, and crq.36,59 While buffy and drice RNA levels are also decreased in E93 mutant salivary glands, drice is not altered and buffy is ectopically expressed in BR-C mutants.36 E74A mutants exhibit reduced levels of hid, crq, buffy, drice, and dream.36,59 Studies of apoptosis gene promoters indicate that rpr is a direct target of the ecdysone receptor54 and that dronc is a direct target of BR-C.61 These data suggest that the ecdysone-regulated early genes BR-C, E74A, and E93 have overlapping and distinct target apoptosis genes that they regulate. Significantly, it appears that proper transciption of apoptosis genes is essential for autophagic cell death, since animals with mutations in BR-C, E74A, and E93 prevent destruction of salivary glands. This is intriguing, as studies of the human mammary carcinoma cell line MCF-7 indicate that caspases are not required for autophagic death of these cells.62
Yeast autophagy genes appear to be conserved in higher animals, and are induced during autophagic programmed cell death
In all, 11 of the 19 genes that are required for yeast autophagy encode related genes in flies and humans (hereafter refered to as apg-related genes)16,63 (Table 1). Nine of these 11 apg-related genes are transcribed in dying Drosophila glands, and the seven genes that are most similar to apg2 (CG1241), apg3 (CG6877), apg4 (CG6194), apg5 (CG1643), apg7 (CG5489), apg9 (CG3615), and aut10/cvt18 (CG7986) exhibit increased transcription following the rise in steroid hormone that triggers autophagic programmed cell death of this tissue.36,64 It is interesting that flies possess genes that are related to the yeast genes involved in both of the autophagy ubiquitin conjugation systems, and that they are also the genes that increase in transcription following the rise in steroid that triggers autophagic cell death. CG1241 encodes a novel protein that is similar to APG2 that functions in the formation and completion of autophagic vacuoles.65 APG2 interacts with the integral membrane protein APG9/AUT9/CVT7,66 which is similar to the predicted protein encoded by CG3615. CG7986 encodes a predicted protein that is most similar to AUT10/CVT18 which has WD40 domains, functions in the formation of autophagic vacuoles, and is required for proper localization of APG2.67 CG6194 encodes a predicted protein that is similar to the AUT2/APG4 cysteine protease, which interacts with and processes AUT7/APG8/CVT5, and is involved in the formation and completion of autophagic vacuoles.68 AUT7/APG8/CVT5 is conjugated with the ubiquitin-E1-like enzyme APG7/CVT2,31 which is similar to the protein encoded by CG5489, indicating that one of the ubiquitin-like conjugation systems may function in salivary gland autophagy. AUT7/APG8/CVT5 is also conjugated with the ubiquitin-E2-like enzyme AUT1/APG3.31 AUT1/APG3 is similar to the predicted protein encoded by CG6877, suggesting the possibility that the second ubiquitin-like conjugation system functions in salivary gland autophagy. CG1643 encodes a predicted protein that is similar to the novel protein APG5,69 which also functions in the formation of autophagic vacuoles. APG5 is conjugated to APG12, and this complex is also activated by the ubiquitin-E1-like enzyme APG7/CVT2,32 further indicating that both ubiquitin-like conjugation systems that are used during yeast autophagy may also function in salivary gland cell death.
