Abstract
Type 1 and type 2 EWS-FLI1 fusion products result from variation in breakpoint locations arising from the t(11;22)(q24;q12) recurrent chromosomal translocation in Ewing’s sarcoma family tumors (EFT). Previously, studies from our institution (updated in the present communication at a median follow-up of more than 6 years) and others suggested a prognostic difference for EFT patients with localized disease depending on the type of EWS-FLI1 fusion present in the tumor. It has been suggested that the observed clinical discrepancies result from different transactivation potentials of the various EWS-FLI1 fusion proteins. In an attempt to identify genes whose expression levels are differentially modulated by structurally different EWS-FLI1 transcription factors, we have used two related PCR-based subtractive approaches, cDNA representational difference analysis (cDNA-RDA) and linker-capture subtraction (LCS) to compare transcript representations in cDNA pools of type 1 versus type 2 EFT cell lines. About 800 clones obtained by the two approaches were analyzed by dot blot hybridization to cDNA pools. Eighty-six clones showing the highest variability in signal intensities on the dot blots were further hybridized to individual EFT cell line RNAs on Northern blots, and four of them were additionally studied by real-time quantitative PCR (RTQ-PCR). Although interindividual variations in gene expression patterns in the range of one- to several-fold were observed, no correlation to specific EWS-FLI1 fusion types could be identified. Among the genes differentially expressed in individual EFT cell lines are several previously implicated in tumor growth, invasion, and metastasis. Although our data may have revealed candidate genes whose composite expression pattern may be relevant for the biology of individual EFT, they do not support a role of distinct EWS-FLI1 fusion types for EFT prognosis based on different transactivation potentials.
Similar content being viewed by others
Introduction
The course of normal cellular development, as well as that of neoplastic transformation leading to cancer, is believed to be driven by changes in gene expression. The identification and characterization of differentially expressed genes associated with different prognostic subtypes of cancer may help us to better adjust treatment to the specific biology of the disease in individual patients and discover new targets for therapeutic intervention. Methods to identify variably expressed genes in tissues have included differential screening of cDNA libraries, various forms of subtractive hybridization, and differential display reverse transcriptase polymerase chain reaction (DDRT-PCR) (Liang and Pardee, 1992). DDRT-PCR relies on random-primed amplification of total RNA populations aiming at the identification and isolation of differential bands in sequencing gels. Although it is a powerful method, it is hampered somewhat by a relatively large number of false positives and its preferential isolation of 3′ untranslated regions. Subtractive hybridization using conventional methods has been a very tedious approach, and this has led to a series of developments in which PCR has been included (Byrne et al, 1995; Hubank and Schatz, 1994; Wang and Brown, 1991; Zeng et al, 1994). Representational difference analysis (RDA) is a combination of subtraction and kinetic enrichment, coupled with subsequent amplification, which was originally developed to isolate differences between complex DNA genomes (Lisitsyn and Wigler, 1993; Lisitsyn et al, 1995). Whereas DDRT-PCR amplifies fragments from all represented mRNA species, RDA has the advantage of eliminating fragments common to both populations, leaving only the differences.
The Ewing’s sarcoma family tumors (EFT) are characterized on the molecular level by the t(11;22) or t(21;22) recurrent chromosomal translocation in about 85% and 10% of cases, respectively. In rare EFT other translocations also involving chromosome 22 have been observed. These chromosomal rearrangements result in the fusion of the DNA-binding domain of a member of the ets transcription factor family to the amino terminal domain of a putative RNA-binding protein, EWS, of as yet unknown function. The resultant chimeric protein is a potent transcriptional activator. The two most common fusions join EWS exon 7 in frame with either exon 6 (type 1 fusion) or exon 5 (type 2 fusion) of FLI1. EWS-FLI1 transforms NIH 3T3 cells, which suggests that this molecule plays an active role in tumorigenesis (Lessnick et al, 1995; May et al, 1993).
Ewing tumor patients can be assigned to one of two different risk groups depending on the extent of the disease at diagnosis. Although about 60% of patients with localized EFT can be cured by modern multimodal treatment regimens, patients with metastases are at an 80% risk of succumbing to the disease (Kovar, 1998). A significantly better treatment outcome was reported for a limited number of patients with localized disease when the tumors expressed a type 1 EWS-FLI1 fusion compared with tumors that expressed a non-type 1 EWS-FLI1 fusion. This introduced the question of whether EWS-FLI1 gene fusion type might serve as a prognostic molecular indicator in this group of patients (Zoubek et al, 1996). More recently, data on independent patient cohorts were obtained that support the idea of a better treatment outcome (de Alava et al, 1998) at the same time that they contradict a putative prognostic impact of EWS-FLI1 fusion type (Delattre, personal communication).
