Abstract
Inactivation of p16 by methylation of CpG islands is a frequent early event in gastric carcinogenesis. The positive relationship between p16 methylation and the clinical characteristics of gastric carcinomas (GC) has not been reported to date. In the present study, a DHPLC assay to quantify p16 methylation was established (detection limit by fluorescence detector: 1:255 (Methlyated vs Unmethylated)). The proportion of methylated p16 in the representative samples was confirmed and standardized by clone sequencing. Then, the DHPLC and two regular methylation-specific PCR (MSP) assays were used to detect p16 methylation in 82 paired, resected GCs and their adjacent normal tissues. Results showed that the average proportion of methylated p16 in GCs was significantly higher than that in their adjacent samples (12.90 vs 0.63%; t-test P=0.005). A much higher proportion of methylated p16 was detected in GCs with metastases (local or distant) than without metastases (14.76 vs 2.61%; t-test P=0.014). A proportional relationship was observed between clinical stages and positive rates of p16 methylation in GCs and/or adjacent tissues: 27.3, 37.5, and 58.8% (by DHPLC) for stage-I, -II, -III–IV of GCs, respectively (two-sided Fisher's exact test P=0.016). To confirm the data obtained by DHPLC, two MSP primer sets (p16-M and p16-M2) were also used to analyze p16 methylation in the same set of samples simultaneously. Data of MSP assay using the primer set p16-M2, but not p16-M, correlated with that of DHPLC. These results imply that the primer set p16-M2 might be more suitable than p16-M to detect p16 methylation in gastric tissues. In conclusion, the present data indicates that p16 methylation correlates with progression of GCs significantly.
Similar content being viewed by others
Main
Methylation of CpG islands is one of the crucial epigenetic pathways used to regulate gene transcription, which is involved in cell differentiation, parasite DNA defending, X-chromosome inactivation, and gene imprinting.1, 2 Aberrant methylation of CpG islands is tumor-specific, which may result in inactivation of tumor suppressor genes.3, 4, 5, 6
P16INK4A (CDKN2A/MTS1) is a cell cycle regulator involved in the inhibition of G1 phase progression.7 Methylation of p16 CpG islands silences transcription of this gene.8 It was reported that the p16-methylated cells could have an advantage in progression and metastasis in non-small-cell lung cancers.9 Aberrant p16 methylation was reported to occur frequently in multiple human cancers and relate to TNM stages of non-small-cell lung cancer and esophageal adenocarcinoma.8, 9, 10, 11, 12, 13
In primary gastric carcinoma (GC), the frequency of p16 inactivation by homozygous deletions ranged from 0–9%, by mutation from 0–2%, whereas by methylation from 32–42%,14, 15, 16, 17, 18, 19, 20 which suggests that methylation is a major mechanism for p16 inactivation in GC. It was reported that methylated p16 was also observed in pre-malignant stages of GC.21 We previously reported a positive association between aberrant p16 methylation and the severity of glandular stomach pathology of Wistar rats induced by chemical carcinogen.22 Recently, we observed that p16 methylation was also associated with malignant transformation of human low-grade gastric dysplasia in a nested case–control study based on a long-term population follow-up screening.23 These studies indicated that p16 methylation might play an important role in gastric carcinogenesis. However, relationship between p16 methylation and clinical characteristics of GCs was not observed.24, 25 Gastric carcinomas is still the leading cause of cancer death in China and the second leading cause in the world.26 It is necessary to clarify whether clinical characteristics of GC are associated with the proportion of methylated p16.
Denatured high-performance liquid chromatography (DHPLC) was successfully used to detect methylation of various CpG islands.27, 28, 29, 30 In the present study, a quantitative DHPLC assay was developed to detect p16 methylation in 82 paired GCs and adjacent tissue samples. We found that the proportion of methylated p16 in GCs was significantly higher than that in their adjacent nonmalignant tissues, and that p16 methylation correlated with the observed clinical stages of GCs.
