Abstract
There exists a need for robust approaches for tandem mass spectrometry (MS/MS)-based identification of proteins in formalin-fixed paraffin-embedded (FFPE) material. We demonstrate herein the identification of proteins in FFPE material using enzymatic cleavage for extraction of peptides from the FFPE specimen and liquid chromatography (LC) followed by MS/MS. We identified 324 proteins from a 3-year-old FFPE cell-block of a human lymphoma cell line. The identified proteins were assigned to the membrane, cytosol and nucleus, with diverse cellular functions. The results were comparable to those obtained with lysates from a fresh specimen of the lymphoma cell line. Western blotting analysis and immunofluorescence microscopy confirmed the expression of selected proteins. The functional significance of one protein (PKC η) was validated using a PKC inhibitory peptide which resulted in lymphoma cell death in vitro. The ability to identify proteins from FFPE specimens has significant implications for MS/MS-based proteomics of vast repositories of archival primary tissue samples for disease-related discovery research.
Similar content being viewed by others
Main
Formalin fixation and tissue embedding in paraffin wax is a universal approach for tissue processing prior to light microscopic evaluation. A major advantage afforded by formalin-fixed paraffin-embedded (FFPE) specimens is the preservation of cellular and architectural morphologic detail in tissue sections.1 The standard buffered formalin fixative in which biopsy specimens are processed is an aqueous solution containing 37% formaldehyde and 10–15% methyl alcohol. Formaldehyde is a highly reactive dipolar compound that results in the formation of protein–nucleic acid and protein–protein crosslinks in vitro.1, 2, 3, 4, 5
FFPE specimens have not been routinely used for global mass spectrometry-based proteomic studies. This relates to the fact that formaldehyde-induced crosslinking also renders proteins relatively insoluble and unsuitable for routine biochemical extraction and analysis. This crosslinking effect has precluded protein extraction from FFPE for routine Western blotting analysis of protein expression.2 Thus, while recently developed enzymatic and heat-induced antigen retrieval methods have facilitated the immunohistologic detection of proteins in tissue, many proteins are still not detectable using these approaches.6, 7 In addition, proteins lacking widely available antibodies to their formalin-resistant epitopes are largely undetectable by immunohistochemistry. Although recent advances in molecular biotechnology have permitted the global analysis of proteins expressed in various cellular and tissue systems, the diminished capacity to extract intact proteins has largely precluded the proteomic analysis of FFPE samples with definitive identification of the proteins present within cells and tissues in biopsy material.
The development of strategies to permit utilization of the universal FFPE specimens will be important in leveraging the application of powerful mass spectrometry-based proteomic approaches into the investigation of archival clinical specimens. In this study, we have used a 3-year-old cell block preparation of a human transformed follicular lymphoma-derived cell line (SUDHL-4) to assess the feasibility of LC-MS/MS analysis on FFPE specimens. Our strategy involved a ‘bottom-up’ approach for identification of proteins in the FFPE sample by trypsin and glutamic C endopeptidase (gluC) digestion and subsequent ‘shotgun’ LC-MS/MS for protein identification. Our results establish the feasibility of utilization of FFPE for MS/MS-based proteomics analysis and with important implications for biomarker discovery and identification of novel therapeutic targets.
Materials and methods
Preparation of Cell Lysates
All cell lines (Karpas 422, OCI Ly-1, SUDHL-4 and SUDHL-16) are derived from human transformed follicular B-cell lymphomas that harbor the chromosomal aberration t(14;18).8 Cells were grown in RPMI 1640 (Gibco-BRL, Gaithersburg, MD, USA) supplemented with 10% heat-inactivated fetal bovine serum (FBS), 2 mM L-glutamine and 100 units/ml of penicillin–streptomycin mixture (Gibco BRL) at 37°C and 5% CO2. Cell cultures of late exponential growth phase (about 2–5 × 105 cells/ml) were pelleted by centrifugation at 1000 rpm at 10°C, followed by washing in PBS × 3 and lysis using RIPA buffer (25 mM Tris HCl, 0.1% SDS, 1% Triton X-100, 1% sodium deoxycholate, 0.15 M NaCl, 1 mM EDTA). Cell lysates were then homogenized by repeated pipetting or using a syringe in ice followed by microcentrifugation. The supernatants were collected as protein extracts for further experiments. Protein concentration of extracts was determined using the Bradford protein assay kit (Pierce, Rockford, Il) and stored at −80°C.
