Abstract
A substantial number of patients with early-stage colorectal cancer relapse from metastatic disease. Identification of these patients by genetic profiling of their primary tumours may allow more informed follow-up and tailored administration of adjuvant therapy. Primary tumours from 70 patients with early-stage and largely microsatellite-stable colorectal cancer were profiled using metaphase-based comparative genomic hybridization (CGH) and the aberrations confirmed independently in a subset of patients using microarray-based CGH. Of the 70 cancers, 61 were amenable to CGH, and follow-up data was available from 56 patients. Genomic aberrations were correlated with patients' survival using univariate, multivariate and Kaplan–Meier survival curves. Metastatic primary tumours exhibited more complex genomic aberrations than non-metastatic primary tumours. Loss of chromosome 4p was an independent prognostic factor in early-stage colorectal cancer using multivariate analysis (Hazard ratio, 9.6; 95% CI, 3.3–28; P=0.0001). Loss of both chromosome arms 8p and 18q had a statistically significant negative effect on disease-free survival. Moreover, primary tumours with loss of both chromosomes 4 and 14q bestowed poorer prognosis than tumours with loss of any one of the two chromosomes (P<0.0001). Genetic profiling of primary tumours of patients with early-stage colorectal cancer is of significant value in identifying the subset of patients who may relapse with metastatic disease. Accordingly, the molecular genetic features of primary tumours should be considered in the mainstream management of patients with this specific stage of the disease.
Similar content being viewed by others
Main
It is estimated that one million cases of colorectal cancer are diagnosed worldwide each year, and half of them die from clinical complications and metastasis.1 The staging systems that associate the clinical–pathological behaviour of colorectal tumours with prognosis are the Astler–Coller modification of Dukes' or TNM classification. In these systems, stage A or I tumours are confined to the mucosa. Stage B or II, where the muscularis propria has been breached, is divided into B1 and B2 depending on whether or not the bowel wall has been breached. Stage C or III is spread to the lymph nodes with subdivision into C1 and C2 depending on whether or not the bowel wall has been breached or whether the apical lymph node is involved (C2). Stage D or IV is spread of the tumour to distant organs.2 This classification remains the most important parameter to direct clinical intervention. Accordingly, Patients presenting with Dukes' stages C and D (advanced disease) receive surgical and adjuvant therapy, whereas patients presenting with Dukes' A and B tumours (early stages without evident metastases) receive surgical therapy alone. Patients presenting with Dukes' stages A or B have better prognosis, but a significant proportion (10–40%) of these patients relapse with metastatic disease and die from it. There have been several attempts to identify this subset of patients both at histopathological and more often at molecular levels. At the genomic level, few studies have focused on early-stage colorectal cancer3, 4, 5, 6 and of those studies that focused on early-stage colorectal cancer, the analysis was limited to few chromosomes.7, 8 Therefore, there is currently a need to profile primary tumours from early-stage colorectal cancer using genome-wide scanning techniques that can also be integrated into current laboratory practices. We describe here the first attempt to study early-stage colorectal cancer at the genomic level using genome-wide scanning DNA methodologies and address the role of chromosomal aberrations in prognosis.
Materials and methods
Patients
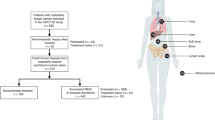
A total of 70 patients with early forms of sporadic colorectal cancers were examined in this study (49 from Glasgow and 21 from Kuwait). The two groups compared comprised of 27 patients with early-stage colorectal cancer who had no evidence of metastatic disease at the time of surgery, but subsequently relapsed with metastasis; and 43 age- and stage-matched patients who remained disease-free postoperatively. All patients received no treatment other than surgery except for 16 Dukes' B2 stage patients from Kuwait who were treated surgically and with six cycles of standard chemotherapy and were followed up prospectively. Patients were followed up at regular intervals for a minimum period of 2 years (range: 2–9 years) with a median of 5.5 years of follow-up for survivors, and 84% of survivors followed up for more than 3.5 years. Patients were clinically assessed for symptoms and signs of recurrence. Metastatic recurrences were confirmed radiologically and/or histologically or at post mortem. For assessing disease-free survival, patients who died of causes unrelated to cancer but had no evidence of metastatic recurrence at the time of death were censored (seven patients). Five patients were lost to follow-up and 10 patients had no recurrence time recorded but died from metastatic disease. For these patients, the date of death was used to calculate the disease-free survival.