Recently, a difference in the transactivation potential of different EWS-FLI1 fusion types has been described. Reporter gene assays performed in EWS-FLI1-transfected Hela and NIH 3T3 cells, as well as in EFT cell lines of different EWS-FLI1 fusion type, revealed an approximately two-fold weaker transcriptional activity of type 1 versus non-type 1 EWS-FLI1 on an ets responsive promoter element. This suggests that if different EWS-FLI1 fusion types are associated with different clinical behavior, this association may be caused by qualitatively or quantitatively different EWS-FLI1-dependent gene expression patterns (Lin et al, 1999). Although some advances have been made in isolating EWS-FLI1 target genes (Braun et al, 1995; Hahm et al, 1999; May et al, 1997; Thompson et al, 1996), no data exist regarding differential expression of genes in EFT carrying the different fusion products. To define the biological basis for the observed differences in clinical behavior, we sought to define discrepancies in gene expression profiles between EFT cell lines of the most frequent fusion types 1 and 2 using two related PCR-based subtraction strategies: cDNA representational difference analysis (cDNA-RDA) and linker-capture subtraction (LCS). We now report the identification of several genes whose composite expression differences between EFT cell lines carrying the two different EWS-FLI1 fusion types may reflect differences relevant to the course of the disease in patients. However, so far, no fusion-type associated differences in gene expression patterns that might account for the reported clinical heterogeneities were identified.
Results
Identification of Genes Differentially Expressed between Type 1 and Type 2 EFT Cell Lines
An earlier clinical study compared type 1 EWS-FLI1–expressing EFT to a heterogenous group of tumors expressing a variety of other fusion types at a median observation time of 30 months (Zoubek et al, 1996). Figure 1A presents an update of treatment results for the cohort of patients with localized disease described previously. After a median observation time of more than 6 years, Kaplan-Meier curves for the two groups still diverged from each other, with a probability for relapse-free survival at 5 years of 0.64 and 0.41, respectively (p = 0.051). However, the predominant gene rearrangements in the non-type 1 group of patients lead to type 2 EWS-FLI1 gene fusions (67%). Figure 1B depicts a comparison between type 1 and type 2 EWS-FLI1–expressing localized tumors only. Kaplan-Meier curves for these more tightly defined groups of patients reflected the results obtained for the whole group of patients with localized disease, although the difference in relapse-free survival (probability at 5 years: 0.64 for type 1 EWS-FLI1 fusions versus 0.40 for type 2 fusions) lacked statistical significance (p = 0.156), possibly because of the relatively small number of patients. For simplicity, we therefore decided to restrict our comparison of gene expression patterns to type 1 and type 2 EWS-FLI1–expressing EFT.
Kaplan-Meier curves for (A) update of relapse-free survival (RFS) of EFT patients with localized disease of either type 1 (7/6) (p* = 0.64 ± 0.07) or other (p* = 0.41 ± 0.08) EWS-FLI1 exon combinations at a medium observation time of 6 years and 4 months, p = 0.051, and (B) RFS of patients with either type 1 (p* = 0.64 ± 0.07) or type 2 (7/5) (p* = 0.40 ± 0.10) EWS-FLI1 exonic fusions at a median observation time of 6 years and 2 months, p = 0.156. p*, Probability of survival at 5 years.
To identify genes differentially expressed in association with distinct EWS-FLI1 fusion types, the starting material for the subtractive screen had to meet three criteria: (a) a high tumor-cell content to avoid the masking of tumor-specific gene expression by contaminating healthy tissue, (b) availability in sufficiently high amounts to allow for the confirmation of differential expression by complementary approaches (ie, repeated Northern blotting and real-time PCR), and (c) minimization of random interindividual differences in gene expression. We therefore chose to use cell lines instead of primary EFT. A mixture of equal amounts of total RNA from each of seven exponentially growing cell lines per individual fusion type has been employed to enrich for fusion-type–associated differences in gene expression relative to intratype variations. Two closely related PCR-coupled subtractive hybridization procedures were conducted, cDNA-RDA and LCS. Subtraction was performed in both directions, type 1 minus type 2 cell lines and vice versa. Initially, the subtraction process in each approach was carried out three times, using the tester:driver amplicon ratios of 1:100, 1:800, and 1:8,000 in three successive rounds of subtraction, respectively. Because the PCR products from the third round of subtraction did not show any enhancement in intensity over the second round products, as was previously observed (Chang et al, 1998; Geng et al, 1998; O’Neill and Sinclair, 1997), PCR fragments from the second round of subtractive hybridization were cloned into pGEM-T Easy (Promega, Madison, Wisconsin) for further analysis to achieve a high degree of sensitivity and diversity (Welford et al, 1998).
Dot-Blot Hybridization and Northern Blot Analysis
Eight hundred white colonies for each subtracted library were picked at random and were analyzed by PCR to confirm the presence and the respective sizes of inserts. Eight clones failed to produce PCR products, and five clones did not have any inserts. With the remaining clones, we prepared replica DNA dot-blots and probed them with the radiolabeled type 1 and type 2 driver cDNA pools, respectively (Fig. 2). A visual comparison of the hybridization intensity for each clone in the two replica membranes allowed for a rough estimate of relative transcript abundancies in type 1 versus type 2 cDNA pools and was used to isolate candidate positives described in Tables 1 and 2. Most of the genes that segregated with the different fusion types are known genes, some of which have been reported to be differentially expressed in cancers and to affect cell proliferation and tumor progression (ie, elongation factor 1-alpha, translational controlled tumor protein, v-fos transformation effector protein). Twelve clones, 11 of which were picked by LCS and 1 by cDNA-RDA, had no matches to any sequence in the Genbank. When used as probes in Northern blot analyses, no signals were obtained. Several high-abundance transcripts, such as those from cytoskeletal-associated genes and from some ribosomal and housekeeping genes, were also picked up by the two differential screening methods, but many of them were found to be constitutively expressed in cells irrespective of the fusion type.