Preparation of Cell Block
The SUDHL-4 cell block was prepared by growing cells to confluency and approximately 107 cells were pelleted then mixed with Thomboplastin-DS (Pacific Hemostasis, Huntersville, NC, USA.) The mixture was allowed to incubate for 5 min while clot formation occurred. Hollande's fixative (∼5 ml) was added, and the tube gently agitated to dislodge the clotted material. After 15 min, the cell clot was transferred to a plastic tissue cassette, placed in formalin and incubated overnight. Paraffin cell blocks were stored at room temperature until analysis.
Excess paraffin was trimmed from the cell block, and the remaining paraffin-embedded cell material was finely diced and placed in a microfuge tube. Paraffin was removed using three washes of xylene, with the remaining material subjected to serial hydration in graded ethanol (100, 90, 70, and 50%) and finally washed in PBS buffer. RIPA lysate buffer (as above) was then added to the pellet and protein concentrations again estimated using the Bradford colormetric method.
Enzymatic Digestion
Duplicate samples were prepared using equal amounts of total protein (600 μg) from both the FFPE SUDHL-4 cell block and fresh SUDHL-4 cell lysate. Each sample was digested by adding freshly prepared sequencing grade modified trypsin or glutamic C endopeptidase (Princeton Separations, Adelphia, NJ, USA) to each tube (1:50) and the tubes incubated at 37°C overnight. Following the enzymatic digest, peptides were desalted and concentrated using Oasis HLB solid phase columns (Waters, Milford, MA, USA). For both samples (FFPE and fresh cell lysate), the trypsin and gluC-digested extracts were prepared in replicates of ten.
Mass Spectrometric Analysis
A 15 μl aliquot of each sample was analyzed by automated nanoflow reverse-phase LC/MS using the LCQ Deca XP ion trap mass spectrometer (ThermoElectron, San Jose, CA, USA). Digested peptides were injected by an autosampler, using an acetonitrile gradient (0–60% B in 80 min; A=5% acetonitrile with 0.4% acetic acid and 0.005% heptafluorobutyric acid (HFBA), B=95% acetonitrile 0.4% acetic acid and 0.005% HFBA) through a reverse phase column (75 μm ID fused silica packed in-house with 10 cm of 5 μm C18 particles) to elute the peptides at a flow rate of ∼200 nl/min into the mass spectrometer. An electrospray voltage of 2.2 kV was used with the ion transfer tube temperature set to 220°C. Peptide analysis was performed using data-dependent acquisition of one MS scan (600–2000 m/z) followed by MS/MS scans of the three most abundant ions in each MS scan. Normalized collision energy for MS/MS was set to 35% with an isolation width of 1.5 amu. Dynamic exclusion was set to a repeat count of 3, with the exclusion duration of 5 min. As previously described, each sample was analyzed multiple times (n=10) to ensure comprehensive identification of proteins.9
Data Analysis
The acquired MS/MS data was searched with TurboSequest against amino acid sequences in the UniProt nonredundant protein database (1/10/05 download). Peptide tolerance was set to 1.4 amu, with a minimum group count of 2 and one missed cleavage site allowed. Two or more peptides matching the criteria of a crosscorrelation score (Xcorr)>1.2 for +1 peptides, >2.2 for +2 peptides, and >3.2 for +3 peptides, and a delta correlation score (ΔCn)>0.100 was used as a threshold of acceptance. Next, all SEQUEST data (.dta) and output (.out) files passing the above criteria from the replicate samples were combined and evaluated using INTERACT and ProteinProphet (Institute for Systems Biology, Seattle, WA, USA).10, 11 All protein identifications with an error rate of less than 5% were summarized and directly exported into Excel. Each sample was then analyzed for unique and shared protein identifications using the Perl script program (iadiff.pl) INTERACT Difference (Eng J, personal communication, January 2004). Finally, UniProt accession numbers were analyzed using GOMiner,12 which maps identified proteins to existing Gene Ontology (GO) terms, with both molecular function and cell location summarized for proteins in each sample.
