Abstract
Congenital mesoblastic nephroma (CMN) and infantile fibrosarcoma (IFS) are two pediatric tumors arising in the kidneys and soft tissues of infants, respectively. Recently, a t(12;15)(p13;q25) resulting in ETV6-NTRK3 gene fusion was detected in patients with IFS and in patients with the cellular type of CMN, suggesting a common pathogenetic pathway. We investigated the presence or absence of ETV6 rearrangements and numerical abnormalities of chromosome 11 by using fluorescence in situ hybridization on paraffin-embedded material from five cases of IFS, two of CMN, and one of mixed type (CMN and IFS) found in our files. In three cases of IFS, we found ETV6 gene rearrangement but a normal copy number of chromosome 11. One case each of IFS, the cellular type of CMN, and the mixed type (CMN and IFS) had both abnormalities. In a case of classic CMN, neither trisomy 11 nor gene rearrangement was found. It is possible that trisomy 11 is a later, nonessential event in the pathogenetic process or that this secondary aberration is associated with still-unrecognized clinical or biological characteristics. We confirmed that IFS and the cellular type of CMN are cytogenetically related and can occur synchronously in the same organ.
Similar content being viewed by others
INTRODUCTION
Congenital mesoblastic nephroma (CMN) was described in 1967 by Bolande et al. (1) based on eight cases of the classic type. The cellular type of CMN, with lesions displaying necrosis, high cellularity, and atypia or mitosis, was reported later (2, 3). Trisomy 11 is the most consistent nonrandom karyotypic finding in the CMNs cytogenetically investigated so far (4), occurring almost exclusively in the cellular type of CMN and the mixed type of CMN and infantile fibrosarcoma (IFS; 5, 6). IFS was characterized by Chung and Enzinger (7) in their series of 53 cases, which showed patients with IFS to have a much better prognosis than adults with fibrosarcomas. A combination of numerical changes (chromosomal gains), including trisomy 11, initially were said to be characteristic cytogenetic findings in IFS (8, 9).
However, Knezevich et al. (10) reported a recurrent cryptic t(12, 15)(p13;q15) resulting in a fusion between the ETV6 gene in the band 12p13 and the NTRK3 gene in the band 15p15. Such cytogenetic features were not seen in any of the 18 control cases, which included cases of adult-type fibrosarcoma and infantile fibromatosis (10). It was subsequently established that both IFS and the cellular type of CMN can carry the same 12;15 translocation, suggesting a possible link between these two pediatric tumors (11, 12).
We retrospectively reviewed tumors coded as CMN or IFS in the Mayo Clinic surgical pathology files and in the consultation files of one of the authors (AGN). We also included a case in which morphologic features of both CMN and IFS were seen in transition-like areas. Molecular cytogenetic technique or fluorescence in situ hybridization (FISH) analysis was used to evaluate the presence or absence of ETV6 gene rearrangements and the copy number of chromosome 11 in these cases without prior knowledge of the histopathologic diagnosis.
MATERIALS AND METHODS
Selection of Cases
Eight cases were selected for study. After review of hematoxylin and eosin–stained slides, cases were classified as IFS (five cases), CMN classic type (one case), CMN cellular type (one case), or mixed type (IFS and CMN, one case). Clinical data are summarized in Table 1. One representative block from each case was sent for molecular cytogenetic analysis.
Preparation of Nuclei From Paraffin Sections
Nuclei were extracted from paraffin-embedded 50-μm-thick tissue sections as described by Kuchinka et al. (13). In brief, the sections were deparaffinized in 2 × 5-minute baths in xylene and 2 × 5-minute baths in ethanol. Digestion was performed by collagenase XI (Sigma, St. Louis, MO) for 2 hours, followed by 0.05% trypsin and EDTA (Life Technologies, Rockville, MD) for 30 minutes at 37° C. The cells were spread on positively charged slides, baked for 2 hours at 50° C, and treated with 30% sodium bisulfite (Pretreatment solution, Oncor, Gaithersburg, MD) for 10 minutes, followed by digestion in 250 μg/mL proteinase K (Oncor) for 10 to 15 minutes at 45° C.