Mutations in the ecdysone-regulated transcription factor genes that prevent proper salivary gland cell death also inhibit proper transcription of the apg-related genes.36 Animals with mutations in BR-C exhibit ectopic transcription of CG6194 (apg4-like) and CG1643 (apg5-like), and decreased transcription of CG5489 (apg7-like) in salivary glands at the stage that they would normally die. While animals with mutations in E74A only exhibit decreased transcription of CG6194, salivary glands dissected from E93 mutants have decreased levels of CG1241 (apg2-like), CG6194, CG1643, and CG5489. Yeasts with mutations in apg4, apg5, and apg7 are defective in autophagic vacuole formation.27 Thus, it is striking that E93 mutants exhibit decreased CG6194, CG1643, and CG5489 RNA levels, since E93 mutant salivary gland and midgut cells are defective in the formation of autophagic vacuoles.40,56
Additional factors that are induced during Drosophila autophagic cell death
Genomewide studies of dying cells have identified numerous interesting genes that are induced just prior to autophagic programmed cell death (Figure 3). For example, several signaling molecules and transcription regulators increase in transcription in dying salivary glands. The Drosophila gene CG8304 encodes a serine/threonine kinase that is most similar to the death-associated protein kinase (DAPk) in humans, and transcription of this gene increases 36-fold in dying salivary glands.36 DAPk mediates membrane blebbing and formation of vesicles in dying human cell lines,70 and suggests that CG8304 may have a related function in Drosophila. Steroids usually function by activating transcription and, therefore, it is not surprising that transcription regulators are induced in dying salivary glands including the corepressors (Smarter and CG4756), the NFkB regulator cactus, the NFkB family member dif, several other genes encoding DNA-binding proteins (bun, sox14, CG8319), and components of transcription complexes (trap95 and hsf).36,64
Salivary gland cells exhibit dynamic changes in cell shape and vacuole localization just before autophagic cell destruction, suggesting that cytoplasmic reorganization and proteolysis are important for their death. Consistent with this notion, genes encoding motor proteins including ctp, ck, and dlc90F, and members of the Rho, Rac, and Rab families of small guanosine triphosphates (GTPases) exhibited increase in transcription just before salivary gland autophagic cell death.36,64 While cell remodeling is known to play an important role in phagocyte engulfment of apoptotic cells,71 the role of these factors in autophagic cell death is less clear. In addition to caspases, several cysteine, serine, and metalloproteases increase in transcription in dying salivary glands, and these changes in RNA level are complemented by decreased transcription of cysteine, serine, and metalloprotease inhibitors.36,64 Furthermore, upregulation of the matrix metalloprotease mmp1 is abolished in mutants that prevent salivary gland cell death.36 These proteases likely complement caspase function during autophagic cell death, as mutations in mmp2 prevent proper destruction of Drosophila midgets,72 this tissue dies by autophagic cell death,56 and inhibition of caspases does not completely prevent autophagic changes in dying cells.40
The most important advances in science often come from the exploration of the unknown. Therefore, it is interesting that many genes encoding novel factors are induced in dying cells. We have identified genes that increase following both radiation-induced apoptosis and steroid-triggered autophagy. Very few common genes were identified using this approach, but the genes CG10965, CG17323, CG7144, EG25E8.4, and CG5254 increased in RNA levels during both ecdysone-and radiation-triggered cell death.36 Transcription of these genes was also altered in salivary glands of BR-C, E74A, and E93 mutants, suggesting a possible functional role for these genes in autophagic programmed cell death.
Concluding Remarks and Future Directions
Studies of developing animals indicate that at least two forms of programmed cell death have been conserved during evolution.2 Studies of apoptosis in cell lines indicate that the biochemical mechanisms underlying programmed cell death are similar in different cell types. Dying cell morphology can be very different,73 however, and identifying the mechanisms that regulate different types of physiological cell death will lead to a better understanding of how defects in this fundamental cellular process lead to aberrations in animal growth and development.
The steroid ecdysone triggers autophagic cell death during Drosophila development, but other factors must provide competence signals for cells to be capable of dying. These ‘competence factors’ may include cell age, status in the cell cycle, the presence of a transcriptional activator, absence of a transcriptional repressor, or the state of chromatin to name a few possibilities. Ultimately, this steroid signal activates a group of transcription regulators that include BR-C, E74A, and E93. These early genes appear to regulate a complex series of downstream cellular events that include cytoplasmic reorganization including cell shape and vacuole transport, proteolysis by caspase and apparently noncaspase proteases, genes related to yeast genes that function in autophagy, and many unknowns.
Many gaps exist in our knowledge of the mechanism underlying autophagic cell death, and this is an exciting time to ask critical questions about this understudied form of cell destruction. How many of the genes that are expressed in dying cells are required, and is it critical for these factors to integrate in a specific temporal manner? Do the apg-related genes serve the same purpose in yeast and higher animals? Are the noncaspase genes that are used during autophagy used during apoptosis? What is the clinical relevance of autophagic programmed cell death? Given our limited knowledge of the mechanisms underlying autophagic programmed cell death, it is clear that this fundamental cellular process will be a fruitful area of investigation for years to come.