Tumour Specifications and DNA Extraction
Haematoxylin–eosin sections were examined by two pathologists and scored for various histopathological parameters. Tumour staging was assessed using Dukes', Astler–Coller and AJCC systems. In the latter system, pT1 are tumours confined to the mucosa with submucosal invasion (two cases). pT2 are tumours that reached the muscularis propria or within it (26 cases), pT3 are tumours that breached the muscularis propria (33 cases) and pT4 (5 cases) are tumours that breached the peritoneal surface/serosa (pT4a) or invaded adjacent organs (pT4b). In all our cases there was no evidence of lymph node or distance metastasis at the time of surgery.
Tumours constituted no less than 50–90% of the used section.
DNA from Formalin-fixed paraffin-embedded tissue was extracted using Purgene kit (Gentra Systems, USA) according to manufacturer's protocol.
MSI Analysis
Microsatellite stability was assessed in all 70 patients using 13 dinucleotide markers (D4S2935, D4S2986, D4S1579, D4S1586, D4S1595, D4S2920, D14S283, D14S275, D14S49, D14S63, D14S267, D14S65 and D14S250). DNA for analysis of microsatellite stability was extracted separately from microdissected tissues as described previously.9
Comparative Genomic Hybridization
A measure of 2 μg of DNA was labeled using biotin nick translation reagent (Roche, GmbH) according to manufacturer's instructions.
Metaphase CGH
The test- and reference-labelled DNAs were mixed in equal concentrations (approx. 2.0 μg) and precipitated with 100 μg human Cot-1 DNA (Invitrogen, CA, USA). The CGH probe was pelleted by centrifugation and washed with 70% alcohol and vacuum dried then was dissolved in 16 μl CGH hybridization buffer (50% dextran sulfate in 2 × SSC containing 50% formamide) and denatured at 75°C for 10 min and reannealed at 37°C for 1 h. CGH metaphase slides were obtained from Vysis Inc (USA), and incubated in denaturation solution (70% deionized formamide in 2 × SSC) at 73°C for 3 min. The slides were then dehydrated and left to dry at 37°C for 10 min. After applying the probe, slides were incubated at 37°C in a humid environment for 72 h. The slides were finally washed with 70% formamide in 2 × SSC buffers and stained with fluorescent antibodies and DAPI as described previously.10 Images were captured by CCD camera (JVL, Japan) and then analysed by ISIS® software (Metasystems, GmbH). Typically, 10–20 metaphases were captured using the appropriate filters. The images were digitally stored and the green to red and vice versa ratios were calculated. Amplifications were scored if the median green to red ratio was above or equal to the 1.25 threshold and deletions were below or equal to the 0.75 threshold. Also, in all experiments, separate normal vs normal DNA hybridisation controls were included. The genetic aberration was scored if the probability representing 99% confidence intervals of the tumour vs normal profiles were outwith the 99% confidence interval of the normal vs normal control profiles.
Microarray CGH
Two human genomic microarrays were chosen for this study. First-generation microarray (Human BAC Array-3 Mb system, Spectral Genomics Inc.) comprising 1003 non-overlapping BAC/PAC clones with an average 3 Mb resolution (Human BAC Array 2–4 Mb system, Spectral Genomics Inc.). The microarrays included subtelomeric as well as the most common microdeletion syndrome regions.
After phenol/chloroform DNA extraction, patients' genomic DNA (test DNA) and normal reference genomic DNA were digested with EcoR1 for 18 h at 37°C and re-purified by Zymo Research's Clean and Concentrator™ (Orange, CA, USA). The reference and test DNAs were labelled with Cy3 and Cy5 by BioPrime random labelling kit (Invitrogen). The Cy3-labelled reference DNA and Cy5-labelled test DNA samples were combined with 55 μg of human Cot-1 DNA and 35 μg of sheared salmon sperm DNA. This mix was precipitated and rinsed in 70% ethanol. The pellets were dissolved in 10 μl of distilled water and mixed with 40 μl of hybridization solution (50% formamide, 10% dextran sulphate in 2 × SSC). The labelled DNAs were denatured at 72°C for 10 min followed by incubation at 37°C for 30 min to block repetitive sequences. Oppositely labelled DNA mixes were added onto duplicate microarray slides, hybridized, scanned and analysed according to the manufacturer's instructions (Spectral Genomics). In brief, the slides were scanned using GenePix 4000B scanner (Axon Ins. Inc., Union City, CA, USA). Cy3 and Cy5 images were scanned independently through two separate channels. Two 16-bit TIFF images were created per array. The data obtained were analysed using the Spectralware 1.0 software (Spectral Genomics). The software recognizes the regions of fluorescent signal, determines signal intensity, and compiles the data into a spreadsheet that links the fluorescent signal of every clone on the array to the clone name, its duplicate position on the array and its position in the genome. The normalized Cy5:Cy3 intensity ratios were computed for each of the two slides and plotted together for each chromosome. Arbitrarily, a ratio plot was assigned such that gains in DNA copy number at a particular locus are observed as the simultaneous deviation of the ratio plots from a modal value of 1.0, with the blue ratio plot showing a positive deviation (to the right) while the red ratio plot shows a negative deviation at the same locus (to the left). Conversely, DNA copy number losses show the opposite pattern.