Materials and methods
Gastric Carcinoma Samples and Cell Lines
Eight-two paired surgical primary gastric carcinoma and adjacent mucosa samples (fresh-frozen at −70°C) were collected from patients at Beijing Cancer Hospital (63 males and 19 females, 35–81 years old, the average age 58 years). All clinical samples and histopathological information for each case were obtained according to approved institutional guidelines. The 1997 UICC-TNM criteria was used for classification of GCs. Human colon cancer cell line RKO and gastric cancer cell line MGC803 were cultured in RPMI 1640 medium (Gibco) supplemented with 10% of fetal bovine serum at 37°C with 5% CO2.
DNA Preparation and Bisulfite Modification
Genomic DNA (2 μg) of cell lines and tissue samples was isolated with phenol/chloroform extraction as previously described.31 The unmethylated-cytosines of the genomic DNA were converted to uridines by the addition of 5 M sodium bisulfite.32 The Wizard® DNA Clean-Up System Kit (Promega) was used before PCR amplification.
Design of Primers and PCR Conditions
The universal primers used to amplify the antisense strand of methylated and unmethylated p16 CpG island (GenBank accession number 527803) after bisulfite modification are 5′-TTTTT AGAGG ATTTG AGGGA TAGG-3′ (sense) and 5′-CTACC TAATT CCAAT TCCCC TACAA ACTTC-3′ (antisense) as previously described.33 35 CpG sites are located in the 392 bp amplicon (Figure 2d). The PCR products of p16 CpG islands were amplified by hot-start PCR. HotStarTaq DNA polymerase (QIAGEN GmbH, Hilden, Germany) was used. A touchdown PCR protocol was used for amplification of p16: 95°C for 15 min → (95°C for 40 s → 70°C for 60 s, −1°C/cycle → 72°C for 60 s) × 10 cycles → (95°C for 40 s → 60°C for 60 s → 72°C for 60 s) × 30 cycles → 72°C for 10 min.
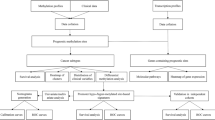
Validation of the proportion of methylated p16 by clone sequencing. (a) Comparison of the exact proportion of methylated p16 by clone sequencing with the non-adjusted proportion of methylated p16 by DHPLC in PCR products from the bisulfite modified genomic DNA of four representative gastric carcinoma samples. (b) Results of clone sequencing of PCR products of four representative gastric carcinoma samples; each dot represents one CpG site; ▪, methylated cytosines; □, unmethylated cytosines; each row of dots represents one clone; each clone block represents one sample; each sample's name is listed on the top of its corresponding clone block; 35 CpG sites in the fragment are highlighted; the underlined, and italic nucleotides are the primer set sequences for p16-M MSP, and p16-M2 MSP assay, respectively. (c) The linearity of the exact proportions and the non-adjusted proportions of methylated p16 presented in (a). The K-value (1.4996) for methylated p16 is calculated as described in method section. (d) The bisulfite modification sequence of the fragment of the methylated p16 CpG island used for DHPLC analysis and clone sequencing; 35 CpG sites in the fragment are highlighted as described in (b) section; the bold sequences are the primer set for DHPLC.
The genomic DNA samples of cell line RKO with methylated p16 and MGC803 with unmethylated p16 (determined by sequencing) were used as positive and negative controls. These DNA samples were also used to optimize the partial denaturing temperature needed for separation of the methylated and unmethylated p16 amplicons for analysis with DHPLC and to calculate the standardization constant K for quantitative analysis of CpG methylation by DHPLC, as described in the next section.
To determine the detection limit for the methylated amplicons in the presence of the unmethylated amplicons for p16, different ratios of the p16-methylated genomic DNA from RKO cells vs the p16-unmethylated genomic DNA from MGC803 cells were prepared (1:1, 1:3, 1:7, 1:15, 1:31, 1:63, 1:127, 1:255), injected onto the WAVE System (Transgenomic, Inc., Omaha, USA), and analyzed at 57.6°C using both UV and fluorescence detection.
