Abstract
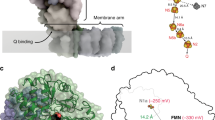
PROTEINS that contain disulphide bonds are often slow to fold in vitro because the oxidation and correct pairing of the cysteine residues is rate limiting1,2. The folding of such proteins is greatly accelerated in Escherichia coli by DsbA3,4, but the mechanism of this rate enhancement is not well understood. Here we report the crystal structure of oxidized DsbA and show that it resembles closely the ubiquitous redox protein thioredoxin5, despite very low sequence similarity. An important difference, however, is the presence of another domain which forms a cap over the thioredoxin-like active site of DsbA. The redox–active disulphide bond, which is responsible for the oxidation of substrates, is thus at a domain interface and is surrounded by grooves and exposed hydrophobic side chains. These features suggest that DsbA might act by binding to partially folded polypeptide chains before oxidation of cysteine residues.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Creighton, T. E. Meth. Enzym. 107, 305–329 (1984).
Weissman, J. S. & Kim, P. S. Cell 71, 841–851 (1992).
Bardwell, J. C. A., McGovern, K. & Beckwith, J. Cell 67, 581–589 (1991).
Kamitani, S., Akiyama, Y. & Ito, K. EMBO J. 11, 57–62 (1992).
Eklund, H., Gleason, F. S. & Holmgren, A. Proteins Struct. Funct. Genet. 11, 13–28 (1991).
Lovejoy, B. et al. Science 259, 1288–1293 (1993).
Brown, L. R. et al. J. molec. Biol. 231, 800–816 (1993).
Katti, S. K., LeMaster, D. M. & Eklund, H. J. molec. Biol. 212, 167–184 (1990).
Tomb, J.-F. Proc. natn. Acad. Sci. U.S.A. 89, 10252–10256 (1992).
Ellis, L. B. M., Saurugger, P. & Woodward, C. Biochemistry 31, 4882–4891 (1992).
Eklund, H. et al. J. molec. Biol. 228, 596–618 (1992).
Sodano, P. et al. J. molec. Biol. 221, 1311–1324 (1991).
Xia, T.-H. et al. Prot. Sci. 1, 310–321 (1992).
Epp, O., Ladenstein, R. & Wendel, A. Eur. J. Biochem. 133, 51–69 (1983).
Reinemer, P. et al. EMBO J. 10, 1997–2005 (1991).
Ji, X., Zhang, P., Armstrong, R. N. & Gilliland, G. L. Biochemistry 31, 10169–10184 (1992).
Thornton, J. M. J. molec. Biol. 151, 261–287 (1981).
Bardwell, J. C. A. et al. Proc. natn. Acad. Sci. U.S.A. 90, 11038–1042 (1993).
Dailey, F. E. & Berg, H. C. Proc. natn. Acad. Sci. U.S.A. 90, 1043–1047 (1993).
Zapun, A., Bardwell, J. C. A. & Creighton, T. E. Biochemistry (in the press).
Wünderlich, M., Jaenicke, R. & Glockshuber, R. J. molec. Biol. (in the press).
Wünderlich, M. & Glockshuber, R. Prot. Sci. 2, 717–726 (1993).
Krause, G., Lundström, J., Lopez-Barea, J., Pueyo de La Cuesta, C. & Holmgren, A. J. biol. Chem. 266, 9494–9500 (1991).
Martin, J. L., Waksman, G., Bardwell, J. C. A., Beckwith, J. & Kuriyan, J. J. molec. Biol. 230, 1097–1100 (1993).
Edman, J. C., Ellis, L., Blacher, R. W., Roth, R. R. & Rutter, W. J. Nature 317, 267–270 (1985).
Wilson, K. S. Acta crystallogr. B34, 1599–1608 (1978).
Terwilliger, T. C. & Eisenberg, D. Acta crystallogr. A39, 813–817 (1983).
Weis, W. I., Kahn, R., Fourme, R., Drickamer, K. & Hendrickson, W. A. Science 254, 1608–1615 (1991).
Jones, T. A., Zou, J. Y., Cowan, S. W. & Kjeldgaard, M. Acta cystallogr. A47, 110–119 (1991).
Wang, B. C. Meth. Enzym. 115, 90–112 (1985).
Zhang, K. Y. J. & Main, P. Acta crystallogr. A46, 41–46 (1990).
Brünger, A. T. X-PLOR (Version 3.1) Manual (The Howard Hughes Medical Institute and Department of Molecular Biophysics and Biochemistry, Yale University, 260 Whitney Avenue, New Haven, CT 06511, 1992).
Bernstein, F. C. et al. J. molec. Biol. 112, 535–542 (1977).
Nicholls, A., Sharp, K. A. & Honig, B. Proteins Struct. Funct. Genet. 11, 281–296 (1991).
Hendrickson, W. A. Science 254, 51–58 (1991).
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Martin, J., Bardwell, J. & Kuriyan, J. Crystal structure of the DsbA protein required for disulphide bond formation in vivo. Nature 365, 464–468 (1993). https://doi.org/10.1038/365464a0
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/365464a0
This article is cited by
-
OsnR is an autoregulatory negative transcription factor controlling redox-dependent stress responses in Corynebacterium glutamicum
Microbial Cell Factories (2021)
-
A shape-shifting redox foldase contributes to Proteus mirabilis copper resistance
Nature Communications (2017)
-
Chemical shift assignments of a reduced N-terminal truncation mutant of the disulfide bond isomerase TrbB from plasmid F, TrbBΔ29
Biomolecular NMR Assignments (2014)
-
Structural and biochemical characterization of the essential DsbA-like disulfide bond forming protein from Mycobacterium tuberculosis
BMC Structural Biology (2013)
-
Structure analysis of the extracellular domain reveals disulfide bond forming-protein properties of Mycobacterium tuberculosis Rv2969c
Protein & Cell (2013)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.