Immunoblot Analysis
For immunoblotting, cell lysates (40 μg of total protein per lane) were resolved in a 10% SDS-PAGE. Separated proteins were then transferred to a nitrocellulose membrane filter (Millipore, Billerica, MA, USA) using semidry transfer and blocked overnight at 4°C. The membrane was incubated at room temperature with the appropriate dilution of primary antibody, then washed and incubated with horseradish peroxidase-conjugated secondary antibodies. Western blot targets were visualized using chemicaluminescence (ECL Plus kit, Amersham, Piscataway, NJ, USA). The following antibodies were used for immunoblotting analysis: Actin (C2), JAK1 (H-106), PKC (H-300), PKC η (C-15), and Raf-B (H-145), and STAT1 (E-23) (Santa Cruz Biotechnology, Santa Cruz, CA, USA).
Immunofluorescence Microscopy
For immunofluorescence, SUDHL-4 cells (1 × 106cells/ml) were washed, plated, and fixed in 24 well plates then subjected to immunofluorescence analysis using an antibody against phospholipase C-gamma-1 (PLCγ1, 503) (Santa Cruz). Control wells were also included for the secondary antibody, and the cells were examined by fluorescent microscopy, with fluorescent signal thresholds set using the control wells.
Functional Validation Studies
To determine the functional significance of protein kinase C eta (PKC η) expression in the human lymphoma-derived SUDHL-4 cells, we performed studies to evaluate the effect of PKC inhibition on the viability of the lymphoma cells. For our experiments, we utilized the cell permeable PKC inhibitor peptide (Myr-RKRTLRRL) which inhibits binding of PKC substrates.13 The PKC inhibitor peptide (Calbiochem, San Diego, CA, USA) was dissolved in PBS as the vehicle and incubated with the SUDHL-4 cells at varying concentrations and time points. Experiments for each drug concentration and time point were performed in triplicate.
Cells at a starting density of 1 × 105 cells/ml were used for MTT studies. Cell viability was determined using the MTT [3-(4,5-dimethyl thiazol-2-yl)-2,5-diphenyl tetrazolium bromide] assay as previously described.14 Cell cycle analysis was performed as previously described.15
Results
Proteins were extracted from a 3-year-old FFPE block of a tFL-derived cell line (SUDHL-4), digested with trypsin or gluC, and analyzed by reverse phase nanoflow LC-MS/MS (Figure 1). To obtain a comprehensive survey of proteins, parallel enzymatic digestion (trypsin and gluC) was performed for each sample type.16, 17 Each digest was analyzed by LC-MS/MS multiple times (n=10) to approach maximal saturation levels of protein identification.9 The cumulative totals of proteins identified in replicate analysis (n=10) were 324 and 514 (Figure 2a) for the paraffin and fresh lysate samples, respectively. Trypsin digestions yielded 133 protein identifications in the FFPE sample and 231 from the fresh lymphoma lysate, while the gluC-digest resulted in 71 proteins identified from the FFPE cells, and 86 from the fresh lysate. Using peptides from both digests (trypsin and gluC), 120 proteins were identified from the FFPE material and 197 proteins identified from fresh cell lysate. Cumulative protein identifications were then combined from each analysis and for both enzyme digests as shown in Figure 2b.
Comprehensive survey of proteins identified using tandem mass spectrometry by replicate analysis (n=10) of (a) SUDHL-4 FFPE cell block (324 proteins) and SUDHL-4 fresh cell lysate (514 proteins). (b) Cumulative protein identifications were combined for both enzyme digests and from each analysis. In some cases, proteins were identified by complementary peptide contributions from trypsin and gluC. The remaining protein identifications were exclusive to either enzyme.
