Abstract
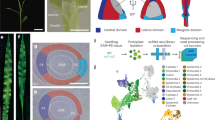
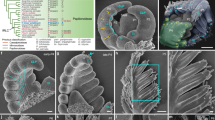
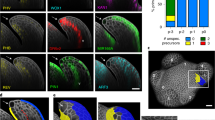
Higher plants elaborate much of their architecture post-embryonically through development initiated at the tips of shoots1,2. During vegetative growth, leaf primordia arise at predictable sites to give characteristic leaf arrangements, or phyllotaxies3,4. How these sites are determined is a long-standing question5,6 that bears on the nature of pattern-formation mechanisms in plants. Fate-mapping studies in several species indicate that each leaf primordium becomes organized from a group of 100–200 cells on the flank of the shoot apex7. Although molecular studies indicate that the regulated expression of specific homeobox genes plays some part in this determination process8,9,10,11, mechanisms that regulate the timing and position of leaf initiation are less well understood. Here we describe a gene from maize, terminal ear 1. Patterns of expression of this gene in the shoot and phenotypes of mutants indicate a role for terminal ear 1 in regulating leaf initiation. The te1 gene product contains conserved RNA-binding motifs, indicating that it may function through an RNA-binding activity.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Walbot, V. On the life strategies of plants and animals. Trends Genet. 1, 165–169 (1985).
Steeves, T. A. & Sussex, I. M. Patterns in Plant Development (Cambridge Univ. Press, Cambridge, 1989).
Sussex, I. M. Developmental programming of the shoot meristem. Cell 56, 225–229 (1989).
Smith, L. G. & Hake, S. The initiation and determination of leaves. Plant Cell 4, 1017–1027 (1992).
Church, A. H. On the Relation of Phyllotaxy to Mechanical Laws (Williams and Norgate, London, 1904).
Richards, F. J. Phyllotaxis: its quantitative expression and relation to growth in the apex. Phil. Trans. R. Soc. Lond. B. 236, 509–564 (1951).
Poethig, R. S. Clonal analysis of cell lineage patterns in plant development. Am. J. Bot. 74, 581–594 (1987).
Smith, L., Greene, B., Veit, B. & Hake, S. Adominant mutation in the maize homeobox gene, Knotted-1, causes its ectopic expression in leaf cells with altered fates. Development 116, 21–30 (1992).
Long, J. A., Moan, E. I., Medford, J. I. & Barton, M. K. Amember of the KNOTTED class of homeodomain proteins encoded by the STM gene of Arabidopsis. Nature 379, 66–69 (1996).
Scanlon, M. J., Schneeburger, R. G. & Freeling, M. The maize mutant narrow sheath fails to establish leaf margin identity in a meristematic domain. Development 122, 1683–1691 (1996).
Jackson, D., Veit, B. & Hake, S. Expression of maize KNOTTED1 related homeobox genes in the shoot apical meristem predicts patterns of morphogenesis in the vegetative shoot. Development 120, 405–413 (1994).
Matthews, D. L., Grogan, C. O. & Manchester, C. E. Terminal ear mutant of maize (Zea mays). J. Agric. Sci. 82, 433–435 (1974).
Kiesselbach, T. A. The Structure and Reproduction of Corn (Univ. Nebraska, Lincoln, 1949).
Chandler, B. L. & Hardeman, K. J. The Mu elements of Zea mays. Adv. Genet. 30, 77–122 (1992).
Watanabe, Y., Iino, Y., Furuhata, K., Shimoda, C. & Yamamoto, M. The S. pombe mei2 gene encoding a crucial molecule for commitment to meiosis is under the regulation of cAMP. EMBO J. 7, 761–767 (1988).
Birney, E., Kumar, S. & Krainer, A. R. Analysis of the RNA-recognition motif and RS and RGG domains: conservation in metazoan pre-mRNA splicing factors. Nucleic Acids Res. 21, 5803–5816 (1993).
Watanabe, Y., Shinozaki-Yabana, S., Chikashige, Y., Hiraoka, Y. & Yamamoto, M. Phosphorylation of RNA-binding protein controls cell cycle switch from mitotic to meiotic in fission yeast. Nature 386, 187–190 (1997).
McDaniel, C. & Poethig, S. Cell-lineage patterns in the shoot apical meristem of the germinating maize embryo. Planta 175, 13–22 (1988).
Doebley, J., Stec, A. & Hubbard, L. The evolution of apical dominance in maize. Nature 386, 485–488 (1997).
Itis, H. From teosinte to maize: the catastrophic sexual transmutation. Science 222, 886–894 (1983).
Burd, C. G. & Dreyfuss, G. Conserved structures and diversity of functions of RNA-binding proteins. Science 265, 615–621 (1994).
Macknight, R.et al. FCA, a gene controlling flowering time in Arabidopsis, encodes a protein containing RNA-binding domains. Cell 89, 737–745 (1997).
Watanabe, Y. & Yamamoto, M. S. pombe mei2+ encodes an RNA-binding protein required for premeiotic DNA synthesis and meiosis I, which cooperates with a novel RNA species meiRNA. Cell 78, 487–498 (1994).
Acknowledgements
This work was supported by NSF and USDA grants (to S.H.), a USDA grant (to R.J.S. and M.F.Y.) and grants from Pioneer Hi-Bred Int. and the Marsden Foundation (to B.V.). Technical assistance in cloning the te1 gene was provided by E. Robbie. te1-mum2 was a gift from P. Chomet. GenBank accession number: AF047852.
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Veit, B., Briggs, S., Schmidt, R. et al. Regulation of leaf initiation by the terminal ear 1 gene of maize. Nature 393, 166–168 (1998). https://doi.org/10.1038/30239
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/30239
This article is cited by
-
Linkage mapping combined with GWAS revealed the genetic structural relationship and candidate genes of maize flowering time-related traits
BMC Plant Biology (2022)
-
Developmental genetics of maize vegetative shoot architecture
Molecular Breeding (2021)
-
Phyllotaxis: from classical knowledge to molecular genetics
Journal of Plant Research (2021)
-
3D genome architecture coordinates trans and cis regulation of differentially expressed ear and tassel genes in maize
Genome Biology (2020)
-
An integrated analysis of QTL mapping and RNA sequencing provides further insights and promising candidates for pod number variation in rapeseed (Brassica napus L.)
BMC Genomics (2017)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.