Apoptosis Assay
The Roche in situ cell death detection kit was used to detect apoptotic cells according to the manufacturer's protocol (Roche, Germany).
Statistical Analysis
The primary outcome was disease-free survival. Disease-free survival was defined as the time from study entry to first confirmed metastatic relapse. Univariate and multivariate Cox's proportional hazards models were used to analyse the effect of clinical characteristics on the survival of patients. Two-sided χ2-test or Fisher's exact tests were used for significance testing of contingency tables containing categorical variables. Mann–Whitney U test was used to compare means of pairs of independent samples from continuous variables and Kruskal–Wallis test was used to compare means of more than two independent samples from continuous variables. Follow-up-to-event outcomes were analysed by Kaplan–Meier survival curves and compared using Log-rank tests. All reported P-values are two-sided. Data were analysed using the SPSS v11.0 software package.
Results
Clinicopathological Correlations
There was no significant association between sex, age and left or right tumour location with respect to metastatic recurrence. Similarly, tumour differentiation, mitotic and apoptotic counts, lymphatic and vascular invasion, and the depth of invasion did not significantly correlate with metastatic relapse (Table 1).
Univariate analysis of the clinicopathological variables and disease-free survival showed that rectal location was significantly associated with shorter disease-free survival (Hazard ratio, 2.3; 95% CI, 1.064–4.983; P=0.034), while Dukes' stage A and extracellular mucin pooling were associated significantly with longer disease-free survival (Hazard ratio, 0.3; 95% CI, 0.118–0.9; P=0.031 and 0.213; 95% CI, 0.05–0.9; P=0.036, respectively). All the remaining variables (age, sex, tumour site (left vs right), differentiation, nuclear grade, apoptosis and mitosis indexes, lymphatic and vascular invasion and depth of invasion showed no significant correlation with survival. Multivariate analysis of all clinicopathological variables demonstrated that only rectal location (Hazard ratio, 4.6; 95% CI, 1.7–12.4; P=0.003) and T-stage 3/4 (Hazard ratio, 4.5; 95% CI, 1.6–12.8; P=0.005) could predict survival independently. These results confirm the limited usefulness of current clinicopathological parameters in identifying patients at risk from metastatic relapse in early-stage CRC.
Microsatellite Analysis of the Primary Tumours
Amplification of 13 dinucleotide repeats was used to study the microsatellite status of the tumours. MSI in ≥30% of the analysed loci were classified as high microsatellite instability (one patient who was excluded from the survival analysis), allelic instability in one or two of the 13 microsatellites or less than 30% of the loci analysed were considered as low microsatellite instability (13 patients). Samples with no MSI were classified as stable (50 patients). In six patients, microsatellite stability could not be assessed owing to the failure of amplifying the DNA from normal or tumour tissues.
Metaphase-CGH Analysis of the Primary Tumours
To seek genomic markers capable of predicting metastatic behaviour in this cohort of largely microsatellite-stable early-staged CRC, we subjected the primary cancers to CGH. Interrogating the genome by metaphase-CGH was informative in 61 primary tumours. In nine cases, the CGH was of poor quality and not amenable to further analysis. The mean number of genetic alterations per sample was 12.8±5.98 (range, 1–23) in disease-free patients and 18.5±7.6 (range, 3–33) in patients who had metastatic relapse (t-test P<0.003).