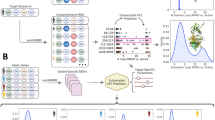
Functional categories of a selected list of proteins identified in common to both the FFPE and fresh cell lysate samples are presented in Table 1. In detail, MS analysis of the extracted proteins from the FFPE specimen yielded 4876 peptides, which matched 1471 known database entries and resulted in the identification of 324 unique proteins (see Supplementary Table 1 for full list) based on a two peptide per protein minimum, standard cross correlation (Xcorr) and delta correlation (ΔCn) scores and INTERACT estimated error rate <5% (see materials and methods). In contrast, MS analysis of proteins extracted from the fresh SUDHL-4 cell lysate resulted in a total of 10043 peptides, which matched 2284 known database entries and 514 unique proteins (see Supplementary Table 2 for full list). As shown in Figure 3 263 proteins were found in common between the two sample types (see Supplementary Table 3 for full list). This represents 81% of the total number identified from the FFPE extraction. Importantly, more than half (52%) of the proteins identified in the fresh cell lysate were also identified from the 3-year-old FFPE material. Figure 4 shows the MS full scan and data-dependent MS/MS scan of the tryptic peptide R.EFIWGVFGK that identified protein kinase C, eta from the FFPE block. Additional peptides are displayed in Table 1 which summarizes the Gene Ontology molecular function of selected proteins identified in both the fresh SUDHL-4 cell lysate and FFPE sample.
MS full scan and data-dependent MS/MS scan showing sequencing of the tryptic peptide R.EFIWGVFGK which identified protein kinase C, eta in the FFPE sample. Peptide sequencing is indicated by matching b ion (red) and y ion (blue) fragments. Additional peptides for protein kinase C are displayed in Table 1.
Figure 5 shows that the total number of proteins identified in the FFPE sample was generally less than that obtained with the fresh cell lysate for both cell location (Figure 5a) and molecular function (Figure 5b). The distribution of proteins identified in the SUDHL-4 FFPE cell block is displayed as a pie chart in Figure 6a, showing both cell location and molecular function. Similarly, Figure 6b shows the cell location and molecular function for the proteins identified in the fresh SUDHL-4 cell lysate. Predicted cell location and molecular function for the 263 proteins identified in common to both samples is provided in Figure 6c.
In order to further validate the proteins identified by MS/MS peptide matching, we performed Western blot analysis and immunofluorescence studies to demonstrate the expression of proteins identified in both the SUDHL-4 FFPE cell block and fresh cell lysate. As shown in Figure 7a, the expression of Raf-B, JAK1, PKC, and STAT1 was observed by immunoblot analysis in the SUDHL-4 cells and not the IgG isotype control lane, confirming the presence of these proteins in the SUDHL-4 proteome. Cytoplasmic expression of PLCγ1 was confirmed by immunofluorescence microscopy in SUDHL-4 cells (Figure 7b).
Validation of selected LC-MS/MS protein identifications by (a) immunoblot analysis (Raf-B, JAK1, PKC, STAT1) and (b) immunofluorescence microscopy (PLCγ1). (c) Western blotting analysis corroborates PKC η expression in the human lymphoma-derived cell line (SUDHL-4). Interestingly, three other human lymphoma cell lines (Karpas 422, OCI-Ly1, and SUDHL-16) demonstrate PKCη expression by Western blotting analysis. (d) MTT cell proliferation assay shows dose- and time-dependent inhibition of SUDHL-4 cell growth. (e) Cell cycle analysis of SUDHL-4 cells after 52 h of inhibitor treatment. M1=dead cells, M2=G0/G1, M3=S-phase and M4=G2/M.
We also validated the expression of protein kinase C eta (PKC η) in the SUDHL-4 cell line, as well as, three other tFL-derived cell lines (Figure 7c). We further investigated the potential functional significance of PKC η expression in the SUDHL-4 cells, by performing studies to evaluate the effect of PKC inhibition on the viability of the lymphoma cells. The PKC inhibitor resulted in significant inhibition of the growth of SUDHL-4 cells as demonstrated by the MTT viability assay (Figure 7d) at a calculated IC50 value of 11 μM. In addition, PKC inhibition resulted in G0/G1 (M2) arrest, decreased S-phase fraction, and induction of cell death (M1) in the SUDHL-4 lymphoma cells 52 h after inhibitor treatment (Figure 7e).