Amplification of the Methylated p16 CpG Islands
The methylation status of the bisulfite-modified p16 CpG islands was analyzed with methylation-specific polymerase chain reactions (MSP) as previously described.34 Briefly, both the p16-M, p16-U, and p16-M2 primer sets for the methylated p16 CpG islands were 5′-TTATT AGAGG GTGGG GCGGA T CGC-3′ (sense one of p16-M and p16-M2), 5′-GACCC CG AAC CGCGA CCGTA A-3′ (antisense one of p16-M), 5′-CCACC TAAAT CG ACC TCCGA CCG-3′ (antisense one of p16-M2), 5′-TTATTAGAGGGTGGGGTGGAT TGT-3′ (sense one of p16-U), and 5′-CAACCCCA AACCACAACCATAA-3′ (antisense one of p16-U). Hot-start MSP was used for amplification of the methylated p16 CpG islands. Thermal cycler conditions were denaturing at 95°C for 15 min, amplification for 35 cycles (95°C for 40 s, 64°C for 40 s, 72°C for 40 s), and extension at 72°C for 10 min. The reaction mixture (20 μl) contained about 10 ng of templates, 4 pmol of each primer, 4 nmol of dNTP, 1 unit of HotStarTaq DNA polymerase (QIAGEN GmbH, Hilden, Germany), and 2 μl of 10 × GC-buffer. Distilled water and genomic DNA of the p16 unmethylated MGC803 cells were used as template for negative control. Genomic DNA of p16-methylated RKO was used as positive control. These controls were used in every individual MSP experiment. The methylated p16 MSP products could be amplified with either the p16-M or p16-M2 primer set only in the positive control, but not in the negative controls. The MSP products were run on 8% PAGE gel.
Quantification Analysis for Methylation by DHPLC
The PCR products amplified by universal primers were analyzed by DHPLC on the WAVE™ DNA Fragment Analysis System coupled to the post-column HSX-3500 Accessory (Transgenomic, Inc., Omaha, USA) and the high-sensitivity fluorescence (FL) detector (excitation at 450 nm, emission at 520 nm) as previously described.35 The PCR products were separated by a DNASep® analytical column (Transgenomic, Inc.) at 57.6°C, the partially denaturing temperature of the amplicon of the unmethylated p16 as predicted by Transgenomic WAVEMaker software. The WAVE-HS1 FL-dye buffer (Transgenomic, Inc.) was used to enhance the FL-intensity of the PCR products (universal post-column labeling). Classical 3 × signal/noise (3S/N) criteria was used to determine the detection limit. Because the peaks for both the methylated and unmethylated p16 CpG islands can be separated in the same DHPLC injection under partially denaturing conditions (57.6°C), the proportion of the methylated p16 CpG island in the sample was calculated based upon the peak height of methylated p16 divided by the sum of the peak heights of the methylated and unmethylated p16 peaks from the DHPLC chromatogram. The proportion of methylated p16 was calculated by the following formula:

where K is the standardization constant which is determined by the ratio of the exact proportion of the methylated CpG islands detected by clone sequencing to the non-adjusted proportion of methylated CpG islands. For p16, the K-value is 1.4996.
Clone Sequencing
Fresh PCR products of p16 for each sample were amplified with the same universal primers, cloned with the AT clone kit (Tianwei Time Company, Beijing, China), and sequenced on ABI PRISM 3730 DNA Analyzer. About 15–20 clones with information of the target sequence were obtained for each sample. Two kinds of p16 MSP products for the representative samples were also confirmed by direct sequencing as described previously.23
Results and discussions
The Proportion of Methylated p16 CpG Islands can be Detected by DHPLC Quantitatively
We first developed a DHPLC assay to detect methylation of hMLH1 CpG islands and subsequently used it to quantify the proportion of methylated MT-3 in tissue samples previously.27, 28, 29 DHPLC was also used to analyze p16 methylation.33 However, quantification of the proportion of methylated p16 in tissue samples was not reported. To quantify the proportion of methylated p16 in tissue samples with DHPLC, we performed a series of experiments. After bisulfite modification, a pair of universal primers was used to amplify both methylated and unmethylated p16 in a same PCR reaction. Amplicons for methylated and unmethylated p16 were separated at 57.6°C within a single DHPLC injection (Figure 1). The retention times (tR) for the methylated and unmethylated p16 peaks were 4.9 and 5.3 min, respectively, as determined by individual injections of the methylated and unmethylated products. A linear relationship (y=0.95x, R2=9.99; for fluorescence (FL)-Detector) was observed between peak heights of methylated p16 vs unmethylated p16 PCR products at various dilution ratios (1:1, 1:3, 1:7, 1:15, 1:31, 1:63, 1:127, 1:255). Such linearity was also obtained among the PCR products amplified from the p16 methylated genomic DNA of RKO cells diluted by the p16 unmethylated genomic DNA of