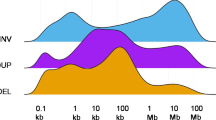
Particular non-random chromosomal aberrations were frequent in early-staged CRC, including increase in copy number of chromosome arms 1q, 7, 8q, 13q, 17q and 20q and loss of chromosomes 1p, 4, 5q, 8p, 9p, 14q, 15q, 17p and 18 (Figure 1). The commonest aberrations were increase in copy number of chromosomes 7 and 20q. Chromosomal deletions were most frequently observed on chromosomes 18 and 4 (Table 2).
Chromosomally selected cumulative comparative genomic hybridization profile of early-stage ((a) 26, delayed metastasis; (b) 35, absence of metastasis) cancer patients. Each bar represents an aberration in a tumour. Bars to the left of the ideograms represent loss, while bars on the right of the ideograms denote gains. Chromosomal number is shown below each ideogram.
Chromosomal aberrations signified by gain of chromosome arm 1q and loss of chromosomal arms 1p, 4, 5q, 8p, 9p and 14q exhibited statistically significant association with metastatic recurrence (Table 2).
Univariate analysis of chromosomal aberrations showed that patients with gain of chromosomal arm 1q and loss of chromosomal arms 1p, 4p, 4q, 5q, 8p, 9p and 14q had significantly shorter disease-free survival than those without them. This association was maintained after adjustment for adjuvant therapy (Table 3).
Multivariate analysis of the overall chromosomal aberrations as covariates in the multiple regression model showed the deletion of chromosome arms 4p and 5q to be independent prognostic factors (Hazard ratio, 3.9; 95% CI, 1.7–8.8; P=0.001 and Hazard ratio, 2.9; 95% CI, 1.3–6.55; P=0.008, respectively). In addition, deletion of chromosome arm 4p was the most significant and independent prognostic factor when all variables were included in the multivariate regression equation (Hazard ratio, 9.6; 95% CI, 3.3–28; P=0.0001). Kaplan–Meier survival curves illustrated the association between reduced disease-free survival and chromosome 4 deletions (Figure 2a). The relationship between reduced disease-free survival and deletion of chromosome 4 was maintained after adjustments for Dukes' stage (limiting the data to Dukes' stage B1; Figure 2b), tumour site (limiting data to colon cancer; Figure 2c) or country of origin (Glasgow patients only; data not shown). The associations between chromosomal aberrations and various clinicopathological parameters are shown in Table 4. Cancers with deleted chromosome arm 9p had significantly higher mitotic index than those with normal copy number of the chromosome arm. Also, tumours with loss of chromosome 4 had a significantly lower apoptotic index than those retaining it (Table 4).
Kaplan–Meier plots of disease-free survival of patients with early-stage colorectal cancers in relation to loss of chromosome 4. (a) Effect of chromosome 4 loss on disease-free survival in 56 patients without stratification. (b) Data limited to Dukes' stage B1. (c) Data limited to colonic cancer. Solid black lines represent patients with loss of chromosome 4 and dashed black lines are patients whose tumours maintained chromosome 4 copy number. P-values represent the Log-rank test.
We then investigated the additive effects of several aberrations on disease-free survival. Patients who lost both chromosome arms 8p and 18q demonstrated significantly shorter disease-free survival (mean, 3.18 years; 95% CI, 2.04–4.31) than those with normal copies or those that lost either chromosomal arm combined (mean, 6.07 years; 95% CI, 4.94–7.2; P=0.015). Also, patients who lost both chromosomes 4 and 14q had a mean disease-free survival of 1.85 years (95% CI, 0.93–2.77) compared to those who lost either one of the chromosomes (mean, 4.58 years; 95% CI, 3.51–5.65) or patients who maintained normal chromosomal copy number (mean, 6.93 years; 95% CI, 5.63–8.23). The above finding was independent of Dukes' stage or tumour location (Figure 3).
Kaplan–Meier plots of disease-free survival of patients with early-stage colorectal cancers in relation to loss of chromosomes 4 and 14q. (a) Effect of chromosome 4 and 14q loss on disease-free survival in 56 patients without stratification. (b) Data limited to Dukes' stage B1. (c) Data limited to colonic cancer. Solid black lines represent patients with loss of both chromosomes 4 and 14q and dashed black lines are patients whose tumours lost either chromosome 4 or 14q and grey lines represent those who maintained both chromosomes. P-values represent the Log-rank test.