Dot-blot screening assay for differentially expressed genes. Replica DNA dot-blots were prepared from randomly picked colonies for type1 minus type2 (A1-D12) and type 2 minus type1 (E1-H12) subtractions and probed with labeled driver cDNAs for type 1 (upper panel) and type 2 cells (lower panel). Clones differentially represented are indicated by arrows.
Northern blot analyses of RNAs from individual type 1- and type 2-expressing cell lines (originally contained in the tester and driver cDNA pools) using selected clones as probes was performed to confirm differential expression of candidate positives from the dot-blot assays (Fig. 3). None of the 86 different clones yielded a consistent EWS-FLI1 fusion type-specific difference in expression levels. Rather, very heterogenous expression levels with intratype and intertype variations were found. The expression of the Fibronectin gene, which was obtained in the type 1 minus type 2 subtraction, was found to be more than 100-fold higher in the type 1 cell line A673 and not expressed in any of the type 2 cell lines. Steady-state transcript levels of individual genes were quantified by phosphor-imaging and normalized by comparison to the hybridization signal obtained with an 18S rDNA probe. The degree of expression differences of 13 confirmed positives was found to range from one- to about two-fold (Fig. 4). Statistical analysis, using the Kruskal-Wallis test, of results obtained by phosphorimaging did not give any significant differences in the expression levels of type 1 versus type 2 cell lines for any isolated clone, indicating lack of correlation with EWS-FLI1 fusion type. For most clones, distinctively high transcript abundance in single cell lines may explain their overrepresentation in starting RNA pools, which enables their isolation by high-sensitivity subtractive screening.
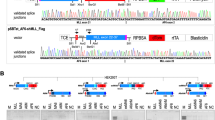
Northern blots to confirm the differential expression of candidate positive clones detected by RDA and LCS (*) in type 1 vs type 2 EWS-FLI1 expressing Ewing tumor cells. A to C, Representative fragments recovered from type 1 minus type 2 subtraction (rpS3, TAXREB107, α-enolase, rpS11, UBE1, α-tubulin, rpL9) and from type 2 minus type 1 subtraction (EF1α, VDAC) are shown. Total RNA form either cell type used in the pool was probed with radiolabeled cDNA from candidate positives as shown in autoradiograms. 18S rRNA was used to control for equal sample loading.
Graphic representations of data obtained by phospho-imager analysis of chosen Northern blots hybridized with two RDA probes. Data are normalized to 18S rRNA control. Type 1 minus type 2 clones: 1-1 (Human mitochondrial gene), 1-4 (UBE1), 1-5 (Human fibronectin gene), 1S-3 (VDAC). Type 2 minus type 1 clones: 2-8 (rpS3a), 2-16 (rpS16), 2-18 (Human mitochondrial cytochrome oxidase subunit II), 2-20 (TAXREB107), 2-23 (Human TEGT gene), 2-25 (CAPN4), 2-29 (Human calmodulin gene), 2-30 (Human MHC gene), 2-32 (Human clone 90L6 on 22q11.21).
Validation of Northern Blot Data with Real-Time Quantitative PCR
To further investigate the reliability of our Northern blot data, we measured the expression levels of four candidate clones whose relative expression levels between cell lines with the different fusion transcripts were close to significance, using the TaqMan 5′nuclease fluorogenic quantitative PCR assay (RTQ-PCR). Figure 5 illustrates the expression pattern and the quantitative expression level of each of the four genes as determined by RTQ-PCR. Statistically, no significant differences in the steady state transcript levels could be detected for all the genes analyzed. Although the transcriptional level of a given gene varies dramatically, no group-specific up- or down-regulation could be detected.
Discussion
In a series of 83 localized EFT, treated according to the European Intergroup Cooperative Ewing’s Sarcoma Study (CESS and EICESS) protocols, we observed an EWS-FLI1 fusion type-associated difference in long-term event-free survival. This difference was also reflected by a comparison between type 1- and type 2-expressing localized tumors only, although statistically the difference was not significant and was restricted to localized disease. Primary metastatic disease was generally associated with an adverse prognosis (data not shown). This study was performed to evaluate the molecular basis for the fusion type-associated difference in the clinical course of the disease. In a previously published study by Lin et al (1999), reporter gene transactivation by type 1 and other types of EWS-FLI1 fusions in EFT cell lines was found to differ up to two-fold. To achieve maximum sensitivity for differences in endogenous gene expression of EFT cells, we focused on homogenous tumor cell populations by using EFT cell lines. Although in patients, several factors may influence prognoses differently and EWS-FLI1 fusion type-associated gene expression differences may impact on localized disease only, we expected these differences to be present in EFT cell lines regardless of the extent of the disease in the patients from which the cell lines were derived. Our primary goals were to rapidly screen pools of EFT cell lines in a pairwise fashion for differential gene expression and to analyze the expression patterns of selected candidate genes in all EFT cell lines individually before testing EFT primary tissues. We further assumed that p53 tumor suppressor gene mutations and INK4A deletions, which have been much more frequently encountered in EFT cell lines than in primary tumors (Kovar et al, 1997), would not influence our study because they were present in the majority of cell lines included in both pools compared. However, we cannot exclude the possibility that the expression of certain genes regulated by EWS-FLI1 is also affected by alterations in p53 and INK4A tumor suppressor genes, which have been proposed to be associated with bad prognoses in a subset of EFT patients (de Alava et al, 2000; Kovar et al, 1997; Wei et al, 2000).