MGC803 cells (data not shown). Methylated p16 was still detected by FL-Detector when it was diluted with unmethylated p16 at ratio 1/256 (0.4%) (Figure 1a); by comparison, the detection limit for the UV-Detector was 1/32 (3.2%) (data not shown). Similar results were observed in two independent repeat experiments (data not shown). These results indicate that DHPLC with FL-Detector may be a very sensitive quantitative assay for the detection of p16 methylation. As the proportion of methylated p16 was directly calculated based on the peak heights of methylated and unmethylated p16 in a single DHPLC injection, a DHPLC assay has an inherent advantage over other quantitative assays such as MSP (MethyLight), in which a separate reference PCR and a labeled probe are generally required to standardize the amount of template used for each sample.36
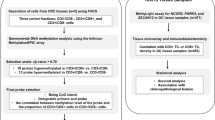
Analysis of methylation of CpG island of p16 by DHPLC quantitatively. (a) Detection of the methylated p16 amplicons diluted with the unmethylated amplicons at various ratios by the fluorescence detector at the partial denaturing temperature 57.6°C. (b) DHPLC chromatograms (fluorescence detection) of PCR products of methylated (if any) and unmethylated p16 CpG islands amplified with universal primers from the bisulfite modified genomic DNA of four representative gastric carcinoma samples.
Validation and Standardization of the Proportion of Methylated p16 by Clone Sequencing
To confirm the result of quantification of CpG methylation by DHPLC, the methylation status of p16 CpG islands in four representative samples of GCs were quantified by DHPLC and clone sequencing simultaneously (Figures 1b and 2b). In order to determine the ratio of methylated p16 in the sample in the DHPLC assay, the peak height for methylated p16 (tR 5.3 min) product was divided by the sum of the peak heights for methylated and unmethylated p16 (tR 4.9 min) products. For comparison using clone sequencing, the non-adjusted proportion of methylated p16 and the ratio of the methylated clones to the total number of sequenced clones were used to calculate an exact proportion of methylated p16. The results showed that the non-adjusted proportion of methylated p16 in GCs by DHPLC with FL-Detector correlated with ratio of the methylated clones linearly (R2=0.977, y=1.4996x; Figure 2a and c). In addition, the non-adjusted proportion of methylated p16 by DHPLC was lower than the exact proportion detected by clone sequencing. Therefore, the non-adjusted proportion of methylated p16 was standardized with the slope constant (K) of 1.4996 during calculation of the proportion of each tested sample (as described in the Materials and methods section).
p16 Methylation is Associated with the Clinical Stages of GCs
p16 methylation is an early, frequent event in gastric carcinogenesis.21, 22, 23 To investigate whether p16 methylation correlates with clinical characteristics of GCs, the methylation status of p16 CpG islands in 82 paired GCs and adjacent tissue samples from the operative cutting margin were quantified with DHPLC and analyzed with MSP as described in the Materials and methods section. Methylated p16 was detected in GCs and/or adjacent tissues from 31 of 82 patients (17 GCs and 20 adjacent tissues) by DHPLC. The adjusted average proportion of methylated p16 in GCs was >20-fold higher than that in their adjacent samples (12.90 vs 0.63%; t-test P=0.005). Among six paired samples, in which methylated p16 was detected both in tumors and normals, the proportion of methylated p16 in tumor was always higher than that in normal (the adjusted average proportion, 14.71 vs 0.76%). Gastric carcinogenesis could be a multi-center process. Precancerous lesions are often observed in different sites within a stomach, especially in the cases with gastric carcinomas. Hence, that aberrant p16 methylation might present in these lesions but not in the sampling site of tumor may account for the detection of p16 methylation only in the adjacent normals.
A much higher positive rate and average proportion of methylated p16 were detected in GCs (n=47) with metastases (local or distant) than GCs (n=35) without metastases (Table 1). The positive rate and adjusted average proportion of p16 methylation in GCs were also associated with their TNM-based clinical stages (Table 2). When the normal and tumors were taken together, there was a significant correlation (P=0.0328, linear trend test by Epi Info 6.0). Thus, detection of methylated p16 in adjacent tissues might also imply the presence of invasive tumor cells and poor prognosis. Correlation between p16 methylation and age, sex, differentiation of GCs was not observed. To the best of our knowledge this is the first report, that correlated p16 methylation with the progression of GCs both qualitatively and quantitatively.