Microarray-CGH Analysis
DNA from 10 patients, known to have specific aberrations by metaphase CGH, were subjected to microarray-CGH comprising 1003 non-overlapping BAC/PAC clones with an average 3 Mb resolution. This step was necessary for two reasons; first, microarray-CGH offers an independent method to confirm the results obtained by metaphase CGH. Second, it is a method that is not prone to operator bias in addition to having a higher resolution than classical CGH. Figure 4 shows the relative copy numbers of chromosomes 4, 8, 14 and 18 compared to a normal reference of the BACs. In all patients who relapsed with metastasis, loss of 4p copy numbers was pronounced within the subregion covered by BAC clones RP11-47C1 to RP11-17I9. Loss of 8p copy numbers was localized to subregions covered by BAC clones RP11-91J19 to RP11-51C1 at 8p23–8p21 and BAC clones RP11-57I3 to RP11-90P5 at 8p12–8p11.2, while amplifications were localized to 8q24 region. Loss of chromosome 18 copy numbers was more extensive and covered the subregions bounded by BAC clones CTB-74G18 to RP11-90L15. Loss of chromosome 14 copy numbers was localized to subregion covered by overlapping BAC clones RP11-81F9 to RP11-79M1 that map to 14q 11.2–14q23 and another subregion covered by overlapping clones RP11-90H21 to CTC-200D12, which has been localized to 14q32–14q32.33. These findings were consistent with the metaphase-CGH data (Figure 4).
Selected microarray-CGH profile from patients with early-stage colorectal cancer that relapsed with metastatic disease. The dotted lines represent modal value of ratio of 1.0. Arrows indicate that change in DNA copy number at a particular locus are observed as the simultaneous deviation of the ratio plots from the modal value of 1.0 (dotted line), with gains showing positive deviation (to the right) while negative deviation to the left indicates a loss. The linear order of the microarray clones, specified in the result section, is reconstituted in the ratio plots consistent with an ideogram, such that the p terminus is to the top and the q terminus is towards the bottom of the plot. The numbers denote chromosomal numbers. Notice the concordance of these aberrations with those illustrated by metaphase CGH (Figure 1a).
Discussion
This paper demonstrated a relationship between loss of chromosomes 1p, 4, 5q, 8p, 9p, 14q, 18p and gain of chromosomes 1q, 8q and 13q and metastatic recurrence of early-staged CRC after surgical intervention. Overall, the chromosomal aberrations described here are consistent with chromosomal changes previously reported for colorectal cancer.10, 11, 12, 13
Although the number of genomic aberrations may potentially substitute for chromosomal instability, it was paramount to establish the MSI status of the tumours because it could have had a profound impact on the analysis of disease-free survival. It is well established that cancers with high microsatellite instability have better survival than microsatellite stable tumours. Most of cancers analysed in this study were microsatellite stable. This excludes microsatellite instability as a significant source of bias in survival data estimation. Dukes' stage (A vs B1 vs B2), tumour site (colon vs rectum) and adjuvant therapy are other confounding factors that could potentially affect the analysis of disease-free survival in this cohort. However, we ensured appropriate data stratification to minimize their effects to the best of our abilities. For example, disease-free survival calculations were limited to Dukes' B1 patients who received no therapy other than surgery and so forth.
The association between loss of chromosome arm 1p and poor survival has been reported by several investigators using different DNA interrogation techniques ranging from allelic imbalance to genome-wide CGH scanning.4, 14, 15, 16 The region of loss we mapped, namely 1p21–31.1 is the same area mapped previously.4
Chromosome 4 loss is a common aberration in advanced colorectal cancers and metastases. In our study, the minimal deleted areas map to 4p14–16, 4q24–28 and 4q32–35 consistent with previously published data.3, 4, 17 The reason for such an association remains to be discovered. However, we and De Angelis et al13 found an inverse relationship between chromosome 4 loss and reduced apoptosis.
Loss of chromosome 5q is common event in adenomas and colorectal cancer. The association of Loss of 5q13.3–23.3 with shorter disease-free survival, to the best of our knowledge, has not been reported before in early-stage colorectal cancer. The study of Choi et al examined loss of 5q in colorectal cancer using five microsatellite markers and found no association between 5q loss and overall survival. However, 5q loss was associated with higher TNM tumour stages in their patients.6 We believe that this discrepancy could have stemmed from the fact that our study focused on early-stage colorectal cancer and benefited from stratifying the data for site (colon vs rectum), treatment and microsatellite stability status.
