Abstract
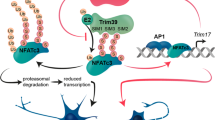
Sumoylation and ubiquitinylation reversibly regulate the activity of transcription factors through covalent attachment to lysine residues of target proteins. We examined whether the Ets-1 transcription factor is modified by sumoylation and/or ubiquitinylation. Among four potential SUMO motifs in Ets-1, we identified lysines 15 and 227 within the LK15YE and IK227QE motifs, as being the sumoylation acceptor sites. Using transfection of Ets-1 wildtype (WT) or its sumoylation deficient version (Ets-1 K15R/K227R), as well as WT or mutant proteins of the SUMO pathway, we further demonstrated that the E2 SUMO-conjugating enzyme Ubc9 and a E3 SUMO ligase, PIASy, can enhance Ets-1 sumoylation, while a SUMO protease, SENP1, can desumoylate Ets-1. We also found that Ets-1 is modified by K48-linked polyubiquitinylation independently of the sumoylation acceptor sites and is degraded through the 26S proteasome pathway, while sumoylation of Ets-1 does not affect its stability. Finally, sumoylation of Ets-1 leads to reduced transactivation and we demonstrated that previously identified critical lysine residues in Synergistic Control motifs are the sumoylation acceptor sites of Ets-1. These data show that Ets-1 can be modified by sumoylation and/or ubiquitinylation, with sumoylation repressing transcriptional activity of Ets-1 and having no clear antagonistic action on the ubiquitin-proteasome degradation pathway.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Bohren KM, Nadkarni V, Song JH, Gabbay KH, Owerbach D . (2004). J Biol Chem 279: 27233–27238.
Bories JC, Willerford DM, Grevin D, Davidson L, Camus A, Martin P et al. (1995). Nature 377: 635–638.
Chakrabarti SR, Sood R, Nandi S, Nucifora G . (2000). Proc Natl Acad Sci USA 97: 13281–13285.
Cheng J, Wang D, Wang Z, Yeh ET . (2004). Mol Cell Biol 24: 6021–6028.
Cowley DO, Graves BJ . (2000). Genes Dev 14: 366–376.
Crepieux P, Coll J, Stehelin D . (1994). Crit Rev Oncogol 5: 615–638.
de Launoit Y, Audette M, Pelczar H, Plaza S, Baert JL . (1998). Oncogene 16: 2065–2073.
Degerny C, Monte D, Beaudoin C, Jaffray E, Portois L, Hay RT et al. (2005). J Biol Chem 280: 24330–24338.
Desterro JM, Rodriguez MS, Hay RT . (1998). Mol cell 2: 233–239.
Desterro JM, Thomson J, Hay RT . (1997). FEBS lett 417: 297–300.
Dittmer J . (2003). Mol Cancer 2: 29.
Dohmen RJ . (2004). Biochim Biophys Acta 1695: 113–131.
Donaldson LW, Petersen JM, Graves BJ, McIntosh LP . (1996). EMBO J 15: 125–134.
Fafeur V, Tulasne D, Queva C, Vercamer C, Dimster V, Mattot V et al. (1997). Cell Growth Differ 8: 655–665.
Haglund K, Dikic I . (2005). EMBO J 24: 3353–3359.
Hahn SL, Wasylyk B, Criqui-Filipe P . (1997). Oncogene 15: 1489–1495.
Hay RT . (2005). Mol Cell 18: 1–12.
Herrera FJ, Triezenberg SJ . (2004). Curr Biol 14: R622–R624.
Hershko A, Ciechanover A . (1998). Annu Rev Biochem 67: 425–479.
Hoege C, Pfander B, Moldovan GL, Pyrowolakis G, Jentsch S . (2002). Nature 419: 135–141.
Iniguez-Lluhi JA, Pearce D . (2000). Mol Cell Biol 20: 6040–6050.
Janknecht R, Nordheim A . (1996). Oncogene 12: 1961–1969.
Johnson ES, Blobel G . (1997). J Biol Chem 272: 26799–26802.
Johnson ES . (2004). Annu Rev Biochem 73: 355–382.
Kurihara I, Shibata H, Kobayashi S, Suda N, Ikeda Y, Yokota K et al. (2005). J Biol Chem 280: 6721–6730.
Leight ER, Glossip D, Kornfeld K . (2005). Development 132: 1047–1056.
Macauley MS, Errington WJ, Scharpf M, Mackereth CD, Blaszczak AG, Graves BJ et al. (2005). J Biol Chem 281: 4164–4172.
Matunis MJ, Coutavas E, Blobel G . (1996). J Cell Biol 135: 1457–1470.