It was reported that p16 methylation by MSP was not associated with pathological characteristics of GCs.24, 25 To determine whether different assays result in different results, the p16 methylation status in 82 above DHPLC-analyzed samples was also determined by MSP as previously described.34 The primer set p16-M and p16-U, the most frequently used primers for detection of p16 methylation, was used to amplify the methylated p16 (150-bp) and the unmethylated p16 (151-bp) MSP products. Methylated p16 was detected in total 37 of the 82 tested cases with GC (22 GCs and 25 adjacent tissues) by MSP. Unmethylated p16 was detected in all tested tissue samples. However, the relationship between p16 methylation by MSP and clinical characteristics of GCs was not observed in the present study. Furthermore, the results by MSP did not correlate with those by DHPLC (Figure 3 bottom block); p16 methylation was detected in only 35.3% (12/34) of DHPLC-positive samples by MSP using the primer set p16-M. A potential reason why the correlation was poor between the DHPLC results and the MSP results using the p16-M primer set is that MSP is a CpG-site specific assay that is used to detect methylation of a few CpG sites complementary to the primers. The MSP assay may work well when these complementary CpG sites are fully methylated or unmethylated such as in the case of in vitro cell lines. However, in the case of tissue samples, fully methylation of target CpG sites may be present in only a small number of copies. For example, full methylation of the complementary CpG sites was observed in some of clones within the antisense p16-M primer (Figure 2b, underlined blocks). This implies that the used primer set for 150-bp fragment of methylated p16 might not be suitable for analysis of p16 methylation in some kinds of tissues, such as stomach tissue. This might contribute to the observation that p16 methylation determination by MSP did not correlate with the clinical characteristics of GCs in the present study as well as previous investigations.24, 25
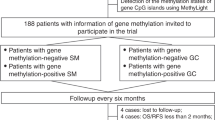
Comparison of detection of methylated p16 in 164 gastric tissue samples with various assays. Blocks within the black frame represent the area of samples containing methylated p16 by DHPLC; blocks within the purple frame represent the area of samples containing methylated p16 by MSP using p16-M2 primer set; blocks within the blue frame represent the area of samples containing methylated p16 by MSP using p16-M primer set; overlapped blocks represent the area of samples in which methylated p16 was detected by two assays simultaneously. The exact positive sample number for each assay was also listed within each block.
Another set of MSP primers (p16-M2, 234-bp amplicon) were used to determine the amount of methylation of p16 CpG islands.34 In the present study, we observed that all CpG sites complementary to the primer set p16-M2 were methylated in all of the methylated clones of three representative GC samples (Figure 2b). To investigate whether p16-M2 is better than p16-M, we used the p16-M2 primer set to detect p16 methylation in the same set of the bisulfite-modified GC and adjacent samples. The result indicated that the positive rate of p16 methylation by p16-M2 was correlated with that determined by DHPLC (P=0.0025). A total of 83.8% (31/37) of DHPLC-positive samples were also p16-M2 MSP positive (Figure 3 Top block). Such a correlation was not observed between determinations by the p16-M and the p16-M2 (Figure 3 middle block). In addition, the positive rate of p16 methylation determined by p16-M2 in GCs was higher than that in their adjacent tissue samples (71.25 vs 54.32%, 2-sided Fisher‘s exact test, P=0.026). A relationship between p16 methylation (by p16-M2) and clinical stages of GCs was observed but not significant (P=0.065, two-sides). These results indicated that the p16-M2 primer set correlates with clone sequencing and DHPLC well, and might be more suitable than the p16-M primer set for detection of p16 methylation in gastric tissues. Whether the p16-M2 primer set is more suitable for other kinds of tissues is unknown.
The present data suggests that DHPLC may be a rapid and convenient quantitative assay for the detection of p16 methylation in human tissue samples and that the progression of GCs is associated with p16 methylation, both qualitatively and quantitatively.
