Abstract
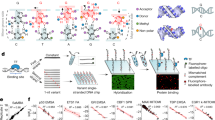
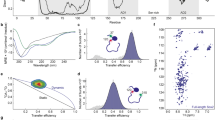
PROTEIN-DNA recognition is often mediated by a small domain containing a recognizable structural motif, such as the helix–turn–helix1 or the zinc-finger2. These motifs are compact structures that dock against the DNA double helix. Another DNA recognition motif, found in a highly conserved family of eukaryotic transcription factors including C/EPB, Fos, Jun and CREB, consists of a coiled-coil dimerization element—the leucine-zipper—and an adjoining basic region which mediates DNA binding3. Here we describe circular dichroism and 1NMR spectroscopic studies of another family member, the yeast transcriptional activator GCN44,5. The 58-residue DNA-binding domain of GCN4, GCN4-p, exhibits a concentration-dependent α-helical transition, in accord with previous studies of the dimerization properties of an isolated leucine-zipper peptide6. The GCN4-p dimer is ∼70% helical at 25 °C, implying that the basic region adjacent to the leucine zipper is largely unstructured in the absence of DNA. Strikingly, addition of DNA containing a GCN4 binding site (AP-1 site) increases the α-helix content of GNC4-p to at least 95%. Thus, the basic region acquires substantial α-helical structure when it binds to DNA. A similar folding transition is observed on GCN4-p binding to the related ATF/CREB site, which contains an additional central base pair. The accommodation of DNA target sites of different lengths clearly requires some flexibility in the GCN4 binding domain, despite its high α-helix content. Our results indicate that the GCN4 basic region is significantly unfolded at 25 °C and that its folded, α-helical conformation is stabilized by binding to DNA.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Pabo, C. O. & Sauer, R. T. A. Rev. Biochem. 53, 293–321 (1984).
Klug, A. & Rhodes, D. Trends Biochem. Sci. 12, 464–467 (1987).
Landschulz, W. H., Johnson, P. F. & McKnight, S. L. Science 243, 1681–1686 (1989).
Hope, I. A. & Struhl, K. Cell 43, 177–188 (1985).
Hope, I. A. & Struhl, K. EMBO J. 6, 2781–2784 (1987).
O'Shea, E. K., Rutkowski, R. & Kim, P. S. Science 243, 538–542 (1989).
Hope, I. A. & Struhl, K. Cell 46, 885–894 (1986).
Oas, T. G., McIntosh, L. P., O'Shea, E. K., Dahlquist, F. W. & Kim, P. S. Biochemistry 29, 2891–2894 (1990).
Kouzarides, T. & Ziff, E. Nature 340, 568–571 (1989).
Sellars, J. W. & Struhl, K. Nature 341, 74–76 (1989).
Landschultz, W. H., Johnson, P. F. & McKnight, S. L. Science 246, 922–926 (1988).
Agre, P., Johnson, P. F. & McKnight, S. L. Scinece 246, 922–926 (1989).
Weiss, M. A. Biochemistry (in the press).
Oliphant, A. R., Brandl, C. J. & Struhl, K. Molec. cell. Biol. 9, 2944–2949 (1989).
Hai, T., Liu, F., Allegretto, E. A., Karin, M. & Green, M. R. Genes Dev. 2, 1216–1226 (1988).
Sadler, J. R., Sasmor, H. & Betz, J. L. Proc. natn. Acad. Sci. U.S.A. 80, 6785–6789 (1983).
Arndt, K., Boschelli, F., Lu, P. & Miller, J. H. Biochemistry 20, 6109–6118 (1981).
Buck, F., Ruterjans, H. & Beyreuther, K. FEBS Lett. 96, 335–338 (1978).
Wade-Jardetzky, N. et al. J. molec. Biol. 128, 259–264 (1979).
Vinson, C. R., Sigler, P. B. & McKnight, S. L. Science 246, 911–916 (1989).
Gartertberg, M. R., Ampe, C., Steitz, T. A. & Crothers, D. M. Proc. natn. Acad. Sci. U.S.A 87, 6034–6038 (1990).
Oakly, M. G. & Dervan, P. B. Science 248, 847–850 (1990).
Jordan, S. & Pabo, C. O. Science 242, 895–899 (1988).
Rosenberg, A. H. et al. Gene 56, 125–135 (1987).
Studier, F. W. & Moffat, B. A. J. molec. Biol. 189, 113–130 (1986).
Bowie, J. U. & Sauer, R. T. Biochemistry 28, 7139–7143 (1989).
O'Shea, E. K., Rutkowski, R., Stafford, W. F. III & Kim, P. S. Science 243, 1689–1694 (1989).
Lehrer, S. S., Qian, Y. & Hvidt, S. Science 246, 926–928 (1989).
Chen, Y.-H., Yang, J. T. & Chou, K. H. Biochemistry 13, 3350–3356 (1974).
Johnson, W. C. Jr Protein Secondary Structure and Circular Dichroism: a Practical Guide 205–214 (1990).
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Weiss, M., Ellenberger, T., Wobbe, C. et al. Folding transition in the DMA-binding domain of GCN4 on specific binding to DNA. Nature 347, 575–578 (1990). https://doi.org/10.1038/347575a0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/347575a0
This article is cited by
-
Dimerization processes for light-regulated transcription factor Photozipper visualized by high-speed atomic force microscopy
Scientific Reports (2022)
-
The transcription factor reservoir and chromatin landscape in activated plasmacytoid dendritic cells
BMC Genomic Data (2021)
-
An improved sequence based prediction protocol for DNA-binding proteins using SVM and comprehensive feature analysis
BMC Bioinformatics (2013)
-
On the segregation of protein ionic residues by charge type
Amino Acids (2012)
-
Ligand-induced conformational changes allosterically activate Toll-like receptor 9
Nature Immunology (2007)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.