Muller S, Ledl A, Schmidt D . (2004). Oncogene 23: 1998–2008.
Muthusamy N, Barton K, Leiden JM . (1995). Nature 377: 639–642.
Oikawa T, Yamada T . (2003). Gene 303: 11–34.
Paumelle R, Tulasne D, Kherrouche Z, Plaza S, Leroy C, Reveneau S et al. (2002). Oncogene 21: 2309–2319.
Perdomo J, Verger A, Turner J, Crossley M . (2005). Mol Cell Biol 25: 1549–1559.
Rabault B, Ghysdael J . (1994). J Biol Chem 269: 28143–28151.
Rabault B, Roussel MF, Quang CT, Ghysdael J . (1996). Oncogene 13: 877–881.
Reveneau S, Paumelle R, Deheuninck J, Leroy C, De Launoit Y, Fafeur V . (2003). Ann NY Acad Sci 1010: 100–103.
Rodriguez MS, Dargemont C, Hay RT . (2001). J Biol Chem 276: 12654–12659.
Saitoh H, Hinchey J . (2000). J Biol Chem 275: 6252–6258.
Sampson DA, Wang M, Matunis MJ . (2001). J Biol Chem 276: 21664–21669.
Schmidt D, Muller S . (2003). Cell Mol Life Sci 60: 2561–2574.
Seeler JS, Dejean A . (2003). Mol Cell Biol 4: 690–699.
Slupsky CM, Gentile LN, Donaldson LW, Mackereth CD, Seidel JJ, Graves BJ et al. (1998). Proc Natl Acad Sci USA 95: 12129–12134.
Takahashi A, Higashino F, Aoyagi M, Yoshida K, Itoh M, Kobayashi M et al. (2005). Biochem Biophys Res Commun 327: 575–580.
Tulasne D, Deheuninck J, Lourenco FC, Lamballe F, Ji Z, Leroy C et al. (2004). Mol Cell Biol 24: 10328–10339.
Tulasne D, Paumelle R, Weidner KM, Vandenbunder B, Fafeur V . (1999). Mol Biol Cell 10: 551–565.
van den Akker E, Ano S, Shih HM, Wang LC, Pironin M, Palvimo JJ et al. (2005). J Biol Chem 280: 38035–38046.
Wasylyk C, Bradford AP, Gutierrez-Hartmann A, Wasylyk B . (1997). Oncogene 14: 899–913.
Wasylyk C, Criqui-Filipe P, Wasylyk B . (2005). Oncogene 24: 820–828.
Wernert N, Gilles F, Fafeur V, Bouali F, Raes MB, Pyke C et al. (1994). Cancer Res 54: 5683–5688.
Wernert N, Raes MB, Lassalle P, Dehouck MP, Gosselin B, Vandenbunder B et al. (1992). Am J Pathol 140: 119–127.
Xiao GH, Jeffers M, Bellacosa A, Mitsuuchi Y, Vande Woude GF, Testa JR . (2001). Proc Natl Acad Sci USA 98: 247–252.
Yang SH, Jaffray E, Hay RT, Sharrocks AD . (2003). Mol cell 12: 63–74.
Yang SH, Sharrocks AD . (2005). EMBO J 24: 2161–2171.
Acknowledgements
This work was supported by CNRS, Institut Pasteur de Lille, University of Lille 1, University of Lille 2, and INSERM, and by grants from the Fondation de France, the FEDER-Région Nord Pas de Calais and the Ligue Contre le Cancer-Comité Nord. ZJ was supported by a Fondation de France fellowship, NV by a Ligue Nationale Contre le Cancer fellowship, BF by a Institut Pasteur/Région Nord- Pas de Calais fellowship and JD by Association pour la Recherche sur le Cancer fellowship.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Ji, Z., Degerny, C., Vintonenko, N. et al. Regulation of the Ets-1 transcription factor by sumoylation and ubiquitinylation. Oncogene 26, 395–406 (2007). https://doi.org/10.1038/sj.onc.1209789
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.onc.1209789
Keywords
This article is cited by
-
A deficiency in SUMOylation activity disrupts multiple pathways leading to neural tube and heart defects in Xenopus embryos
BMC Genomics (2019)
-
Usp9x regulates Ets-1 ubiquitination and stability to control NRAS expression and tumorigenicity in melanoma
Nature Communications (2017)
-
A genetic screen to discover SUMOylated proteins in living mammalian cells
Scientific Reports (2017)
-
An improved ChIP-seq peak detection system for simultaneously identifying post-translational modified transcription factors by combinatorial fusion, using SUMOylation as an example
BMC Genomics (2014)
-
Review of Ets1 structure, function, and roles in immunity
Cellular and Molecular Life Sciences (2013)