Abstract
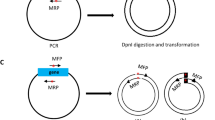
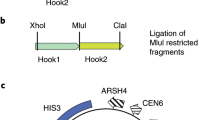
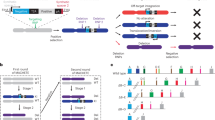
A basic difficulty in the molecular analysis of genes identified by mutations in the mammalian genome is the need to cover genetic distances corresponding to several hundred kilobases or more by molecular techniques like chromosome walking. In chromosome jumping1–3, this limitation is overcome by the deletion of all but the extreme ends of large DNA molecules before cloning. We describe here the construction and characterization of a NotI 'jumping library' from human DNA. To characterize this library, random clones were analysed by restriction mapping. Clones carrying unique end fragments were characterized further by hybridization to Southern blots of NotI-cleaved human DNA separated on pulsed field gradient (PFG) gels4–6. As a first step in a directional walk, the library was screened with a clone containing a NotI site cleaved in genomic DNA ('NotI linking clone')2,3,7 localized to the distal third of the short arm of human chromosome 4 (A.-M.F. & T.P., unpublished data). Starting and end points of two identified clones were positioned within a restriction map covering 850 kilobases.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Collins, F. S. & Weissman, S. M. Proc. natn. Acad. Sci. U.S.A. 81, 6812–6816 (1984).
Poustka, A. & Lehrach, H. Trends Genet. 2, 174–179 (1986).
Poustka, A. et al. Cold Spring Harb. Symp. quant. Biol. (in the press).
Schwartz, D. C. & Cantor, C. R. Cell 37, 67–75 (1984).
Smith, C. L. et al. Genet. Engng. 8, 45–70 (1986).
Smith, C. L. & Cantor, C. R. Meth. Enzym. (in the press).
Frischauf, A.-M., Murray, N. E. & Lehrach, H. Meth. Enzym. (in the press).
Steinmetz, M., Stephan, D., Dastoomikoo, G. R., Gibb, E. & Romaniuk, R. in Immunological Methods Vol. III, (eds Lefkovits, I. & Pernis, B.) 2–14 (Academic, 1985).
Dente, L., Solazzo, H., Baldari, C., Cesareui, G. & Cortese, R. in DNA Cloning Vol. I (ed. Glover, D. M.) 101–106 (IRL Press, 1985).
Bird, A. P. Nature 321, 209–213 (1986).
Brown, W. R. A. & Bird, A. P. Nature 322, 477–481 (1986).
Huang, H. V., Little, P. F. R. & Seed, B. in Vectors, a Survey of Molecular Cloning Vectors and their Uses (ed. Rodriguez, R.), (Butterworth, in the press).
Murray, N. E. in Lambda II (eds Hendrix, R. W., Roberts, J. W., Stahl, F. W. & Weisberg, R. A.) 395–398 (Cold Spring Harbor Laboratory, New York, 1983).
Scherer, G., Telford, J., Baldari, L. & Pirrotta, V. Devl Biol. 86, 438–447 (1981).
Casadaban, M. J. & Cohen, S. N. J. molec. Biol. 138, 179–207 (1980).
Wasmuth, J. J., Carlock, L. R., Smith, B. & Immken, L. L. Am. J. hum. Genet. 39, 397–403 (1986).
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Poustka, A., Pohl, T., Barlow, D. et al. Construction and use of human chromosome jumping libraries from NotI-digested DNA. Nature 325, 353–355 (1987). https://doi.org/10.1038/325353a0
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/325353a0
This article is cited by
-
Pig genome analysis: differential distribution of SINE and LINE sequences is less pronounced than in the human and mouse genomes
Mammalian Genome (1996)
-
Genetic and physical mapping of the fitness 1 (fit1) locus within the Fes-Hbb region of mouse Chromosome 7
Mammalian Genome (1995)
-
A Notl restriction map of the entire long arm of human chromosome 21
Nature Genetics (1993)
-
Chromosomal location and structure of amplicons in two human cell lines with coamplification of c-myc and Ki-ras oncogenes
Somatic Cell and Molecular Genetics (1993)
-
The pseudoautosomal regions of the human sex chromosomes
Human Genetics (1993)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.