Abbreviations
- dark:
-
fly Apaf-1 homolog ark
- Nc:
-
the caspase dronc
References
Schweichel J-U and Merker H-J (1973) The morphology of various types of cell death in prenatal tissues. Teratology 7: 253–266
Clarke PGH (1990) Developmental cell death: morphological diversity and multiple mechanisms. Anat. Embryol. 181: 195–213
Kerr JF, Wyllie AH and Currie AR (1972) Apoptosis: a basic biological phenomenon with wide-ranging implications in tissue kinetics. Br. J. Cancer 26: 239–257
Johnstone RW, Ruefli AA and Lowe SW (2002) Apoptosis: a link between cancer genetics and chemotherapy. Cell 108: 153–164
Hengartner MO (2000) The biochemistry of apoptosis. Nature 407: 770–776
Shi Y (2002) Mechanisms of caspase activation and inhibition during apoptosis. Mol. Cell 9: 459–470
Weissman AM (2001) Themes and variations on ubiquitylation. Nat. Rev. Mol. Cell Biol. 2: 169–178
Ohsumi Y (2001) Molecular dissection of autophagy: two ubiquitin-like systems. Nat. Rev. Mol. Cell Biol. 2: 211–216
Ashford TP and Porter KR (1962) Cytoplasmic components in hepatic cell lysosomes. J. Cell. Biol. 12: 198–202
Essner E and Novikoff AB (1962) Cytological studies on two functional hepatomas, interrelations of endoplasmic reticulum, golgi apparatus, and lysosomes. J. Cell. Biol. 15: 289–312
Klionsky DJ and Emr SD (2000) Autophagy as a regulated pathway of cellular degradation. Science 290: 1717–1721
Dunn WAJ (1990) Studies on the mechanisms of autophagy: formation of the autophagic vacuole. J. Cell. Biol. 110: 1923–1933
Scharrer B (1966) Ultrastructural study of the regressing prothoracic glands of blattarian insects. Z. Zellerforsch. 69: 1–21
Schin KS and Clever U (1965) Lysosomal and free acid phosphatase in salivary glands of Chironomus tentans. Science 150: 1053–1055
Scheib D (1965) Sur la regression du canal de de Müller male de l'embryon de poulet: localisation de la phosphatase acide au microscope electronique. C. R. Acad. Sci. Hebd. Seances. Acad. Sci. D 261: 5219–5221
Baehrecke EH (2002) How death shapes life during development. Nat. Rev. Mol. Cell. Biol. 3: 779–787
Tanida I, Tanida-Miyake E, Komatsu M, Ueno T and Kominami E (2002) Human Apg3p/Aut1p homologue is an authentic E2 enzyme for multiple substrates, GATE-16, GABARAP, and MAP-LC3, and facilitates the conjugation of hApg12p to hApg5p. J. Biol. Chem. 277: 13739–13744
Tanida I, Tanida-Miyake E, Ueno T and Kominami E (2001) The human homolog of Saccharomyces cerevisiae Apg7p is a protein-activating enzyme for multiple substrates including human Apg12p, GATE-16, GABARAP, and MAP-LC3. J. Biol. Chem. 276: 1701–1706
Schulte-Hermann R, Bursch W, Grasl-Kraupp B, Marian B, Torok L, Kahl-Rainer P and Ellinger A (1997) Concepts of cell death and application to carcinogenesis. Toxicol. Pathol. 25: 89–93
Bursch W, Ellinger A, Kienzl H, Torok L, Pandey S, Sikorska M, Walker R and Hermann RS (1996) Active cell death induced by the anti-estrogens tamoxifen and ICI164384 in human mammary carcinoma cells (MCF-7) in culture: the role of autophagy. Carcinogenesis 17: 1595–1607
Liang XH, Jackson S, Seaman M, Brown K, Kempkes B, Hibshoosh H and Levine B (1999) Induction of autophagy and inhibition of tumorigenesis by beclin 1. Nature 402: 672–676
Jia L, Dourmashkin RR, Allen PD, Gray AB, Newland AC and Kelsey SM (1997) Inhibition of autophagy abrogates tumour necrosis factor alpha induced apoptosis in human T-lymphoblastic leukaemic cells. Br. J. Haematol. 98: 673–685
Kegel KB, Kim M, Sapp E, McIntyre C, Castano JG, Aronin N and DiFiglia M (2000) Huntingtin expression stimulates endosomal–lysosomal activity, endosome tubulation, and autophagy. J. Neurosci. 20: 7268–7278
