Abstract
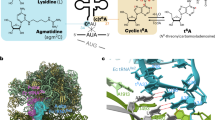
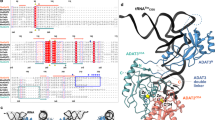
AMINOACYL-RNA synthetases can be divided into two classes according to structural features inferred from sequence alignments1–3. This classification correlates almost perfectly with the attachment of the amino acid to the 2'-OH (class I) or 3'-OH (class II) group of the terminal adenosine4–6. Six subgroups of higher homology can be inferred from sequence analysis7,8. The five aminoacyl-tRNA synthetases whose crystal structures are known (MetRS, TyrRS and GlnRS in class I, SerRS and AspRS in class II) 9–13 belong to different subgroups. Two of them, GlnRS and AspRS, have been cocrystallized with their cognate tRNA11,13. AspRS, like six other members of class II, is an a2 dimer. Yeast tRNAAsp exhibits five identity determinants: the three anticodon bases, the discriminator base G73 and the base pair G10-U2514. We report here that the refined crystal structure of AspRS com-plexed with tRNAAsp at 2.9 A resolution reveals three regions of contact, each involving a domain of AspRS and at least one identity determinant of tRNAAsp. The mode of binding of the acceptor stem of tRNAAsp by AspRS can be generalized to class II aminoacyl-tRNA synthetases, whereas the deciphering of the anticodon, which involves a large conformational change of the loop and the formation of a bulge, is more specific to the aspartic system.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Eriani, G., Delarue, M., Poch, O., Gangloff, J. & Moras, D. Nature 347, 203–206 (1990).
Webster, T., Tsai, H., Kula, M., Mackie, G. A. & Schimmel, P. Science 226, 1315–1317 (1984).
Hountondji, C., Dessen, P. & Blanquet, S. Biochimie 68, 1071–1078 (1986).
Fraser, T. H. & Rich, A. Proc. natn. Acad Sci. U.S.A. 72, 3044–3048 (1975).
Von der Haar, F. & Cramer, F. Biochemistry 15, 4131–4136, (1976).
Hecht, S. M. Transfer RNA: Structures, Properties and Recognition (eds P. P. Schimmel, D. Söll & J. N. Abelson) 345–360 (Cold Spring Harbor Laboratory Press, New York, 1979).
Cusack, S., Härtlein, M. & Leberman, R. Nucleic Acids Res. 19, 3489–3498 (1991).
Moras, D. Trends biochem. Sci. 17, 159–164 (1992).
Brick, P., Bhat, T. N. & Blow, D. M. J. molec. Biol. 208, 83–98 (1989).
Brunie, S., Zelwer, C. & Risler, J. L. J. molec. Biol. 216, 411–424 (1990).
Rould, M. A., Perona, J. J., Söll, D. & Steitz, T. A. Science 246, 1135–1142 (1989).
Cusack, S., Berthet-Colominas, C., Härtlein, M., Nassar, N. & Leberman, R. Nature 347, 249–255 (1990).
Ruff, M. et al. Science 252, 1682–1689 (1991).
Pütz, J., Puglisi, J. D., Florentz, C. & Giegé, R. Science 252, 1696–1699 (1991).
Hynes, T. R. & Fox, R. O. Proteins 10, 92–105 (1991).
Sixma, T. K. Nature 355, 561–564 (1992).
Stein, P. E., Boodhoo, A., Tyrrell, G. J., Brunton, J. L. & Read, R. J. Nature 355, 748–750 (1992).
Eriani, G., Dirheimer, G. & Gangloff, J. Nucleic Acids Res. 18, 7109–7117 (1990).
Anselme, J. & Hartlein, M. Gene 84, 481–485 (1989).
Leveque, F., Plateau, P., Dessen, P. & Blanquet, S. Nucleic Acids Research 18, 305–312 (1990).
Moras, D. et al. Nature 288, 669–674 (1980).
Westhof, E., Dumas, P. & Moras, D. J. molec. Biol. 184, 119–145 (1985).
McClain, W. H. & Foss, K. Science 241, 1804–1807 (1988).
Hou, Y. M. & Schimmel, P. Nature 333, 140–145 (1988).
Kraulis, P. J. J. appl. Crystallogr. 24, 946–950 (1991).
Eriani, G., et al. Eur. J. Biochem. 200, 337–343 (1991).
Brünger, A. T. J. molec. Biol. 203, 803–816 (1988).
Carson, M. K. J. appl. Crystallogr. 24, 958–961 (1991).
Garcia, A. & Giegé, R. Biochem. biophys. Res. Commun. 186, 956–962 (1992).
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Cavarelli, J., Rees, B., Ruff, M. et al. Yeast tRNAAsp recognition by its cognate class II aminoacyl-tRNA synthetase. Nature 362, 181–184 (1993). https://doi.org/10.1038/362181a0
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/362181a0
This article is cited by
-
Structural basis for mRNA recognition by elongation factor SelB
Nature Structural & Molecular Biology (2005)
-
Structure of the trp RNA-binding attenuation protein, TRAP, bound to RNA
Nature (1999)
-
Structural basis for recognition of the tra mRNA precursor by the Sex-lethal protein
Nature (1999)
-
The NMR structure of the RNA binding domain of E.coli rho factor suggests possible RNA–protein interactions
Nature Structural Biology (1998)
-
RNA chiropractics
Nature Structural Biology (1998)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.