Wild mice around Chicago may have switched genotype to keep pace with modern living.
Abstract
We have compared the sequences of mitochondrial DNA extracted from museum skins of white-footed mice caught in the Chicago area since 1855 and from modern mice trapped alive in the same locations. We found a consistently similar directional change of mouse genotype over this period at each of five collection sites that were separated by 10–70 km. The genotype most common 100 years ago is now extremely rare, indicating that the mammalian mitochondrial genome can undergo rapid evolution.
Similar content being viewed by others
Main
Museums from Alaska to Switzerland were searched for specimens of the white-footed mouse, Peromyscus leucopus, originally collected from two counties in northeastern Illinois, USA. The white-footed mouse is the most common rodent in the deciduous forests of North America. We borrowed 61 museum-specimen skins of mice collected from five locations in the Chicago area, and in 1999 we trapped 52 white-footed mice in the same locations and in three others.
Small (12 mm × 1.5 mm) strips were cut from the ventral suture of the museum-specimen mouse skins. We found that shaving the hair from these strips before extraction, combined with a freeze–thaw step, reduced the dark colour of the extract and prevented inhibition of the polymerase chain reaction (PCR)1. The Chelex-100 DNA-extraction protocol2 was modified by the addition of proteinase K (ref. 3).
We used a 340-base-pair polymorphic region within the coding portion of the cytochrome oxidase II gene to genotype the museum mouse skins. This region was amplified for sequencing of DNA from 56 of 61 skins by using KlentaqLA DNA polymerase and PCR buffer (DNA Polymerase Technology)4,5,6,7, extremely long (20-min) extension cycles5 and betaine8, in conjunction with three nested PCR primer sets spanning 559, 496 and 400 base pairs in turn.
Three haplotypes (named A, M and Mw) were identified: Mw was found in only three mice. M and Mw differ at just a single position, but they both differ from A at four positions in this 340-base-pair region (see supplementary information); none of the base changes alters the amino-acid sequence of the gene.
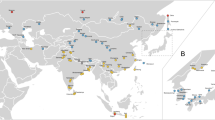
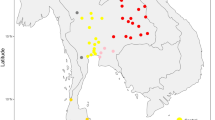
At four of the five collection sites, the oldest white-footed mice were predominantly of genotype A. At each geographical location there was a monotonic decrease in the proportion of A (Fig. 1). Pooling sequencing data for all five locations, the proportion of A was 5/5 in 1850–99, 18/30 in 1900–49 and 4/73 in 1950–99. The observed frequency change requires less than a 1% per generation advantage of M over A. None of the live mice trapped at these five locations had the A type, and only one A mouse was trapped in 1999–2000.
The geographic relation is shown between the five locations (arrows) from which pre-1950 museum specimens of Peromyscus leucopus were obtained. Major highways are shown in black; natural areas are shown in green. The frequencies (vertical scale) of three different mitochondrial haplotypes (A, green; M, red; Mw, yellow) are shown for each location for the periods 1850–99, 1900–49 and 1950–2000.
As the change in the mitochondrial genotype of the wild mouse coincides with a marked increase in human activity in the region, we presume that this caused the replacement of the A haplotype by the M. The M genotype might have become advantageous in the altered habitat, or it could be unconditionally advantageous and have been introduced by people from elsewhere.
In the Chicago region, the white-footed mouse (Fig. 2) has displaced the prairie deer mouse from the few remaining prairies9,10. We suggest that the M haplotype has not only spread through the white-footed mouse population but might also have contributed to the displacement of the prairie deer mouse by the white- footed mouse.
References
Goodyear, P. D., MacLaughlin-Black, S. & Mason, I. J. Biotechniques 16, 232–233 (1994).
Walsh, P. S., Metzger, D. A. & Higuchi, R. Biotechniques 10, 506–513 (1991).
Steinberg, E. K. Mol. Ecol. 8, 1075–1092 (1999).
Barnes, W. M. Gene 112, 29–35 (1992).
Barnes, W. M. Trends Biochem. Sci. 19, 342 (1994).
Barnes, W. M. Proc. Natl Acad. Sci. USA 91, 2216–2220 (1994).
Barnes, W. M. US Patent No. 5,436,149 (1995).
Baskaran, N. et al. Genome Res. 6, 633–638 (1996).
Pergams, O. R. W. & Nyberg, D. J. Mammalogy 82, 984–992 (2001).
Pergams, O. R. W. & Nyberg, D. Proc. R. Soc. Lond. B (submitted).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
Competing financial interests: W.M.B. is an inventor and provider or Klentaq1 DNA polymerase (corrected on 27 May 2003 from "declared none", as was previously stated).
Supplementary information
Rights and permissions
About this article
Cite this article
Pergams, O., Barnes, W. & Nyberg, D. Rapid change in mouse mitochondrial DNA. Nature 423, 397 (2003). https://doi.org/10.1038/423397a
Issue Date:
DOI: https://doi.org/10.1038/423397a
This article is cited by
-
Mitochondrial lineage sorting in action – historical biogeography of the Hyles euphorbiae complex (Sphingidae, Lepidoptera) in Italy
BMC Evolutionary Biology (2013)
-
Ancient DNA studies: new perspectives on old samples
Genetics Selection Evolution (2012)
-
Re-examination of the historical range of the greater prairie chicken using provenance data and DNA analysis of museum collections
Conservation Genetics (2006)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.