Abstract
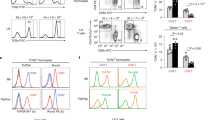
RECOMBINATION of V-, D- and J-gene segments can generate an enormous diversity of T-cell antigen receptor (TCR) gene sequences1,2. Although many γδ T cells fully exploit this diversification process, those in the epidermal and vaginal epithelium do not3,4, predominantly expressing invariant γδ receptors in which the V–(D)–J junctional sequences in almost all the productive rearrangements are identical. The almost exclusive use of identical TCRs by cells in these sites is thought to reflect recognition of a stress-induced autologous antigen5–8. To explain the prevalence of the invariant junctional sequences, it has been proposed that thymic selection operates on a population of originally diverse progenitor cells, resulting in a homogeneous repertoire9,10. Alternatively the invariant sequences may result from biases in the recombination machinery in the fetal thymic progenitors of these cells8,11,12. We report here the use of mice into which mutated TCR γ-gene rearrangement substrates have been introduced as transgenes to demonstrate directly that the canonical TCR Vγ3–Jγl and Vγ4–Jγl sequences occur at high frequency in the absence of the possibility of selection for the protein products.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Davis, M. M. & Bjorkman, P. J. Nature 334, 395–402 (1988).
Tonegawa, S. Nature 302, 575–581 (1983).
Asarnow, D. M., Goodman, T., LeFrancois, L. & Allison, J. P. Nature 341, 60–62 (1989).
Itohara, S. et al. Nature 343, 754–757 (1990).
Asarnow, D. M. et al. Cell 55, 837–847 (1988).
Havran, W. L., Chien, Y. H. & Allison, J. P. Science 252, 1430–1432 (1991).
Janeway, C. A. Jr, Jones, B. & Hayday, A. Immun. Today 9, 73–76 (1988).
Allison, J. P. & Havran, W. L. A. Rev. Immun. 9, 679–705 (1991).
Lafaille, J. J., DeCloux, A., Bonneville, M., Takagaki, Y. & Tonegawa, S. Cell 59, 859–870 (1989).
Itohara, S. & Tonegawa, S. Proc. natn. Acad Sci. U.S.A. 87, 7935–7938 (1990).
Gu, H., Forster, I. & Rajewsky, K. EMBO J. 9, 2133–2140 (1990).
Raulet, D. et al. Immun. Rev. 120, 185–204 (1991).
Garman, R. D., Doherty, P. J. & Raulet, D. H. Cell 45, 733–742 (1986).
Spencer, D. M., Hsiang, Y.-H., Goldman, J. P. & Raulet, D. H. Proc. natn. Acad Sci. U.S.A. 88, 800–804 (1991).
Kappes, D., Browne, C. P. & Tonegawa, S. Proc. natn. Acad. Sci. U.S.A. 88, 2204–2208 (1991).
Nandi, D. & Allison, J. P. J. Immun. 147, 1773–1778 (1991).
McCormack, W. T. et al. Cell 56, 785–791 (1989).
Engler, P., Klotz, E. & Storb, U. J. exp. Med. 176, 1399–1404 (1992).
Alt, F. W. & Baltimore, D. Proc. natn. Acad. Sci. U.S.A. 79, 4118–4122 (1982).
Goldman, J., Spencer, D. & Raulet, D. J. exp. Med. (in the press).
Heilig, J. S. & Tonegawa, S. Nature 322, 836–840 (1986).
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Asarnow, D., Cado, D. & Raulet, D. Selection is not required to produce invariant T-cell receptor γ-gene junctional sequences. Nature 362, 158–160 (1993). https://doi.org/10.1038/362158a0
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/362158a0
This article is cited by
-
Origin and evolutionary malleability of T cell receptor α diversity
Nature (2023)
-
Selection of the cutaneous intraepithelial γδ+ T cell repertoire by a thymic stromal determinant
Nature Immunology (2006)
-
Interleukin 15 controls the generation of the restricted T cell receptor repertoire of γδ intestinal intraepithelial lymphocytes
Nature Immunology (2005)
-
Characterization of the T-cell receptor gamma locus and analysis of the variable gene segment expression in rabbit
Immunogenetics (2005)
-
Selection of evolutionarily conserved mucosal-associated invariant T cells by MR1
Nature (2003)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.