Some modifications of both the cDNA-RDA and LCS procedures were applied because the selectivity and sensitivity of the two strategies applied is greatly influenced by the ratio of tester to driver cDNA pools and by the number of consecutive subtractions (Welford et al, 1998). Although the validity of the RDA and LCS methods in detecting expression differences with high sensitivity could be confirmed, the majority of the clones derived from our screen showed interindividual variation rather than EWS-FLI1 fusion type specificity. Expression differences were found to be in the range of up to two-fold, as indicated by phosphor-imaging of hybridization signals on Northern blots and by RTQ-PCR. Based on the method’s sensitivity, significant differences in gene expression patterns segregating with specific EWS-FLI1 fusion type should have been selected. However, if only a small set of genes exist that are differentially affected by the different EWS-FLI1 fusion proteins, we may have missed the relevant candidates by analyzing only about 800 clones on dot blots. The 112 genes that were used for individual analyses on Northern blots were selected because they displayed the highest discrepancies in signal intensities on dot blots. In fact, when tested on individual cell line RNAs, even these clones showed only two-fold variations in gene expression. Their overrepresentation in one, as opposed to the other, cDNA pool was most likely caused by increased expression levels in single cell lines, as has been observed for the Fibronectin gene in the A673 type 1 cell line. Thus, among the 800 clones analyzed, it is unlikely that we have missed clones that are significantly more highly expressed in cell lines of one as compared to those of the other EWS-FLI1 fusion type with consistency.
The results presented here serve to illustrate the wide array of genes expressed to different degrees in EFT. Interestingly, genes identified thus far are from several functional categories, including cytoskeletal components, chromatin remodeling molecules, nuclear proteins (transcription factors and DNA processing enzymes), ion channels, protein processing, protein transport and folding molecules, extracellular proteins, signaling molecules, and growth and differentiation regulators.
The elucidation of the role of each of these candidate genes in the EFT cell phenotype is beyond the scope of this series of experiments. Several genes identified in our study have been reported to be differentially expressed in cancers and to affect cell proliferation and tumor progression. For example, the expression of elongation factor 1-alpha (Edmonds et al, 1996), translationally controlled tumor protein (Baudet et al, 1998), and the v-fos transformation effector protein (Lecomte et al, 1997) have been reported to be altered in several tumors. It has been recently reported that cleavage of p35, the neuron-specific activator of cyclin-dependent kinase 5 (cdk5), to p25 by calpain (one of the isolated clones) may be involved in the pathogenesis of Alzheimer’s disease (Lee et al, 2000). It is therefore possible that some of these genes contribute to the biology of individual EFT independently of EWS-FLI1 fusion type. It cannot be established at this stage whether composite expression patterns of these genes may differ in EFT of different prognostic subgroups. The different clones obtained for each fusion type from both cDNA-RDA and LCS could be used on cDNA microarrays for hybridization to a large number of individual EFT to identify such composite expression patterns. It may also be possible that there is no difference in the transcriptional activity of the various EWS-FLI1 chimeric transcription factor types on endogenous gene expression in EFT and that the transactivation potentials of type 1 and type 2 fusion types, so far identified in reporter gene assays only, do not contribute to different courses of disease. Instead, there may be a transactivation independent function for EWS-FLI1, as suggested recently (Jaishankar et al, 1999), that may be influenced by different fusion type structures. Alternatively, it cannot be excluded that variability in EWS-FLI1 architecture does not affect EFT biology at all, because the clinical results previously obtained by Zoubek et al (1996) and others (de Alava et al, 1998) could not be confirmed so far for a cohort of patients treated in France (Olivier Delattre, personal communication). Thus, the impact of EWS-FLI1 fusion type on prognosis remains disputable and requires prospective evaluation on greater numbers of patients.
Based on the findings presented in this study, it could be deduced that if we had chosen at random a cell line for each fusion type for both the RDA and LCS analyses, the probability of obtaining clones that will clearly show significant differences in their expression level between the two materials would have been very high. However, our results suggest that the majority of these differences would have presumably resulted from intraindividual, rather than from EWS fusion type-associated, variation in gene expression; that is, they would have reflected only the individual situation of the cell line. On this basis, we have abstained from analyzing some of the candidate clones by RTQ-PCR, using only the primary tumor samples at our disposal. Our data, therefore, call for caution in interpreting the results from comparisons of single tumor specimens as revealing general biological differences in tumor behavior.