Fluorescence In Situ Hybridization Analysis
FISH analysis was performed to assess the status of the ETV6 gene using the yeast artificial chromosome clone 964c10, spanning the ETV6 gene region in 12p13 (14). To increase the specificity of the analysis, 964c10 was hybridized to interphase nuclei together with another yeast artificial chromosome clone probe, 753f12, localized to the neighboring KRAS2 region in 12p12 (Fig. 1A). The clones 964c10 and 753f12 were obtained from Fondation Jean Dausset (CEPH, Paris, France). Labeling was performed by random-priming octanucleotides (Life Technologies).
A, ideogram of a G-banded chromosome 12 with positions of the ETV6 and KRAS2 probes. B, schematic description of hybridization pattern resulting from a 12;15 translocation involving the ETV6 gene region: the ETV6 signal (green) is split between the derivative (der) chromosomes 12 and 15, whereas the KRAS2 signal remains on the der(12). C, schematic interphase pattern in nuclei without and with rearrangement of the ETV6 region. D, interphase nucleus with a normal hybridization pattern; in other words, two pairs of colocalized ETV6 (green) and KRAS2 (red) signals; blending of red and green results in yellow fluorescence. E, nucleus from Case 3, showing two copies of the KRAS2 region (red) and three signals for the ETV6 region; the smaller green signal is located some distance from the red-green-yellow signals, indicating that it is translocated to another chromosome. F, nucleus from Case 7, showing one intact and one split signal for ETV6 (red) and three signals for the centromere of chromosome 11 (green).
With use of these probes, normal interphase nuclei show four signals arranged as two pairs (Fig. 1B–C). Nuclei harboring a translocation involving ETV6 show two signals for KRAS2, two signals for ETV6 adjacent to the KRAS2 signals, and an additional ETV6 signal. The centromeric alpha-satellite probes for chromosomes 11 (D11Z1) and 12 (D12Z1) were detected by biotinylated and digoxigenin-labeled alpha-satellite probes (Oncor).
Probes and extracted nuclei were denatured simultaneously on a HYBrite slide warmer (Vysis, Downers Grove, IL). This was followed by hybridization at 37° C overnight and stringency washing in 50% formamide/2 × SSC for 15 minutes and 2 × SSC for 8 minutes at 42° C. Biotinylated probes were detected by fluorescein isothiocyanate-avidin or Alexa594-streptavidin, and digoxigenin-labeled probes were detected by rhodamine or fluorescein isothiocyanate-conjugated anti-digoxigenin antibodies (Oncor). At least 50 cells with signals were analyzed for each hybridization. Nuclei from the karyotypically normal human fibroblast line GM498B (NIGMS Human Cell Repository, Camden, NJ) were used to determine normal hybridization patterns. Of 500 interphase cells hybridized with centromeric and single-copy probes, more than 90% showed only two signals per cell, and fewer than 5% showed three or more signals (Fig. 1D).
RESULTS
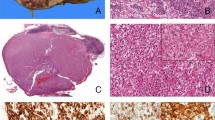
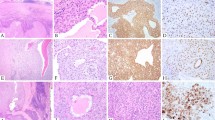
The conditions studied occurred in four boys and three girls (records did not indicate the sex of the patient in Case 6). Mean age was 10 months (range, 4 months to 2 years; records did not indicate patient age in three cases). The five patients with IFS were affected in the extremities (three cases) and the head (two cases). The two patients with CMN were affected in the kidneys, as was the one patient with the mixed type, IFS/CMN (Table 1). The tumors in Cases 1, 2, 3, and 8 were composed of a hypercellular malignant spindle cell population arranged in a herringbone pattern, with numerous mitotic figures, consistent with the diagnosis of IFS. The tumor in Case 4 was composed of a multinodular hypocellular spindle cell population in a myxoid background, with moderately cellular areas and rare mitotic figures (Fig. 2). The neoplasm in Case 7 arose from the left kidney. On low-power view, two distinct areas were seen. One area was composed of hyperchromatic, malignant-looking spindle cells arranged haphazardly in a fascicular pattern with a high mitotic index. The other area was composed of a bland-looking spindle cell population with an absence of mitotic figures, consistent with CMN (Figs. 3 and 4). The tumors in Cases 5 and 6 had morphologic features consistent with the cellular type and the classic type of CMN, respectively.