Discussion
We report the direct identification of proteins isolated from an approach involving deparaffinization, peptide extraction by enzymatic digestion, and analysis by reversed-phase LC-MS/MS. We identified 324 proteins in the SUDHL-4 FFPE cell block, including proteins known to localize primarily to the membrane, cytosol and nucleus. Comparative proteomic analysis performed on protein extracts from fresh lymphoma cells revealed an overlap of 263 proteins identified from the FFPE samples (52% of fresh lysate total IDs). Western blotting and immunofluorescence studies confirmed the expression of selected proteins identified by peptide extraction and LC-MS/MS in the FFPE specimen. Notably, the extraction time for LC-MS/MS sample preparation of FFPE samples was similar to DNA/RNA extraction from paraffin-embedded tissue samples, with the deparaffinizing step as the only additional labor required beyond that of preparing traditional samples for analysis by LC-MS/MS. The overall yield of extracted protein, peptide chromatography, and quality of MS/MS spectra was comparable to that of fresh cell lysate.
Detailed examination of the cellular compartments of the proteins identified revealed relative biases for detection of proteins from particular compartments (Figure 5). For example, the number of extracellular compartment proteins identified from the FFPE sample (n=29) was comparable to that identified in the fresh lymphoma cell lysate (n=34), and represented only a 15% reduction in protein identifications (Figure 5a). In either sample, proteins assigned as ‘extracellular’ are considered to be precursor proteins destined for export to the extracellular compartment. The number of cytosolic proteins identified in the FFPE sample (n=125) was also reduced in comparison to those identified in the fresh lymphoma cell lysate (n=183), representing a 32% reduction in proteins identified in the fresh cell lysate. Membrane protein identifications were also under represented in the FFPE sample (n=100) as compared to the fresh cell lysate (n=162), representing a 39% reduction in the membrane proteins identified in the fresh cell lysate. Correspondingly, proteins assigned to signaling, catalytic or enzymatic (cytosolic), and transport (membrane) functions were less represented in the FFPE samples (Figure 5b). On the other hand, analysis of the FFPE sample yielded 65 proteins assigned to the nuclear compartment as compared to 125 proteins in the fresh lymphoma cell lysate, representing a 48% reduction of the proteins identified in fresh cell lysate. Accordingly, proteins assigned to transcription (nuclear) function were represented least in the FFPE specimen, as compared to the fresh cell lysate (Figure 5b).
Interestingly, some proteins were identified exclusively in the FFPE specimen (n=61), as compared to 251 proteins identified exclusively in the fresh cell lysate (Figure 3). In addition, analysis of the FFPE sample afforded identification of important low-abundance proteins including transcription factors such as NF-κB p105 subunit, cytokines such as leukocyte cell-derived chemotaxin 2, kinases such as leukocyte tyrosine kinase, phosphatases such as protein-tyrosine phosphatase nonreceptor type 22, receptors such as ryanodine receptor 1 and signaling proteins such as Rho-GTPase-activating protein 6 (Table 1 and Supplementary Table 1). Moreover, several proteins previously reported to play an important role in the pathogenesis of tFL were identified in the FFPE specimen. These include NF-κB,18 BCL-11A,19 BAL-associated protein,20 and RhoF.21 Importantly, many proteins previously unknown to be expressed in tFL such as ribosomal S6 kinase, Wnt2 protein, beta-catenin like protein1, and the eta (η) isoform of protein kinase C (PKC) were identified in the FFPE sample.
With regard to protein kinase C, the PKC isoenzyme family members are signal transduction regulators that exhibit cell and tissue-specific expression. PKC η has been implicated in the mediation of proliferative signals important in the pathogenesis and progression of some human cancers.22, 23, 24 However, such a role has not been established in the regulation of growth of transformed follicular lymphoma. Hence, we sought to establish a functional role for PKC η expression in tFL. In this regard, we demonstrated evidence for a dose- and time-dependent inhibition of SUDHL-4 cell growth, following exposure of the lymphoma cells to a PKC inhibitor. These results further validate the utility of FFPE-based proteomics approaches for the identification of biologically relevant proteins which may serve as rational targets for novel therapeutic strategies.