Accession codes
References
Jones PA, Laird PW . Cancer epigenetics comes of age. Nature Genet 1999;21:163–167.
Robertson KD . DNA methylation and human disease. Nat Rev Genet 2005;6:597–610.
Costello JF, Frühwald MC, Smiraglia DJ, et al. Aberrant CpG-island methylation has non-random and tumour-type-specific patterns. Nat Genet 2000;24:132–138.
Esteller M . CpG island hypermethylation and tumor suppressor genes: a booming present, a brighter future. Oncogene 2002;21:5427–5440.
Herman JG, Baylin SB . Mechanisms of disease: gene silencing in cancer in association with promoter hypermethylation. N Engl J Med 2003;349:2042–2054.
Jones PA, Baylin SB . The fundamental role of epigenetic events in cancer. Nat Rev Genet 2002;3:415–428.
Serrano M, Hannon GJ, Beach D . A new regulatory motif in cell-cycle control causing specific inhibition of cyclinD/CDK4. Nature 1993;366:704–707.
Merlo A, Herman JG, Mao L, et al. 5′CpG island methylation is associated with transcriptional silencing of the tumor suppressor p16/CDKN2/MTS1 in human cancer. Nat Med 1995;1:686–692.
Seike M, Gemma A, Hosoya Y, et al. Increase in the frequency of p16INK4 gene inactivation by hypermethylation in lung cancer during the process of metastasis and its relation to the status of p53. Clin Cancer Res 2000;6:4307–4313.
Gonzales-Zulueta M, Bender CM, Yang AS, et al. Methylation of the 5′CpG island of the p16/CDKN2 tumor suppressor gene in normal and transformed human tissues correlates with gene silencing. Cancer Res 1995;55:4531–4535.
Herman JG, Merlo A, Mao L, et al. Inactivation of the CDKN2/p16/MTS1 gene is frequently associated with aberrant DNA methylation in all common human cancers. Cancer Res 1995;55:4525–4530.
Ng CS, Zhang J, Wan S, et al. Tumor p16M is a possible marker of advanced stage in non-small cell lung cancer. J Surg Oncol 2002;79:101–106.
Brock MV, Gou M, Akiyama Y, et al. Prognostic importance of promoter hypermethylation of multiple genes in esophageal adenocarcinoma. Clin Cancer Res 2003;9:2912–2919.
Wu MS, Lin YW, Sheu JC, et al. Intragenic homozygous deletions of MTS1 gene in gastric cancer in Taiwan. Jpn J Cancer Res 1996;87:1052–1055.
Takaoka AS, Kakiuchi H, Itoh F, et al. Infrequent alterations of the p16 (MTS-1) gene in human gastric cancer. Tumour Biol 1997;18:95–103.
Gunther T, Schneider-Stock R, Pross M, et al. Alteration of p16/MTS1-tumor suppressor gene in gastric cancer. Pathol Res Pract 1998;194:809–813.
Lee YY, Kang SH, Seo JY, et al. Alterations of p16INK4A and p15INK4B genes in GC. Cancer 1997;80:1889–1896.
Suzuki H, Itoh F, Toyota M, et al. Distinct methylation pattern and microsatellite instability in sporadic gastric cancer. Int J Cancer 1999;83:309–313.
Toyota M, Ahuja N, Suzuki H, et al. Aberrant methylation in gastric cancer associated with the CpG island methylator phenotype. Cancer Res 1999;59:5438–5442.
Shim YH, Kang GH, Ro JY . Correlation of p16 hypermethylation with p16 protein loss in sporadic gastric carcinomas. Lab Invest 2000;80:689–695.
Kang GH, Shim YH, Jung HY, et al. CpG island methylation in premalignant stages of gastric carcinoma. Cancer Res 2001;61:2847–2851.
Bai H, Gu LK, Zhou J, et al. p16 hypermethylation during gastric carcinogenesis of Wistar rats by N-methyl-N′-nitro-N-nitrosoguanidine. Mutat Res 2003;535:73–78.
Sun Y, Deng D, You W-C, et al. Methylation of p16 CpG islands associated with malignant transformation of gastric dysplasia in a population-based study. Clin Cancer Res 2004;10:5087–5093.