Yue Z, Horton A, Bravin M, DeJager PL and Selimi FNH (2002) A novel protein complex linking the delta 2 glutamate receptor and autophagy: implications for neurodegeneration in lurcher mice. Neuron 35: 921–933
Stefanis L, Larsen KE, Rideout HJ, Sulzer D and Greene LA (2001) Expression of A53 T mutant but not wild-type alpha-synuclein in PC12 cells induces alterations of the ubiquitin-dependent degradation system, loss of dopamine release, and autophagic cell death. J. Neurosci. 21: 9549–9560
Mastroberardino PG, Iannicola C, Nardacci R, Bernassola F, De Laurenzi V, Melino G, Moreno S, Pavone F, Oliverio S, Fesus L and Piacentini M (2002) ‘Tissue’ transglutaminase ablation reduces neuronal death and prolongs survival in a mouse model of Huntington's disease. Cell Death Differ. 9: 873–880
Tsukada M and Ohsumi Y (1993) Isolation and characterization of autophagy-defective mutants of Saccharomyces cerevisiae. FEBS Lett. 1–2: 169–174
Thumm M, Egner R, Koch B, Schlumpberger M, Straub M, Veenhuis M and Wolf DH (1994) Isolation of autophagocytosis mutants of Saccharomyces cerevisiae. FEBS Lett. 349: 275–280
Harding TM, Morano KA, Scott SV and Klionsky DJ (1995) Isolation and characterization of yeast mutants in the cytoplasm to vacuole protein targeting pathway. J. Cell. Biol. 131: 591–602
Harding TM, Hefner-Gravink A, Thumm M and Klionsky DJ (1996) Genetic and phenotypic overlap between autophagy and the cytoplasm to vacuole protein. J. Biol. Chem. 271: 17621–17624
Ichimura Y, Kirisako T, Takao T, Satomi Y, Shimonishi Y, Ishihara N, Mizushima N, Tanida I, Kominami E, Ohsumi M, Noda T and Ohsumi Y (2000) A ubiquitin-like system mediates protein lipidation. Nature 408: 488–492
Mizushima N, Noda T, Yoshimori T, Tanaka Y, Ishii T, George MD, Klionsky DJ, Ohsumi M and Ohsumi Y (1998) A protein conjugation system essential for autophagy. Nature 395: 395–398
Tanida I, Mizushima N, Kiyooka M, Ohsumi M, Ueno T, Ohsumi Y and Kominami E (1999) Apg7p/Cvt2p: a novel protein-activating enzyme essential for autophagy. Mol. Biol. Cell 10: 1367–1379
Kim J, Dalton VM, Eggerton KP, Scott SV and Klionsky DJ (1999) Apg7p/Cvt2p is required for the cytoplasm-to-vacuole targeting, macroautophagy, and peroxisome degradation pathways. Mol. Biol. Cell 10: 1337–1351
Shintani T, Mizushima N, Ogawa Y, Matsuura A, Noda T and Ohsumi Y (1999) Apg10p, a novel protein-conjugating enzyme essential for autophagy in yeast. EMBO J. 18: 5234–5241
Lee C-Y, Clough EA, Yellon P, Teslovich TM, Stephan DA and Baehrecke EH (2003) Genome-wide analyses of steroid- and radiation-triggered programmed cell death in Drosophila. Curr. Biol. 13: 350–357
Bodenstein D (1965) The postembryonic development of Drosophila. In Biology of Drosophila Demerec M (ed) New York: Hafner Publishing Co., pp. 275–367
Robertson CW (1936) The metamorphosis of Drosophila melanogaster, including an accurately timed account of the principal morphological changes. J. Morphol. 59: 351–399
Jiang C, Baehrecke EH and Thummel CS (1997) Steroid regulated programmed cell death during Drosophila metamorphosis. Development 124: 4673–4683
Lee CY and Baehrecke EH (2001) Steroid regulation of autophagic programmed cell death during development. Development 128: 1443–1455
Jochova J, Zakeri Z and Lockshin RA (1997) Rearrangement of the tubulin and actin cytoskeleton during programmed cell deathin Drosophila salivary glands. Cell Death Differ. 4: 140–149