Materials and Methods
Cell Lines and Fusion Types
EFT cell lines were routinely maintained in RPMI-1640 (GIBCO BRL, Gaithersburg, Maryland) supplemented with 10% fetal bovine serum, penicillin (0.1 mg/ml), and streptomycin (0.1 mg/ml) in 5% CO2 at 37° C. All cell lines used in this study expressed EWS-FLI1, as determined by reverse transcriptase polymerase chain reaction (RT-PCR) (Zoubek et al, 1996). Cell lines SK-N-MC, a generous gift from Dr. J. Biedler (Memorial Sloan-Kettering Cancer Center, New York), WE-68 and WE-M2-68, kindly provided by Dr. F. van Valen (Department of Pediatrics, University of Münster, Münster, Germany), and TC-252, a gift of Dr. T. Triche (Department of Pathology, Children’s Hospital, Los Angeles, California), as well as A673, from the American Type Culture Collection (ATCC, Rockville, Maryland), expressed type 1 EWS-FLI1. Cell lines IARC-EW2, kindly supplied by Dr. G. M. Lenoir (International Agency for Research on Cancer, Lyon, France), VH64, provided by Dr. F. van Valen (Department of Pediatrics, University of Münster), RDES and SK-ES1 (ATCC) carried type 2 EWS-FLI1 fusions. Among the cell lines established in our institute, STA-ET-1, STA-ET-3, and STA-ET-9 expressed type 1, and STA-ET-6, STA-ET-7, and STA-ET-8 expressed type 2 EWS-FLI1 chimeric products.
RNA Isolation and cDNA Synthesis
Poly(A)+ RNA was extracted from the total RNA of a pool of seven different RNAs for each fusion type through oligo(dT)25-Dynabeads (Dynal, Hamburg, Germany) according to the manufacturer’s recommendations. cDNA was synthesized from 1 μg of poly(A)+ RNA employing the SMART cDNA synthesis technology (Clontech, Palo Alto, California). Briefly, a modified oligo dT primer (cDNA synthesis [CDS] primer, with an RsaI site to facilitate removal of these identical sequences from the PCR-amplified cDNA molecules) primes the first strand cDNA synthesis reaction. When reverse transcriptase (RT) reaches the 5′ end of the mRNA, the enzyme’s terminal transferase activity adds a few additional nucleotides, primarily deoxycytidine, to the 3′ end of the cDNA. A SMARTTM oligonucleotide, which has an oligo(G) sequence at its 3′ end, base-pairs with the deoxycytidine stretch, creating an extended template. RT then switches templates and continues replicating to the end of the oligonucleotide. The resulting full-length, single-stranded cDNA contains the complete 5′ end of the mRNA, as well as sequences that are complementary to the SMART oligonucleotide. Second-strand cDNA synthesis was accomplished by long distance PCR amplification of the first-strand cDNA using a specific SMART PCR primer according to instructions from the manufacturer. The SMART oligonucleotide, CDS primer, and PCR primer all contain a stretch of identical sequence. The final cDNA product was extracted with phenol-chloroform and then ethanol-precipitated and resuspended in 20 μl of water. Tester and driver cDNA samples were prepared in parallel at all times.
cDNA-RDA
Representational difference analysis was performed as described (Braun et al, 1995) with some modifications. Tester and driver cDNAs were digested with DpnII (New England Biolabs, Beverly, Massachusetts), and for adaptor ligation or PCR, oligonucleotides used were as described (Hubank and Schatz, 1994). For the first cycle of subtractive hybridization, tester:driver ratios of 1:2 and 1:100 were simultaneously performed (for both type1 − type2 and type2 − type1). After hybridization for 20 hours at 67° C, the ends of the resulting hybrids were filled in, and a PCR amplification was done. The second and the third cycles of subtraction were performed similarly with ratios of 1:50 and 1:800 as well as 1:100 and 1:8000, respectively. Amplicons from the second or third rounds of subtractive hybridization were gel-purified by means of the QIAquick PCR purification kit (QIAGEN, Hilden, Germany), inserted into the pGEM-T Easy vector (Promega) at the EcoRV site, and transformed into Escherichia coli JM 109 cells.