(Case 4). Photomicrograph of a bona fide IFS with mild to low cellular areas. Left, low-power view showing areas with moderate and high cellularity consistent with the diagnosis of IFS. (Hematoxylin and eosin stain; original magnification 4×.) Right, higher power view of cellular areas with fascicles of spindle cells. (Hematoxylin and eosin stain; original magnification 20×.)
Higher-power view of portion of field shown in Figure 3. Compare the cell population of the congenital mesoblastic nephroma (left side) with the more atypical cell population and higher mitotic index of the infantile fibrosarcoma (right side). (Hematoxylin and eosin stain; original magnification 20×.)
FISH analysis was performed on interphase nuclei from all eight cases (Table 1). An increased proportion of cells exhibited three signals positive for ETV6 probe (964c10) per nucleus in Cases 1, 2, 3, 5, 7, and 8 (44%, 47%, 36%, 27%, 13%, and 26% of nuclei, respectively). In these cells, dual-color FISH with the ETV6 and KRAS2 probes showed two colocalized ETV6-KRAS2 signals and an additional ETV6 signal (Fig. 1E). One of the three ETV6 signals usually exhibited weaker fluorescence than the other 2, indicating an asymmetric split. Three signals for chromosome 11 were observed in Cases 2, 5, and 7 (63%, 32%, and 17% of cells, respectively; Fig. 1F). In Case 4, both the chromosome 11 centromeric probe and the ETV6 probe showed increased frequencies of cells with three (22% and 26%) and four (15% and 13%) signals. An additional hybridization with a centromeric probe for chromosome 12 also showed three or four signals in a substantial percentage (20% and 36%) of cells, indicating Polyploidy or Polysomies 11 and 12 in this case. Case 6 had a normal hybridization pattern for ETV6 and centromere 11.
DISCUSSION
Previous authors have suggested that IFS and the cellular and mixed type of CMN are histogenetically linked (11, 12), and a t(12, 15)(p13;q25) resulting in ETV6-NTRK3 gene fusion has been observed in both entities.
In this study, FISH analysis demonstrated that rearrangements of the ETV6 region occurred in four of the five cases of IFS and in two of the three cases of CMN. A combination of ETV6 rearrangement and trisomy 11 occurred in one case each of IFS (Case 2), the cellular type of CMN (Case 5), and the mixed type, IFS and CMN (Case 7), but not in the classic type of CMN (Case 6). These findings are consistent with previously reported findings in cases of CMN and IFS (4, 5, 6, 8, 9, 10, 11, 12, 15, 16, 17).
In Case 4, two abnormal populations of cells carrying trisomy and tetrasomy for both chromosomes 11 and 12 were found in the absence of ETV6 rearrangement. This may reflect the presence of cells with additional copies of these two chromosomes or with near-triploid and near-tetraploid modal numbers. It is interesting to note that this tumor had a lower-grade morphology than the tumors in the other cases of IFS. Previous studies have shown that multiplication of the chromosome complement, tetraploidy in particular, may be a very early stage in tumor evolution that is followed by more specific chromosomal changes, usually occurring in a near-diploid cell population (18, 19). FISH analysis may thus have detected an earlier stage in the development of IFS.