Loss of chromosome arm 8p is a frequent aberration in colorectal cancer. Consistent with our finding, numerous studies established an association between loss of chromosome arm 8p and reduced survival.4, 18, 19, 20
Loss of chromosome arm 9p is not an infrequent event in colorectal cancer.13, 21 We identified a significant association between 9p loss and high mitotic index in early-stage colorectal cancer. This is consistent with the localization of cyclin-dependent kinase inhibitor 2A (p16 INK4A and p14 ARF) and cyclin-dependent kinase inhibitor 2B (p15) to chromosome arm 9p.
Loss of chromosome 14q has been reported in colorectal cancer.22 More importantly it has been documented in advanced colorectal cancer stages as well as in metastases.23
Our data indicated that metastatic primary tumours had higher number of genetic aberrations than non-metastatic tumours. Accordingly, primary tumours that lost both chromosome arms 8p and 18q had worse prognosis than tumours with loss of any one of the chromosomes or those that retain them. This is consistent with recent digital SNP analysis.8 Also, loss of both chromosomes 4 and 14q was associated with shorter disease-free survival probably indicates a selective advantage for tumours harbouring these losses. The precise molecular and cellular mechanisms behind such selection remain to be elucidated and await further refinement of these areas.
Our results showed, albeit modestly, that administering chemotherapy could be of benefit to patients with early-stage colorectal cancer who have lost chromosome 4 or chromosome arm 8p (Table 3). Therefore, loss of these chromosomes is not only of prognostic value but may also be predictive in that the aberrations could indicate the extent of benefit from adjuvant therapies (Figure 5).
Graphical illustrations of prognostic and predictive powers for loss of chromosomes 4 (a) and 8p (b). Notice that loss of either marker can predict prognosis after surgical intervention alone. However, the 5-year disease-free survival improved after treatment with adjuvant therapy particularly in patients with loss of chromosome 4 indicating its usefulness as a marker for predicting response to treatment. Adapted from Hayes et al.24
Our study identified several independent genetic markers that could predict metastatic behaviour of early-stage CRC. This highly suggests that patients with tumours harbouring these aberrations had micrometastases at the time of surgery that were undetectable by current clinical and radiological methods, and eventually grew to form clinically detectable secondary tumours. The novel aspect of this study was to show that the identification of a metastatic signature by profiling primary tumours at the DNA level might be possible. This finding gains in significance given the high prevalence of CRC and the urgent demand for molecular markers that can predict metastasis especially in early-staged cancer. To that end, a larger prospective study addressing the influence of detecting these aberrations on prognosis and targeted therapy is now indicated.
References
Parkin DM, Bray F, Ferlay J, et al. Global cancer statistics, 2002. CA Cancer J Clin 2005;55:74–108.
Hermanek P . pTNM and residual tumor classifications: problems of assessment and prognostic significance. World J Surg 1995;19:184–190.
Bardi G, Fenger C, Johansson B, et al. Tumor karyotype predicts clinical outcome in colorectal cancer patients. J Clin Oncol 2004;22:2623–2634.
De Angelis PM, Stokke T, Beigi M, et al. Prognostic significance of recurrent chromosomal aberrations detected by comparative genomic hybridization in sporadic colorectal cancer. Int J Colorectal Dis 2001; 16:38–45.
Diep CB, Thorstensen L, Meling GI, et al. Genetic tumor markers with prognostic impact in Dukes' stages B and C colorectal cancer patients. J Clin Oncol 2003;21:820–829.
Choi SW, Lee KJ, Bae YA, et al. Genetic classification of colorectal cancer based on chromosomal loss and microsatellite instability predicts survival. Clin Cancer Res 2002;8:2311–2322.
Martinez-Lopez E, Abad A, Font A, et al. Allelic loss on chromosome 18q as a prognostic marker in stage II colorectal cancer. Gastroenterology 1998;114:1180–1187.
Zhou W, Goodman SN, Galizia G, et al. Counting alleles to predict recurrence of early-stage colorectal cancers. Lancet 2002;359:219–225.
Al-Mulla F, Going JJ, Sowden ET, et al. Heterogeneity of mutant vs wild-type Ki-ras in primary and metastatic colorectal carcinomas, and association of codon-12 valine with early mortality. J Pathol 1998;185:130–138.
Al-Mulla F, Keith WN, Pickford IR, et al. Comparative genomic hybridization analysis of primary colorectal carcinomas and their synchronous metastases. Genes Chromosomes Cancer 1999;24:306–314.