Linker Capture Subtraction
Restriction enzyme digestion, linker ligation, and PCR amplification were done according to the method of Yang and Sytkowsky (1996) with minor modifications. Briefly, the double-stranded cDNA was digested with AluI and RsaI and then ligated to linkers that had a blunt end and a two-base 3′ protruding end, ACTCTTGCTTGGACGAGCTCT and ACTGAGAACGAACCTG CTCGAGA-p. The linker contained an AluI/SacI site near the blunt end. The top strand was designated the AP. The linker was prepared by combining an equal mass of each of the two oligodeoxynucleotides, heating the mixture to 90° C for 2 minutes, and allowing it to cool slowly to room temperature to enable annealing of the two strands. Ligation was carried out by mixing 1 μg of cut cDNA, 5 μg of linker, 1× ligation buffer (Stratagene, La Jolla, California) and 4 Weiss units of T4 DNA-ligase (Stratagene) in a volume of 10 μl, for 20 hours at 8° C. The reaction mixture was electrophoresed through a 2% low-melt agarose gel to remove unligated linkers, and linker-ligated cDNA fragments in the size range of 0.1–1.0 kb were collected. Linker-ligated cDNA fragments in agarose were amplified directly by PCR using the top strand sequence of the linker as the primer. The reaction (100 μl) contained 10 mm Tris-HCl, pH 8.9, 50 mm KCl, 0.1% Triton X-100, 200 μm dNTPs, 1μm AP primer, 2 mm MgCl2, 1 μl of molten agarose, and 5 U Taq polymerase (Promega), running for 30 cycles (94° C, 1 minute; 55° C, 1 minute; 72° C, 1 minute). The amplified cDNA fragments were purified using a QIAEX purification kit (QIAGEN) and were used as the initial material for subtractive hybridization. After two rounds of subtraction, the PCR-amplified products were gel-purified, inserted into pGEM-T Easy vector (Promega), and transformed into Escherichia coli JM109 cells.
Dot Blot Hybridization and PCR and Sequence Analysis
Forty-eight white colonies from each subtracted cell type were picked at random and inoculated into LB + Ampicillin medium in individual wells of a 96-well plate. Two replica blots were prepared on HybondTM-N nylon filters using 25 μl of bacterial cells per well. The replica dot-blots were processed according to the method of Brown and Knudson (1991) and probed with random-primed–labeled driver DNAs from each fusion type.
Inserts were analyzed by direct PCR amplification of a 1 μl aliquot of each colony inoculum employing vector-specific primers (T7 and Sp6) for 30 cycles of 95° C, 1 minute; 46° C, 1 minute; and 72° C, 1 minute. The PCR products were electrophoresed on 2% agarose gels. The desired bands were excised and purified using the QIAquick PCR purification kit (QIAGEN). The products were then subjected to direct DNA sequencing and were employed to prepare probes for Northern blot analyses.
Northern Blotting and Phosphor-Imager Analysis
Ten μg of total RNA was fractionated on a 1.2% agarose/37% formaldehyde gel and transferred overnight onto a Hybond N membrane (Amersham, Aylesbury, United Kingdom) according to standard protocols. Prehybridization and hybridization reactions were performed at 42° C in 50% (v/v) formamide, 5 × SSC, 50 mm Tris-HCl, pH 7.5, 5 × Denhardt’s solution, 5% SDS, and 250 μg/ml denatured salmon sperm DNA, and washed at a final stringency of 0.1 × SSC, 0.1% SDS at 55° C. Blots were hybridized to 32P-labeled cDNA probes and hybridization was detected by autoradiography. Quality and comparable loading of RNA samples were confirmed by including ethidium bromide in the gels and by rehybridizing blots to an 18S rDNA probe. After autoradiography, the filters were then exposed overnight to a Packard screen and scanned at 50 μm resolution in a phosphorimager instrument for quantification (Cyclone Instrument; Packard, Meriden, Connecticut).
RTQ-PCR Analysis of Clones
RTQ-PCR is based on the continuous optical monitoring of the progress of a fluorogenic PCR (Heid et al, 1996; Lo et al, 1999). The specificity of amplicon sequence selection was determined using primer and probe sequences that specifically detect the experimental gene sequence, as determined by means of the ABS module (Applied Biosysytems, Foster City, California). Analysis of gene expression was generated using an ABI Prism 7700 Sequence Detection System (TaqMan; Applied Biosystems, Foster City, California), which uses the 5′ nuclease activity of Taq DNA polymerase to generate a real-time quantitative DNA analysis assay (Gelmini et al, 1997; Heid et al, 1996). A nonextendable oligonucleotide hybridization probe with 5′ fluorescent and 3′ rhodamine (quench) moieties is present during the extension phase of the PCR. Degradation and release of the fluorescent moiety attributable to the 5′ nuclease activity results in peak emission at 518 nm and is monitored every 8.5 seconds by a sequence detector. The increase in fluorescence is monitored during the complete amplification process (real-time). A relative standard curve representing four-fold dilutions of stock cDNA of the gene of interest (1:2.5, 1:10, 1:40, and 1:160) was used for linear regression analysis of unknown samples. The expression of the housekeeping gene, GAPDH, was used to normalize for variances in input cDNA. The sequences of the PCR primer pairs and fluorogenic probe (5′ to 3′), respectively, that were used for each gene are as follows:
1S-3(VDAC): (forward) ccttggttgtgatgttgactttga, (reverse) gccctcataaccaaagacagct, and FAM-tttgctgg acctgcaatccatggtt-TAMRA
2-20(TAXREB107): (forward) ttacccccgggaccattct, (reverse) cttcaggaaaaccaccctcttg, and FAM-atcatcct cactggacgccacaggg-TAMRA
CAPN: (forward) cccacgcacacattactcca, (reverse) caaagagtctccggaactgcc, and TET-cattgaggccaacgag agtgaggagg-TAMRA
UBE1: (forward) gcaccacgtcaccagtactataacc, (reverse) aactgtttgagggtcatctcctca, and TET-agagtgga cattgtgggatcgctttga-TAMRA.