Becker HJ (1959) Die puffs der speicheldrüsenchromosomen von Drosophila melanogaster I. Beobachtungen zur verhalten des puffmusters in normalstamm und bei zwei mutanten, giant und lethal-giant-larvae. Chromosoma 10: 654–678
Ashburner M (1967) Patterns of puffing activity in the salivary gland chromosomes of Drosophila. I. Autosomal puffing patterns in a laboratory stock of Drosophila melanogaster. Chromosoma 21: 398–428
Andres AJ and Thummel CS (1992) Hormones, puffs, and flies: the molecular control of metamorphosis by ecdysone. Trends Genet. 8: 132–138
Koelle MR, Talbot WS, Segraves WA, Bender MT, Cherbas P and Hogness DS (1991) The Drosophila EcR gene encodes an ecdysone receptor, a new member of the steroid receptor superfamily. Cell 67: 59–77
Shea MJ, King DL, Conboy MJ, Mariani BD and Kafatos FC (1990) Proteins that bind to Drosophila chorion cis-regulatory elements: a new C2H2 zinc finger protein and a C2C2 steroid receptor-like component. Genes Dev. 4: 1128–1140
Henrich VC, Sliter TJ, Lubahn DB, MacIntyre A and Gilbert LI (1990) A steroid/thyroid hormone receptor superfamily member in Drosophila melanogaster that shares extensive sequence similarity with a mammalian homologue. Nucleic Acids Res. 18: 4143–4148
Oro AE, McKeown M and Evans RM (1990) Relationship between the product of the Drosophila ultraspiracle locus and vertebrate retinoid X receptor. Nature 347: 298–301
Thomas HE, Stunnenberg HG and Stewart AF (1993) Heterodimerization of the Drosophila ecdysone receptor with retinoid X receptor and ultraspiracle. Nature 362: 471–475
Yao T-P, Segraves WA, Oro AE, McKeown M and Evans RM (1992) Drosophila ultraspiracle modulates ecdysone receptor function via heterodimer formation. Cell 71: 63–72
Woodard CT, Baehrecke EH and Thummel CS (1994) A molecular mechanism for the stage-specificity of the Drosophila prepupal genetic response to ecdysone. Cell 79: 607–615
Broadus J, McCabe JR, Endrizzi B, Thummel CS and Woodard CT (1999) The Drosophila βFTZ-F1 orphan nuclear receptor provides competence for stage-specific responses to the steroid hormone ecdysone. Mol. Cell 3: 143–149
Restifo LL and White K (1992) Mutations in a steroid hormone-regulated gene disrupt the metamorphosis of internal tissues in Drosophila: salivary glands, muscle, and gut. Wilhelm Roux's Arch. Dev. Biol. 201: 221–234
Jiang C, Lamblin A-FJ, Steller H and Thummel CS (2000) A steroid-triggered transcriptional hierarchy controls salivary gland cell death during Drosophila metamorphosis. Mol. Cell 5: 445–455
Lee C-Y, Wendel DP, Reid P, Lam G, Thummel CS and Baehrecke EH (2000) E93 directs steroid-triggered programmed cell death in Drosophila. Mol. Cell 6: 433–443
Lee C-Y, Cooksey BAK and Baehrecke EH (2002) Steroid regulation of midgut cell death during Drosophila development. Dev. Biol. 250: 101–111
Burtis KC, Thummel CS, Jones CW, Karim FD and Hogness DS (1990) The Drosophila 74EF early puff contains E74, a complex ecdysone-inducible gene that encodes two ets-related proteins. Cell 61: 85–99
DiBello PR, Withers DA, Bayer CA, Fristrom JW and Guild GM (1991) The Drosophila Broad-Complex encodes a family of related proteins containing zinc fingers. Genetics 129: 385–397
Lee C-Y, Simon CR, Woodard CT and Baehrecke EH (2002) Genetic mechanism for the stage-and tissue-specific regulation of steroid-triggered programmed cell death in Drosophila. Dev. Biol. 252: 138–148
Dorstyn L, Colussi PA, Quinn LM, Richardson H and Kumar S (1999) DRONC, an ecdysone-inducible Drosophila caspase. Proc. Natl. Acad. Sci. USA 96: 4307–4312