In conclusion, the identification of proteins of diverse function and subcellular localization from FFPE material demonstrates the feasibility of mass spectrometry-based proteomic studies for the investigation of archived pathologic specimens previously inaccessible to direct proteomic analysis. The utilization of proteolytic cleavage for peptide extraction from FFPE samples circumvents the problems associated with routine biochemical extraction of intact proteins from FFPE, or recovery and analysis of insoluble or crosslinked proteins from gel electrophoresis-based approaches. Our current approach utilized two enzymes with distinct cleavage specificities which improved the overall number and validity of the protein identifications. We identified several proteins known to be expressed in human-transformed follicular B-cell lymphoma cells, from which the SUDHL-4 cell line is derived. Importantly, several proteins of potential functional and pathogenetic significance, which were not known to be expressed in this subtype of lymphoma, were identified by analysis of the 3-year-old FFPE specimen. Application of multidimensional chromatography separation techniques would increase the number of definitive identifications, and improve the identification of low-abundance proteins.25 Utilization of specific cell isolation and enrichment techniques such as laser capture microdissection will permit characterization of pure cell populations in their precise microanatomic localization.26 Future studies examining FFPE specimens that have been stored for varying lengths of time will be necessary to determine the effects of storage on peptide recovery and protein identification by bottom-up proteomics approaches as described herein. The ability to perform mass spectrometry-based proteomic analysis on FFPE samples provides significant opportunities for biomarker and therapeutic target discovery in archival samples with well-documented clinical follow-up.
References
Fox CH, Johnson FB, Whiting J, et al. Formaldehyde fixation. J Histochem Cytochem 1985;33:845–853.
Clark RK, Tani Y, Damjanov I . Suppression of nonspecific binding of avidin-biotin complex (ABC) to proteins electroblotted to nitrocellulose paper. J Histochem Cytochem 1986;34:1509–1512.
McGhee JD, von Hippel PH . Formaldehyde as a probe of DNA structure. I. Reaction with exocyclic amino groups of DNA bases. Biochemistry 1975;14:1281–1296.
McGhee JD, von Hippel PH . Formaldehyde as a probe of DNA structure. II. Reaction with endocyclic imino groups of DNA bases. Biochemistry 1975;14:1297–1303.
Kunkel GR, Mehrabian M, Martinson HG . Contact-site cross-linking agents. Mol Cell Biochem 1981;34:3–13.
Shi SR, Key ME, Kalra KL . Antigen retrieval in formalin-fixed, paraffin-embedded tissues: an enhancement method for immunohistochemical staining based on microwave oven heating of tissue sections. J Histochem Cytochem 1991;39:741–748.
van den Berg FM, Baas IO, Polak MM, et al. Detection of p53 overexpression in routinely paraffin-embedded tissue of human carcinomas using a novel target unmasking fluid. Am J Pathol 1993;142:381–385.
Tsujimoto Y, Cossman J, Jaffe E, et al. Involvement of the bcl-2 gene in human follicular lymphoma. Science 1985;228:1440–1443.
Liu H, Sadygov RG, Yates III JR . A model for random sampling and estimation of relative protein abundance in shotgun proteomics. Anal Chem 2004;76:4193–4201.
Keller A, Nesvizhskii AI, Kolker E, et al. Empirical statistical model to estimate the accuracy of peptide identifications made by MS/MS and database search. Anal Chem 2002;74:5383–5392.
Nesvizhskii AI, Keller A, Kolker E, et al. A statistical model for identifying proteins by tandem mass spectrometry. Anal Chem 2003;75:4646–4658.
Zeeberg BR, Feng W, Wang G, et al. GoMiner: a resource for biological interpretation of genomic and proteomic data. Genome Biol 2003;4:R28.