The fluorogenic probes are FAM, TET, and TAMRA.
Statistical Analyses
Duration of relapse-free survival (RFS) for patients with localized disease was computed from the date of diagnosis of EFT to the first occurrence of a disease event, defined as local or systemic relapse of EFT. Distributions of RFS were estimated according to the method of Kaplan and Meier, and comparisons of RFS were performed by means of log-rank tests. The Kruskal-Wallis analyses of variance was applied to test for differences in gene expression levels as monitored by phosphorimaging of Northern blots or by RTQ-PCR.
Accession codes
Accessions
GenBank/EMBL/DDBJ
References
Baudet C, Perret E, Delpech B, Kaghad M, Brachet P, Wion D, and Caput D (1998). Differentially expressed genes in C6.9 glioma cells during vitamin D-induced cell death program. Cell Death Differ 5: 116–125.
Braun BS, Frieden R, Lessnick SL, May WA, and Denny CT (1995). Identification of target genes for the Ewing’s sarcoma EWS/FLI fusion protein by representational difference analysis. Mol Cell Biol 15: 4623–4630.
Brown SE and Knudson DL (1991). 96-well plates for recombinant library maintenance and dot-blot production. Biotechniques 10: 718–722.
Byrne JA, Tomasetto C, Garnier JM, Rouyer N, Mattei MG, Bellocq JP, Rio MC, and Basset P (1995). A screening method to identify genes commonly overexpressed in carcinomas and the identification of a novel complementary DNA sequence. Cancer Res 55: 2896–2903.
Chang DD, Park NH, Denny CT, Nelson SF, and Pe M (1998). Characterization of transformation related genes in oral cancer cells. Oncogene 16: 1921–1930.
de Alava E, Antonescu CR, Panizo A, Leung D, Meyers PA, Huvos AG, Pardo-Mindan FJ, Healey JH, and Ladanyi M (2000). Prognostic impact of P53 status in Ewing sarcoma. Cancer 89: 783–792.
de Alava E, Kawai A, Healey JH, Fligman I, Meyers PA, Huvos AG, Gerald WL, Jhanwar SC, Argani P, Antonescu CR, Pardo-Mindan FJ, Ginsberg J, Womer R, Lawlor ER, Wunder J, Andrulis I, Sorensen PH, Barr FG, and Ladanyi M (1998). EWS-FLI1 fusion transcript structure is an independent determinant of prognosis in Ewing’s sarcoma. J Clin Oncol 16: 1248–1255.
Edmonds BT, Wyckoff J, Yeung YG, Wang Y, Stanley ER, Jones J, Segall J, and Condeelis J (1996). Elongation factor-1 alpha is an overexpressed actin binding protein in metastatic rat mammary adenocarcinoma. J Cell Sci 109: 2705–2714.
Gelmini S, Orlando C, Sestini R, Vona G, Pinzani P, Ruocco L, and Pazzagli M (1997) Quantitative polymerase chain reaction-based homogeneous assay with fluorogenic probes to measure c-erbB-2 oncogene amplification. Clin Chem 43: 752–758.
Geng M, Wallrapp C, Muller-Pillasch F, Frohme M, Hoheisel JD, and Gress TM (1998). Isolation of differentially expressed genes by combining representational difference analysis (RDA) and cDNA library arrays. Biotechniques 25: 434–438.
Hahm KB, Cho K, Lee C, Im YH, Chang J, Choi SG, Sorensen PH, Thiele CJ, and Kim SJ (1999). Repression of the gene encoding the TGF-beta type II receptor is a major target of the EWS-FLI1 oncoprotein. Nat Genet 23: 481.
Heid CA, Stevens J, Livak KJ, and Williams PM (1996). Real time quantitative PCR. Genome Res 6: 986–994.
Hubank M and Schatz DG (1994). Identifying differences in mRNA expression by representational difference analysis of cDNA. Nucleic Acids Res 22: 5640–5648.
Jaishankar S, Zhang J, Roussel MF, and Baker SJ (1999). Transforming activity of EWS/FLI is not strictly dependent upon DNA-binding activity. Oncogene 18: 5592–5597.
Kovar H (1998). Progress in the molecular biology of Ewing tumors. Sarcoma 2: 3–17.
Kovar H, Jug G, Aryee DN, Zoubek A, Ambros P, Gruber B, Windhager R, and Gadner H (1997). Among genes involved in the RB dependent cell cycle regulatory cascade the p16 tumor suppressor gene is frequently lost in the Ewing family of tumors. Oncogene 15: 2225–2232.
Lecomte F, Szpirer J, and Szpirer C (1997). The S3a ribosomal protein gene is identical to the Fte-1 (v-fos transformation effector) gene and the TNF-alpha-induced TU-11 gene, and its transcript level is altered in transformed and tumor cells. Gene 186: 271–277.
Lee MS, Kwon YT, Li M, Peng J, Friedlander RM, and Tsai LH (2000). Neurotoxicity induces cleavage of p35 to p25 by calpain. Nature 405: 360–364.
Lessnick SL, Braun BS, Denny CT, and May WA (1995). Multiple domains mediate transformation by the Ewing’s sarcoma EWS/FLI-1 fusion gene. Oncogene 10: 423–431.