Studies have distinguished CMN from Wilms tumor on the basis of dissimilar molecular profiles (20, 21). In 1980, Snyder et al. (22) distinguished the classic type of CMN from its cellular variant. Based on the results of a clinicopathologic study, these authors considered the cellular type of CMN to be a precursor of sarcoma. The finding of an ETV6-NTRK3 fusion gene in only the cellular type of CMN supports the notion that the classic and the cellular types of CMN are genetically distinct entities. Besides CMN/IFS, ETV6 rearrangements have been found in several lymphoid and myeloid hematologic neoplasms. In these conditions, as in CMN/IFS, ETV6 is frequently fused to genes coding for tyrosine-kinase receptors, such as, PDGFRB, JAK2, and ABL1 (23). However, trisomy 11, which often is concurrent with the 12;15 translocation in CMN/IFS, rarely is found in leukemias with ETV6 rearrangements, suggesting that trisomy 11 is a typical rearrangement for solid tumors with the ETV6-NTRK3 fusion. In the present study, trisomy 11 was found in only three of the six cases with ETV6 rearrangement. Therefore, gain of chromosome 11 may be a later event in tumor progression. On the other hand, this secondary aberration may be associated with still-unrecognized clinical or biological characteristics. Like other authors, we believe that the cellular type of CMN is the renal counterpart of IFS (11, 12).
CONCLUSION
Our results also indicate that interphase FISH may be highly useful for detecting genetic abnormalities in archived materials. Recently, several reports have indicated that reverse transcription-polymerase chain reaction (RT-PCR) analysis is more sensitive in detecting ETV6-NTRK3 gene fusion than conventional cytogenetic and immunohistochemical analysis (15, 16, 17). The present study demonstrates that interphase FISH analysis may be a comparably sensitive method for detecting ETV6 rearrangements in CMN/IFS. Compared with RT-PCR analysis, the FISH technique allows analysis of very small sample volumes, even if they consist of a few single nuclei. Because FISH may be used to detect rearrangements of a single gene, by disruption of the fluorescence pattern, it also may detect ETV6 rearrangements involving partners other than NTRK3. Such variant rearrangements would be missed by an ETV6-NTRK3-specific RT-PCR approach. Further analysis of cases exhibiting ETV6 rearrangements detected by FISH in the absence of ETV6-NTRK3 fusions detected by RT-PCR could lead to the identification of other oncogenic fusion products in CMN/IFS.
References
Bolande RP, Brough AJ, Izant RJ Jr . Congenital mesoblastic nephroma of infancy. A report of eight cases and the relationship to Wilms' tumor. Pediatrics 1967; 40: 272–278.
Bolande RP . Congenital mesoblastic nephroma of infancy. Perspect Pediatr Pathol 1973; 1: 227–250.
Beckwith JB . Mesenchymal renal neoplasms of infancy revisited. J Pediatr Surg 1974; 9: 803–805.
Dal Cin P, Lipcsei G, Hermand G, Boniver J, Van den Berghe H . Congenital mesoblastic nephroma and trisomy 11. Cancer Genet Cytogenet 1998; 103: 68–70.
Mascarello JT, Cajulis TR, Krous HF, Carpenter PM . Presence or absence of trisomy 11 is correlated with histologic subtype in congenital mesoblastic nephroma. Cancer Genet Cytogenet 1994; 77: 50–54.
Schofield DE, Yunis EJ, Fletcher JA . Chromosome aberrations in mesoblastic nephroma. Am J Pathol 1993; 143: 714–724.
Chung EB, Enzinger FM . Infantile fibrosarcoma. Cancer 1976; 38: 729–739.
Speleman F, Dal Cin P, De Potter K, Laureys G, Roels HJ, Leroy J, et al. Cytogenetic investigation of a case of congenital fibrosarcoma. Cancer Genet Cytogenet 1989; 39: 21–24.
Mitelman F . Catalog of chromosome aberrations in cancer (CD-ROM). New York: Wiley-Liss; 1998.
Knezevich SR, McFadden DE, Tao W, Lim JF, Sorensen PH . A novel ETV6-NTRK3 gene fusion in congenital fibrosarcoma. Nat Genet 1998; 18: 184–187.
Knezevich SR, Garnett MJ, Pysher TJ, Beckwith JB, Grundy PE, Sorensen PH . ETV6-NTRK3 gene fusions and trisomy 11 establish a histogenetic link between mesoblastic nephroma and congenital fibrosarcoma. Cancer Res 1998; 58: 5046–5048.