Ward NE, O’Brian CA . Inhibition of protein kinase C by N-myristoylated peptide substrate analogs. Biochemistry 1993;32:11903–11909.
Lin Z, Lim S, Viani MA, et al. Down-regulation of telomerase activity in malignant lymphomas by radiation and chemotherapeutic agents. Am J Pathol 2001;159:711–719.
Lin Z, Lim S, Lim MS . Growth regulation by p27Kip1 is abrogated by multiple mechanisms in aggressive malignant lymphomas. Br J Haematol 2003;12:739–748.
MacCoss MJ, McDonald WH, Saraf A, et al. Shotgun identification of protein modifications from protein complexes and lens tissue. Proc Natl Acad Sci USA 2002;99:7900–7905.
Gatlin CL, Eng JK, Cross ST, et al. Automated identification of amino acid sequence variations in proteins by HPLC/microspray tandem mass spectrometry. Anal Chem 2000;72:757–763.
Heckman CA, Mehew JW, Boxer LM . NF-kappaB activates Bcl-2 expression in t(14;18) lymphoma cells. Oncogene 2002;21:3898–3908.
Satterwhite E, Sonoki T, Willis TG, et al. The BCL11 gene family: involvement of BCL11A in lymphoid malignancies. Blood 2001;98:3413–3420.
Aguiar RC, Yakushijin Y, Kharbanda S, et al. BAL is a novel risk-related gene in diffuse large B-cell lymphomas that enhances cellular migration. Blood 2000;96:4328–4334.
Gouw LG, Reading NS, Jenson SD, et al. Expression of the Rho-family GTPase gene RHOF in lymphocyte subsets and malignant lymphomas. Br J Haematol 2005;129:531–533.
Aeder SE, Martin PM, Soh JW, et al. PKC-eta mediates glioblastoma cell proliferation through the Akt and mTOR signaling pathways. Oncogene 2004;23:9062–9069.
Sonnemann J, Gekeler V, Ahlbrecht K, et al. Down-regulation of protein kinase Ceta by antisense oligonucleotides sensitises A549 lung cancer cells to vincristine and paclitaxel. Cancer Lett 2004;209:177–185.
Brenner W, Farber G, Herget T, et al. Protein kinase C eta is associated with progression of renal cell carcinoma (RCC). Anticancer Res 2003;23:4001–4006.
Wolters DA, Washburn MP, Yates III JR . An automated multidimensional protein identification technology for shotgun proteomics. Anal Chem 200;73:5683–5690.
Emmert-Buck MR, Bonner RF, Smith PD, et al. Laser capture microdissection. Science 1996;274:998–1001.
Acknowledgements
This work was supported by the ARUP Institute for Clinical and Experimental Pathology.
Author information
Authors and Affiliations
Corresponding author
Additional information
Supplementary Information accompanies the paper on Laboratory Investigation website (http://www.nature.com/labinvest)
Supplementary information
Rights and permissions
About this article
Cite this article
Crockett, D., Lin, Z., Vaughn, C. et al. Identification of proteins from formalin-fixed paraffin-embedded cells by LC-MS/MS. Lab Invest 85, 1405–1415 (2005). https://doi.org/10.1038/labinvest.3700343
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/labinvest.3700343
Keywords
This article is cited by
-
Comparative proteomic analysis of children FSGS FFPE tissues
BMC Pediatrics (2022)
-
Laser capture microdissection coupled mass spectrometry (LCM-MS) for spatially resolved analysis of formalin-fixed and stained human lung tissues
Clinical Proteomics (2020)
-
Quantitative Proteomic Analysis of Optimal Cutting Temperature (OCT) Embedded Core-Needle Biopsy of Lung Cancer
Journal of the American Society for Mass Spectrometry (2017)
-
MALDI imaging mass spectrometry of N-linked glycans on formalin-fixed paraffin-embedded murine kidney
Analytical and Bioanalytical Chemistry (2015)
-
Extraction of tamoxifen and its metabolites from formalin-fixed, paraffin-embedded tissues: an innovative quantitation method using liquid chromatography and tandem mass spectrometry
Cancer Chemotherapy and Pharmacology (2014)