Abstract
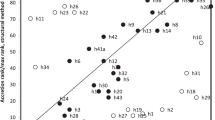
THERE is no doubt that the large ribosomal RNAs play specific roles in ribosome function1. Yet the thrust of experimentation during the past 5 years indicates clearly that the biologist tends to view these roles as structural, function being reserved by and large for the ribosomal protein components2,3. Whether or not this prejudice can be maintained remains to be seen. In any case, the existence of an approximate sequence for a 16S rRNA provides a basis for detailed characterisations of this molecule3,4. Here we begin to define 16S rRNA function by localising the major conserved regions in the molecule through a comparative analysis of 27 prokaryotic 16S rRNA primary structures.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Traub, P., and Nomura, M., Proc. natn. Acad. Sci. U.S.A., 59, 777–784 (1968).
Garrett, R. A., and Wittmann, H. G., Adv. Protein Chem., 27, 277–347 (1973).
Fellner, P., Ehresmann, C., Stiegler, P., and Ebel, J.-P., Nature new Biol., 239, 1–5 (1972)
Ehresmann, C., Stiegler, T., Mackie, G., Zimmerrmann, R. A., Ebel, J. -P., and Fellner, P., in Handbook of nucleic acid sequences, 73 (Joynson-Bruvvers, Oxford 1974).
Pechman, K. J., and Woese, C. R., J. molec. Evol., 1, 230–240 (1972).
Sogin, S. J., Sogin, M. L., and Woese, C. R., J. molec. Evol., 1, 173–184 (1972).
Uchida, T., Bonen, L., Schaup, H. W., Lewis, B. J., Zablen, L., and Woese, C. R., J. molec. Evol., 3, 63–77 (1974).
Woese, C. R., Sogin, M. L., and Sutton, L. A., J. molec. Evol., 3, 293–299 (1974).
Zablen, L., Bonen, L., Meyer, R., and Woese, C., J. molec. Evol. (in the press).
Zablen, L. thesis, Univ. Illinois (1975).
Doolittle, W. F., Woese, C. R., Sogin, M. L., Bonen, L., and Stahl, D., J. molec. Evol., (in the press).
Zablen, L., and Woese, C. R., J. molec. Evol. (in the press).
Senior, B. W., and Holland, I. B., Proc. natn. Acad. Sci. U.S.A., 68, 959–963 (1971).
Bowman, C. M., Dahlberg, J. E., Ikemura, T., Konisky, J., and Nomura, M., Proc. natn. Acad. Sci. U.S.A., 68, 964–968 (1971).
Helser, T. L., Davies, J., and Dahlberg, J. E., Nature new Biol., 235, 6–9 (1972).
Noller, H. F., Biochemistry, 13, 4694–4703 (1974).
Santer, M., and Santer, U. V., J. Bact., 116, 1304–1313 (1973).
Schaup, H. W., Sogin, M., Woese, C., and Kurland, C. G., Molec. gen. Genetics, 114, 1–8 (1971).
Zimmerman, R. A., Muto, A., Fellner, P., Ehresmann, C., and Branlant, C., Proc. natn. Acad. Sci. U.S.A., 69, 1282–1286 (1972).
Schaup, H. W., Sogin, M. L., Kurland, C. G., and Woese, C. R., J. Bact., 115, 82–87 (1973).
Muto, A., Ehresmann, C., Fellner, P., and Zimmermann, R. A., J. molec. Biol., 86, 411–432 (1974).
Székely, M., Brimacombe, R., and Morgan, J., Eur. J. Biochem., 35, 574–581 (1973).
Nomura, M., Traub, P., and Bechmann, H., Nature, 219, 793–799 (1968).
Wrede, P., and Erdmann, V. A., FEBS Letts., 33, 315–319 (1973).
Gray, P. N., Bellemare, G., Monier, R., Garrett, R. A., and Stöffler, G., J. molec. Biol., 77, 133–152 (1973).
Fox, G. E., and Woese, C. R., J. molec. Evol., 4, 000 (1974).
Fellner, P., Ehresmann, C., and Ebel, J. P., Biochimie, 54, 853–900 (1972).
Santer, M., and Santer, U. V., Biochemistry, 11, 786–791 (1972).
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
WOESE, C., Fox, G., ZABLEN, L. et al. Conservation of primary structure in 16S ribosomal RNA. Nature 254, 83–86 (1975). https://doi.org/10.1038/254083a0
Received:
Revised:
Issue Date:
DOI: https://doi.org/10.1038/254083a0
This article is cited by
-
Ultrafast and accurate 16S rRNA microbial community analysis using Kraken 2
Microbiome (2020)
-
Diversity, ecology and evolution of Archaea
Nature Microbiology (2020)
-
A highly adaptive microbiome-based association test for survival traits
BMC Genomics (2018)
-
The heat shock protein 70 gene as a new alternative molecular marker for the taxonomic identification of Streptomyces strains
AMB Express (2018)
-
A powerful microbiome-based association test and a microbial taxa discovery framework for comprehensive association mapping
Microbiome (2017)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.