Vo QN, Geradts J, Boudreau DA, et al. CDKN2A promoter methylation in gastric adenocarcinomas: clinical variables. Hum Pathol 2002;33:1200–1204.
Liu YH, Zhang LH, Ren H, et al. Promoter hypermethylation of the p16 gene in pre- and post-gastrectomy plasma of patients with gastric adenocarcinoma. Beijing Da Xue Xue Bao 2005;37:257–260.
Sun XD, Mu R, Zhou YS, et al. Analysis of mortality rate of stomach cancer and its trend in twenty years in China. Zhonghua Zhong Liu Za Zhi 2004;26:4–9.
Deng D, Deng G, Lu Y, et al. Analysis of the methylation in CpG island by denaturing high-performance liquid chromatography. Zhonghua Yi Xue Za Zhi 2001;81:158–161.
Deng D, Deng G, Smith MF, et al. Simultaneous detection of CpG methylation and single nucleotide polymorphism (SNP) by denaturing high performance liquid chromatography (DHPLC). Nucleic Acids Res 2002;30:e13.
Deng D, El-Rifai W, Ji J, et al. Hypermethylation of metallothionein-3 CpG island in gastric carcinoma. Carcinogenesis 2003;24:25–29.
Baumer A, Wiedemann U, Hergersberg M, et al. A novel MSP/DHPLC method for the investigation of the methylation status of imprinted genes enables the molecular detection of low cell mosaicisms. Human Mutat 2001;17:423–430.
Sambrook J, Fritsch EF, Maniatis T . Molecular Cloning: A Laboratory Manual, 2nd edn. Cold Spring Harbor Laboratory Press: Cold Spring Harbor, New York, 1989.
Eads CA, Laird PW . Combined bisulfite restriction analysis (COBRA). In: Mills KI, Ramsahoye BH (eds). DNA Methylation Protocols (Methods in Molecular Biology, Vol. 200). Humana Press: Totowa, New Jersey, 2002, pp 53–70.
Betz B, Florl AR, Seifert HH, et al. Denaturing high-performance liquid chromatography (DHPLC) as a reliable high-throughput prescreening method for aberrant promoter methylation in cancer. Hum Mutat 2004;23:612–620.
Herman JG, Graff JR, Myöhänen S, et al. Methylation-specific PCR: a novel PCR assay for methylation status of CpG islands. Proc Natl Acad Sci USA 1996;93:9821–9826.
Bahrami AR, Dickman MJ, Matin MM, et al. Use of fluorescent DNA-intercalating dyes in the analysis of DNA via ion-pair reversed-phase denaturing high-performance liquid chromatography. Anal Biochem 2002;309:248–252.
Eads CA, Danenberg KD, Kawakami K, et al. MethyLight: a high-throughput assay to measure DNA methylation. Nucleic Acids Res 2000;28:E32.
Acknowledgements
We thank the expert technical assistance of Jing Zhou, Liankun Gu, Ruming Wang, Guiguo Li at Beijing Institute for Cancer Research. We also appreciate Dr Benjamin Legendre Jr (Transgenomic, Inc.) for language editing. The present work was supported by National Basic Research Program of China 2005CB522403, NSFC grant 30471996, and 211 grant from Peking University.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Luo, D., Zhang, B., Lv, L. et al. Methylation of CpG islands of p16 associated with progression of primary gastric carcinomas. Lab Invest 86, 591–598 (2006). https://doi.org/10.1038/labinvest.3700415
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/labinvest.3700415
Keywords
This article is cited by
-
A similar effect of P16 hydroxymethylation and true-methylation on the prediction of malignant transformation of oral epithelial dysplasia: observation from a prospective study
BMC Cancer (2018)
-
Clinical and biological significance of a − 73A > C variation in the CDH1 promoter of patients with sporadic gastric carcinoma
Gastric Cancer (2018)
-
Down-regulated expression of OPCML predicts an unfavorable prognosis and promotes disease progression in human gastric cancer
BMC Cancer (2017)
-
P16-specific DNA methylation by engineered zinc finger methyltransferase inactivates gene transcription and promotes cancer metastasis
Genome Biology (2015)
-
RETRACTED ARTICLE: Role of p16 gene promoter methylation in gastric carcinogenesis: a meta-analysis
Molecular Biology Reports (2014)