Abstract
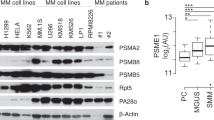
Proteasomal proteolysis relies on the activity of six catalytically active proteasomal subunits (β1, β2, β5, β1i, β2i and β5i). Applying a functional proteomics approach, we used a recently developed activity-based, cell-permeable proteasome-specific probe that for the first time allows differential visualization of individual active proteasomal subunits in intact primary cells. In primary leukemia samples, we observed remarkable variability in the amounts of active β1/1i-, β2/2i- and β5/5i-type of subunits, contrasting with their constant protein expression. Bortezomib inhibited β5- and β1-type, but to a lesser extend β2-type of subunits in live primary cells in vitro and in vivo. When we adapted the bortezomib-sensitive human acute myeloid leukemia cell line HL-60 to bortezomib 40 nM (HL-60a), proteasomal activity profiling revealed an upregulation of active subunits, and residual β1/β5-type of activity could be visualized in the presence of bortezomib 20 nM, in contrast to control cells. In a panel of cell lines from hematologic malignancies, the ratio between β2-type and (β1+β5)-type of active proteasomal polypeptides mirrored different degrees of bortezomib sensitivity. We thus conclude that the proteasomal activity profile varies in primary leukemia cells, and that the pattern of proteasomal subunit activity influences the sensitivity of hematologic malignancies toward bortezomib.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Hideshima T, Richardson P, Chauhan D, Palombella VJ, Elliott PJ, Adams J et al. The proteasome inhibitor PS-341 inhibits growth, induces apoptosis, and overcomes drug resistance in human multiple myeloma cells. Cancer Res 2001; 61: 3071–3076.
Rajkumar SV, Richardson PG, Hideshima T, Anderson KC . Proteasome inhibition as a novel therapeutic target in human cancer. J Clin Oncol 2005; 23: 630–639.
Glickman MH, Ciechanover A . The ubiquitin–proteasome proteolytic pathway: destruction for the sake of construction. Physiol Rev 2002; 82: 373–428.
Wilkinson KD . Ubiquitination and deubiquitination: targeting of proteins for degradation by the proteasome. Semin Cell Dev Biol 2000; 11: 141–148.
Kovalenko A, Chable-Bessia C, Cantarella G, Israel A, Wallach D, Courtois G . The tumour suppressor CYLD negatively regulates NF-kappaB signalling by deubiquitination. Nature 2003; 424: 801–805.
Stein RL, Melandri F, Dick L . Kinetic characterization of the chymotryptic activity of the 20S proteasome. Biochemistry 1996; 35: 3899–3908.
Lightcap ES, McCormack TA, Pien CS, Chau V, Adams J, Elliott PJ . Proteasome inhibition measurements: clinical application. Clin Chem 2000; 46: 673–683.
Dantuma NP, Lindsten K, Glas R, Jellne M, Masucci MG . Short-lived green fluorescent proteins for quantifying ubiquitin/proteasome-dependent proteolysis in living cells. Nat Biotechnol 2000; 18: 538–543.
Lindsten K, Menendez-Benito V, Masucci MG, Dantuma NP . A transgenic mouse model of the ubiquitin/proteasome system. Nat Biotechnol 2003; 21: 897–902.
Luker GD, Pica CM, Song J, Luker KE, Piwnica-Worms D . Imaging 26S proteasome activity and inhibition in living mice. Nat Med 2003; 9: 969–973.
Altun M, Galardy PJ, Shringarpure R, Hideshima T, LeBlanc R, Anderson KC et al. Effects of PS-341 on the activity and composition of proteasomes in multiple myeloma cells. Cancer Res 2005; 65: 7896–7901.
Berkers CR, Verdoes M, Lichtman E, Fiebiger E, Kessler BM, Anderson KC et al. Activity probe for in vivo profiling of the specificity of proteasome inhibitor bortezomib. Nat Methods 2005; 2: 357–362.
Greiner A, Lautwein A, Overkleeft HS, Weber E, Driessen C . Activity and subcellular distribution of cathepsins in primary human monocytes. J Leukocyte Biol 2003; 73: 235–242.
Kessler BM, Tortorella D, Altun M, Kisselev AF, Fiebiger E, Hekking BG et al. Extended peptide-based inhibitors efficiently target the proteasome and reveal overlapping specificities of the catalytic beta-subunits. Chem Biol 2001; 8: 913–929.
Lautwein A, Kraus M, Reich M, Burster T, Brandenburg J, Overkleeft HS et al. Human B lymphoblastoid cells contain distinct patterns of cathepsin activity in endocytic compartments and regulate MHC class II transport in a cathepsin S-independent manner. J Leukocyte Biol 2004; 75: 844–855.
Shevchenko A, Wilm M, Vorm O, Mann M . Mass spectrometric sequencing of proteins silver-stained polyacrylamide gels. Anal Chem 1996; 68: 850–858.
Perkins DN, Pappin DJ, Creasy DM, Cottrell JS . Probability-based protein identification by searching sequence databases using mass spectrometry data. Electrophoresis 1999; 20: 3551–3567.
Froment C, Uttenweiler-Joseph S, Bousquet-Dubouch MP, Matondo M, Borges JP, Esmenjaud C et al. A quantitative proteomic approach using two-dimensional gel electrophoresis and isotope-coded affinity tag labeling for studying human 20S proteasome heterogeneity. Proteomics 2005; 5: 2351–2363.
Glas R, Bogyo M, McMaster JS, Gaczynska M, Ploegh HL . A proteolytic system that compensates for loss of proteasome function. Nature 1998; 392: 618–622.
Gavioli R, Frisan T, Vertuani S, Bornkamm GW, Masucci MG . c-myc overexpression activates alternative pathways for intracellular proteolysis in lymphoma cells. Nat Cell Biol 2001; 3: 283–288.
Chen L, Madura K . Increased proteasome activity, ubiquitin-conjugating enzymes, and eEF1A translation factor detected in breast cancer tissue. Cancer Res 2005; 65: 5599–5606.
Adams J . Development of the proteasome inhibitor PS-341. Oncologist 2002; 7: 9–16.
Acknowledgements
This work was supported by the Deutsche Krebshilfe (project 106739), the fortune-Programm at the University of Tübingen, the Sonderforschungsbereich 685 (project B2) and the Netherlands Organization for Scientific Research (NWO) to HO.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Kraus, M., Rückrich, T., Reich, M. et al. Activity patterns of proteasome subunits reflect bortezomib sensitivity of hematologic malignancies and are variable in primary human leukemia cells. Leukemia 21, 84–92 (2007). https://doi.org/10.1038/sj.leu.2404414
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.leu.2404414
Keywords
This article is cited by
-
Negligible role of TRAIL death receptors in cell death upon endoplasmic reticulum stress in B-cell malignancies
Oncogenesis (2023)
-
Bortezomib resistance in multiple myeloma is associated with increased serine synthesis
Cancer & Metabolism (2017)
-
Proteasome subunit expression analysis and chemosensitivity in relapsed paediatric acute leukaemia patients receiving bortezomib-containing chemotherapy
Journal of Hematology & Oncology (2016)
-
Synergistic effects of proteasome inhibitor carfilzomib in combination with tyrosine kinase inhibitors in imatinib-sensitive and -resistant chronic myeloid leukemia models
Oncogenesis (2014)
-
The resistance mechanisms of proteasome inhibitor bortezomib
Biomarker Research (2013)