Cakouros D, Daish T, Martin D, Baehrecke EH and Kumar S (2002) Ecdysone-induced expression of the caspase DRONC during hormone-dependent programmed cell death in Drosophila is regulated by Broad-Complex. J. Cell Biol. 157: 985–995
Paglin S, Hollister T, Delohery T, Hackett N, McMahill M, Sphicas E, Domingo D and Yahalom J (2001) A novel response of cancer cells to radiation involves autophagy and formation of acidic vesicles. Cancer Res. 61: 439–444
Reggiori F and Klionsky DJ (2002) Autophagy in the eukaryotic cell. Eukaryot. Cell 1: 11–21
Gorski SM, Chittaranjan S, Pleasance ED, Freeman JD, Anderson CL, Varhol RJ, Coughlin SM, Zuyderduyn SD, Jones SJ and Marra MA (2003) A SAGE approach to discovery of genes involved in autophagic cell death. Curr. Biol. 13: 358–363
Wang CW, Kim J, Huang WP, Abeliovich H, Stromhaug PE, Dunn WAJ and Klionsky DJ (2001) Apg2 is a novel protein required for the cytoplasm to vacuole targeting, autophagy, and pexophagy pathways. J. Biol. Chem. 276: 30442–30451
Noda T, Kim J, Huang WP, Baba M, Tokunaga C, Ohsumi Y and Klionsky DJ (2000) Apg9p/Cvt7p is an integral membrane protein required for transport vesicle formation in the Cvt and autophagy pathways. J. Cell Biol. 148: 465–480
Guan J, Stromhaug PE, George MD, Habibzadegah-Tari P, Bevan A, Dunn WAJ and Klionsky DJ (2001) Cvt18/Gsa12 is required for cytoplasm-to-vacuole transport, pexophagy, and autophagy in Saccharomyces cerevisiae and Pichia pastoris. Mol. Biol. Cell 12: 3821–3838
Kirisako T, Ichimura Y, Okada H, Kabeya Y, Mizushima N, Yoshimori T, Ohsumi M, Takao T, Noda T and Ohsumi Y (2000) The reversible modification regulates the membrane-binding state of Apg8/Aut7 essential for autophagy and the cytoplasm to vacuole targeting pathway. J. Cell. Biol. 151: 263–276
Kametaka S, Matsuura A, Wada Y and Ohsumi Y (1996) Structural and functional analyses of APG5, a gene involved in autophagy in yeast. Gene 178: 139–143
Inbal B, Bialik S, Sabanay I, Shani G and Kimchi A (2002) DAP kinase and DRP-1 mediate membrane blebbing and the formation of autophagic vesicles during programmed cell death. J. Cell Biol. 157: 455–468
Hengartner MO (2001) Apoptosis: coralling the corpses. Cell 104: 325–328
Page-McCaw A, Serano J, Sante JM and Rubin GM (2003) Drosophila matrix metalloproteinases are required for tissue remodeling, but not embryonic development. Dev. Cell 4: 95–106
Leist M and Jäätelä M (2001) Four deaths and a funeral: from caspases to alternative mechanisms. Nat. Rev. Mol. Cell Biol. 2: 1–10
Acknowledgements
I thank many colleagues and the members of the Baehrecke laboratory for helpful discussions, and Deborah Berry for comments on the manuscript. Work on this subject has been supported by NIH grant GM59136.
Author information
Authors and Affiliations
Corresponding author
Additional information
Edited by M Piacentini
Rights and permissions
About this article
Cite this article
Baehrecke, E. Autophagic programmed cell death in Drosophila. Cell Death Differ 10, 940–945 (2003). https://doi.org/10.1038/sj.cdd.4401280
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.cdd.4401280
Keywords
This article is cited by
-
Autophagy in animal development
Cell Death & Differentiation (2020)
-
Thin is required for cell death in the Drosophila abdominal muscles by targeting DIAP1
Cell Death & Disease (2018)
-
Drosophila MOF regulates DIAP1 and induces apoptosis in a JNK dependent pathway
Apoptosis (2016)
-
Autophagy in the fat body cells of the cave cricket Troglophilus neglectus Krauss, 1878 (Rhaphidophoridae, Saltatoria) during overwintering
Protoplasma (2016)