Ried T, Knutzen R, Steinbeck R, et al. Comparative genomic hybridization reveals a specific pattern of chromosomal gains and losses during the genesis of colorectal tumors. Genes Chromosomes Cancer 1996; 15:234–245.
Nakao K, Shibusawa M, Ishihara A, et al. Genetic changes in colorectal carcinoma tumors with liver metastases analyzed by comparative genomic hybridization and DNA ploidy. Cancer 2001;91:721–726.
De Angelis PM, Clausen OP, Schjolberg A, et al. Chromosomal gains and losses in primary colorectal carcinomas detected by CGH and their associations with tumour DNA ploidy, genotypes and phenotypes. Br J Cancer 1999;80:526–535.
Thorstensen L, Qvist H, Heim S, et al. Evaluation of 1p losses in primary carcinomas, local recurrences and peripheral metastases from colorectal cancer patients. Neoplasia 2000;2:514–522.
Ogunbiyi OA, Goodfellow PJ, Gagliardi G, et al. Prognostic value of chromosome 1p allelic loss in colon cancer. Gastroenterology 1997;113:761–766.
Bardi G, Sukhikh T, Pandis N, et al. Karyotypic characterization of colorectal adenocarcinomas. Genes Chromosomes Cancer 1995;12:97–109.
Arribas R, Ribas M, Risques RA, et al. Prospective assessment of allelic losses at 4p14-16 in colorectal cancer: two mutational patterns and a locus associated with poorer survival. Clin Cancer Res 1999;5:3454–3459.
Halling KC, French AJ, McDonnell SK, et al. Microsatellite instability and 8p allelic imbalance in stage B2 and C colorectal cancers. J Natl Cancer Inst 1999;91:1295–1303.
Kelemen PR, Yaremko ML, Kim AH, et al. Loss of heterozygosity in 8p is associated with microinvasion in colorectal carcinoma. Genes Chromosomes Cancer 1994;11:195–198.
Kochhar R, Halling KC, McDonnell S, et al. Allelic imbalance and microsatellite instability in resected Duke's D colorectal cancer. Diagn Mol Pathol 1997; 6:78–84.
Weber TK, Conroy J, Keitz B, et al. Genome-wide allelotyping indicates increased loss of heterozygosity on 9p and 14q in early age of onset colorectal cancer. Cytogenet Cell Genet 1999;86:142–147.
Bando T, Kato Y, Ihara Y, et al. Loss of heterozygosity of 14q32 in colorectal carcinoma. Cancer Genet Cytogenet 1999;111:161–165.
Korn WM, Yasutake T, Kuo WL, et al. Chromosome arm 20q gains and other genomic alterations in colorectal cancer metastatic to liver, as analyzed by comparative genomic hybridization and fluorescence in situ hybridization. Genes Chromosomes Cancer 1999;25:82–90.
Hayes DF, Trock B, Harris AL . Assessing the clinical impact of prognostic factors: when is ‘statistically significant’ clinically useful? Breast Cancer Res Treat 1998;52:305–319.
Acknowledgements
This study was supported by Kuwait Foundation for the Advancement of Sciences grant number 99-07-07 and Kuwait University Shared Facility Grant GM/0101. We thank Professor David Hole for the follow-up data from the Scottish patients. Our thanks go for Bincy Ancy Mathew for her help with CGH, and Dr Shirley George for performing the TUNEL assay.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Al-Mulla, F., Behbehani, A., Bitar, M. et al. Genetic profiling of stage I and II colorectal cancer may predict metastatic relapse. Mod Pathol 19, 648–658 (2006). https://doi.org/10.1038/modpathol.3800564
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/modpathol.3800564
Keywords
This article is cited by
-
Whole genome analysis identifies the association of TP53 genomic deletions with lower survival in Stage III colorectal cancer
Scientific Reports (2020)
-
Copy number alterations and allelic ratio in relation to recurrence of rectal cancer
BMC Genomics (2015)
-
The relationship between genetic profiling, clinicopathological factors and survival in patients undergoing surgery for node-negative colorectal cancer: 10-year follow-up
Journal of Cancer Research and Clinical Oncology (2013)
-
Deletion of chromosome 4q predicts outcome in Stage II colon cancer patients
Cellular Oncology (2011)
-
High Frequency of Chromosome 14 Deletion in Early-Onset Colon Cancer
Diseases of the Colon & Rectum (2007)