Rubin BP, Chen CJ, Morgan TW, Xiao S, Grier HE, Kozakewich HP, et al. Congenital mesoblastic nephroma t(12;15) is associated with ETV6-NTRK3 gene fusion: cytogenetic and molecular relationship to congenital (infantile) fibrosarcoma. Am J Pathol 1998; 153: 1451–1458.
Kuchinka BD, Kalousek DK, Lomax BL, Harrison KJ, Barrett IJ . Interphase cytogenetic analysis of single cell suspensions prepared from previously formalin-fixed and paraffin-embedded tissues. Mod Pathol 1995; 8: 183–186.
Kobayashi H, Montgomery KT, Bohlander SK, Adra CN, Lim BL, Kucherlapati RS, et al. Fluorescence in situ hybridization mapping of translocations and deletions involving the short arm of human chromosome 12 in malignant hematologic diseases. Blood 1994; 84: 3473–3482.
Bourgeois JM, Knezevich SR, Mathers JA, Sorensen PH . Molecular detection of the ETV6-NTRK3 gene fusion differentiates congenital fibrosarcoma from other childhood spindle cell tumors. Am J Surg Pathol 2000; 24: 937–946.
Argani P, Fritsch M, Kadkol SS, Schuster A, Beckwith JB, Perlman EJ . Detection of the ETV6-NTRK3 chimeric RNA of infantile fibrosarcoma/cellular congenital mesoblastic nephroma in paraffin-embedded tissue: application to challenging pediatric renal stromal tumors. Mod Pathol 2000; 13: 29–36.
Dubus P, Coindre JM, Groppi A, Jouan H, Ferrer J, Cohen C, et al. The detection of Tel-TrkC chimeric transcripts is more specific than TrkC immunoreactivity for the diagnosis of congenital fibrosarcoma. J Pathol 2001; 193: 88–94.
Heselmeyer K, Schrock E, du Manoir S, Blegen H, Shah K, Steinbeck R, et al. Gain of chromosome 3q defines the transition from severe dysplasia to invasive carcinoma of the uterine cervix. Proc Natl Acad Sci U S A 1996; 93: 479–484.
Steinbeck RG . The DNA content of chromosome division figures and interphase nuclei classifies ulcerative colitis. Eur J Cancer 1998; 34: 175–181.
Tomlinson GE, Argyle JC, Velasco S, Nisen PD . Molecular characterization of congenital mesoblastic nephroma and its distinction from Wilms tumor. Cancer 1992; 70: 2358–2361.
Kaneko Y, Homma C, Maseki N, Sakurai M, Hata J . Correlation of chromosome abnormalities with histological and clinical features in Wilms' and other childhood renal tumors. Cancer Res 1991; 51: 5937–5942.
Snyder HM III, Lack EE, Chetty-Baktavizian A, Bauer SB, Colodny AH, Retik AB . Congenital mesoblastic nephroma: relationship to other renal tumors of infancy. J Urol 1981; 126: 513–516.
Bohlander SK . Fusion genes in leukemia: an emerging network. Cytogenet Cell Genet 2000; 91: 52–56.
Author information
Authors and Affiliations
Corresponding author
Additional information
David Gisselsson is presently at the Department of Clinical Genetics, University Hospital, SE-221 85, Lund, Sweden.
Rights and permissions
About this article
Cite this article
Adem, C., Gisselsson, D., Dal Cin, P. et al. ETV6 Rearrangements in Patients with Infantile Fibrosarcomas and Congenital Mesoblastic Nephromas by Fluorescence In Situ Hybridization. Mod Pathol 14, 1246–1251 (2001). https://doi.org/10.1038/modpathol.3880469
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/modpathol.3880469
Keywords
This article is cited by
-
Recurrent EML4–NTRK3 fusions in infantile fibrosarcoma and congenital mesoblastic nephroma suggest a revised testing strategy
Modern Pathology (2018)
-
Fibrous hamartoma of infancy: a clinicopathologic study of 145 cases, including 2 with sarcomatous features
Modern Pathology (2017)
-
A rare malignant tumor of scalp in a 3-month-old Taiwanese infancy: case report of primitive myxoid mesenchymal tumor of infancy with molecular study
Medical Molecular Morphology (2013)