Liang P and Pardee AB (1992). Differential display of eukaryotic messenger RNA by means of the polymerase chain reaction. Science 257: 967–971.
Lin PP, Brody RI, Hamelin AC, Bradner JE, Healey JH, and Ladanyi M (1999). Differential transactivation by alternative EWS-FLI1 fusion proteins correlates with clinical heterogeneity in Ewing’s sarcoma. Cancer Res 59: 1428–1432.
Lisitsyn N and Wigler M (1993). Cloning the differences between two complex genomes. Science 259: 946–951.
Lisitsyn NA, Lisitsina NM, Dalbagni G, Barker P, Sanchez CA, Gnarra J, Linehan WM, Reid BJ, and Wigler MH (1995). Comparative genomic analysis of tumors: Detection of DNA losses and amplification. Proc Natl Acad Sci USA 92: 151–155.
Lo YM, Chan LY, Lo KW, Leung SF, Zhang J, Chan AT, Lee JC, Hjelm NM, Johnson PJ, and Huang DP (1999). Quantitative analysis of cell-free Epstein-Barr virus DNA in plasma of patients with nasopharyngeal carcinoma. Cancer Res 59: 1188–1191.
May WA, Arvand A, Thompson AD, Braun BS, Wright M, and Denny CT (1997). EWS/FLI1-induced manic fringe renders NIH 3T3 cells tumorigenic. Nat Genet 17: 495–497.
May WA, Gishizky ML, Lessnick SL, Lunsford LB, Lewis BC, Delattre O, Zucman J, Thomas G, and Denny CT (1993). Ewing sarcoma 11;22 translocation produces a chimeric transcription factor that requires the DNA-binding domain encoded by FLI1 for transformation. Proc Natl Acad Sci USA 90: 5752–5756.
O’Neill MJ and Sinclair AH (1997). Isolation of rare transcripts by representational difference analysis. Nucleic Acids Res 25: 2681–2682.
Thompson AD, Braun BS, Arvand A, Stewart SD, May WA, Chen E, Korenberg J, and Denny C (1996). EAT-2 is a novel SH2 domain containing protein that is up regulated by Ewing’s sarcoma EWS/FLI1 fusion gene. Oncogene 13: 2649–2658.
Wang Z and Brown DD (1991). A gene expression screen. Proc Natl Acad Sci USA 88: 11505–11509.
Wei G, Antonescu CR, de Alava E, Leung D, Huvos AG, Meyers PA, Healey JH, and Ladanyi M (2000). Prognostic impact of INK4A deletion in Ewing sarcoma. Cancer 89: 793–799.
Welford SM, Gregg J, Chen E, Garrison D, Sorensen PH, Denny CT, and Nelson SF (1998). Detection of differentially expressed genes in primary tumor tissues using representational differences analysis coupled to microarray hybridization. Nucleic Acids Res 26: 3059–3065.
Yang M and Sytkowski AJ (1996). Cloning differentially expressed genes by linker capture subtraction. Anal Biochem 237: 109–114.
Zeng J, Gorski RA, and Hamer D (1994). Differential cDNA cloning by enzymatic degrading subtraction (EDS). Nucleic Acids Res 22: 4381–4385.
Zoubek A, Dockhorn-Dworniczak B, Delattre O, Christiansen H, Niggli F, Gatterer-Menz I, Smith TL, Jurgens H, Gadner H, and Kovar H (1996). Does expression of different EWS chimeric transcripts define clinically distinct risk groups of Ewing tumor patients? J Clin Oncol 14: 1245–1251.
Acknowledgements
We are grateful to Dr. Olivier Delattre for providing us with unpublished data and Ms. Ulrike Poetschger, Dr. Michael Paulussen, and Dr. Susanne Ahrens for the statistical analyses and Mr. Herwig Machat for excellent technical assistance.
This work was supported in part by grants 6137 and P14299GEN of the “Jubiläumsfonds der Oesterreichischen Nationalbank” and the Austrian Science Fund, respectively, and by private donations to the Children’s Cancer Research Institute.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Aryee, D., Sommergruber, W., Muehlbacher, K. et al. Variability in Gene Expression Patterns of Ewing Tumor Cell Lines Differing in EWS-FLI1 Fusion Type. Lab Invest 80, 1833–1844 (2000). https://doi.org/10.1038/labinvest.3780194
Received:
Published:
Issue Date:
DOI: https://doi.org/10.1038/labinvest.3780194
This article is cited by
-
Diagnostic and Prognostic Sarcoma Signatures
Molecular Diagnosis & Therapy (2008)
-
Differentiating Ewing's sarcoma from other round blue cell tumors using a RT-PCR translocation panel on formalin-fixed paraffin-embedded tissues
Modern Pathology (2007)
-
EWS/FLI-1 oncoprotein subtypes impose different requirements for transformation and metastatic activity in a murine model
Journal of Molecular Medicine (2007)
-
Rapamycin induces the fusion-type independent downregulation of the EWS/FLI-1 proteins and inhibits Ewing's sarcoma cell proliferation
Oncogene (2003)