Abstract
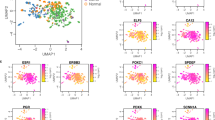
Comparison of gene expression changes between cancer cells at the periphery and in the centre of breast cancers was performed using a combination of microdissection and microarray analysis. Cancer cells from the two areas were pooled separately from five patients with ductal carcinoma in situ and separately from five patients with frankly invasive cancer. Limited total RNA, 100–200 ng, from this microdissected tissue required use of the Atlas SMART™ Probe Amplification Kit to synthesize and amplify cDNA and make 33P-labelled probes. Probes were then hybridized to Atlas Human Cancer 1.2 Arrays containing 1176 known genes. Triplicate analysis revealed that 22 genes changed their expression levels in the periphery relative to the central region: 15 upregulated and seven downregulated (arbitrary threshold of 1.5-fold or greater). Differences in RNA levels were confirmed by quantitative real-time PCR for two of the genes and by changes in protein levels, detected by immunohistochemistry, for a couple of representative gene products. Thus, changes in gene expression associated with variation in microanatomical location of neoplastic cells can be detected within even small developing tumour masses.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
Abbreviations
- CK 19:
-
cytokeratin 19
- DCIS:
-
ductal carcinoma in situ
- EF-1 α:
-
elongation factor 1 alpha
- TSG101:
-
tumour susceptibility gene 101
References
Assmann V, Gillett CE, Poulsom R, Ryder K, Hart IR and Hanby AM . (2001). J. Pathol., 195, 191–196.
Chang C and Werb Z . (2001). Trends Cell Biol., 11, S37–S43.
Condeelis J . (1995). Trends Biochem. Sci., 20, 169–170.
DeRisi J, Penland L, Brown PO, Bittner ML, Meltzer PS, Ray M, Chen Y, Su YA and Trent JM . (1996). Nat. Genet., 14, 457–460.
Edmonds BT, Wyckoff J, Yeung YG, Wang Y, Stanley ER, Jones J, Segall J and Condeelis J . (1996). J. Cell Sci., 109, 2705–2714.
Gehlsen KR, Argraves WS, Pierschbacher MD and Ruoslahti E . (1988). J. Cell Biol., 106, 925–930.
Hall PA, Coates PJ, Goodlad RA, Hart IR and Lane DP . (1994). J. Cancer, 70, 244–247.
Li L and Cohen SN . (1996). Cell, 85, 319–329.
Luzzi V, Holtschlag V and Watson MA . (2001). Am. J. Pathol., 158, 2005–2010.
Mariani L, McDonough WS, Hoelzinger DB, Beaudry C, Kaczmarek E, Coons SW, Giese A, Moghaddam M, Seiler RW and Berens ME . (2001). Cancer Res., 61, 4190–4196.
Marshall JF and Hart IR . (1996). Semin. Cancer Biol., 7, 129–138.
Meyer T and Hart IR . (1998). Eur. J. Cancer, 34, 214–221.
Nocito A, Bubendorf L, Maria Tinner E, Suess K, Wagner U, Forster T, Kononen J, Fijan A, Bruderer J, Schmid U, Ackermann D, Maurer R, Alund G, Knonagel H, Rist M, Anabitarte M, Hering F, Hardmeier T, Schoenenberger AJ, Flury R, Jager P, Luc Fehr J., Schraml P, Moch H, Mihatsch MJ, Gasser T and Sauter G . (2001). J. Pathol., 194, 349–357.
Parker SL, Tong T, Bolden S and Wingo PA . (1997). Cancer J. Clin., 47, 5–27.
Paweletz CP, Charboneu L, Bichsel VE, Simone NL, Chen T, Gillespie JW, Emmert-Buck MR, Roth MJ, Petricoin III EF and Liotta LA . (2001). Oncogene, 20, 1981–1989.
Perou CM, Jeffrey SS, van de Rijn M, Rees CA, Eisen MB, Ross DT, Pergamenschikov A, Williams CF, Zhu SX, Lee JC, Lashkari D, Shalon D, Brown PO and Botstein D . (1999). Proc. Natl. Acad. Sci. USA, 96, 9212–9217.
Saha S, Bardelli A, Buckhaults P, Velculescu VE, Rago C, Croix BS, Romans KE, Choti MA, Lengauer C, Kinzler KW and Vogelstein B . (2001). Science, 294, 1343–1346.
Scheidl SJ, Nilsson S, Kalen M, m Hellstrom M, Takemoto M, Hakansson J and Lindahl P . (2002). Am. J. Pathol., 160, 801–813.
Schena M, Shalon D, Davis RW and Brown PO . (1995). Science, 270, 467–470.
Schutze K, Posl H and Lahr G . (1998). Cell Mol. Biol., 44, 735–746.
Suwa H, Ohshio G, Imamura T, Watanabe G, Arii S, Imamura M, Narumiya S, Hiai H and Fukumoto M . (1998). Br. J. Cancer, 77, 147–152.
Talbot DC and Brown PD . (1996). Eur. J. Cancer, 32A, 2528–2533.
Van Trappen PO, Gyselman VG, Lowe DG, Ryan A, Oram DH, Bosze P, Weekes AR, Shepherd JH, Dorudi S, Bustin SA and Jacobs IJ . (2001). Lancet, 357, 15–20.
Zhumabayeva B, Diatchenko L, Chenchik A and Siebert PD . (2001). Biotechniques, 30, 158–163.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Zhu, G., Reynolds, L., Crnogorac-Jurcevic, T. et al. Combination of microdissection and microarray analysis to identify gene expression changes between differentially located tumour cells in breast cancer. Oncogene 22, 3742–3748 (2003). https://doi.org/10.1038/sj.onc.1206428
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.onc.1206428
Keywords
This article is cited by
-
Discovery of Missing Methylation Sites on Endogenous Peptides of Human Cell Lines
Journal of the American Society for Mass Spectrometry (2019)
-
Identification of unique expression signatures and therapeutic targets in esophageal squamous cell carcinoma
BMC Research Notes (2012)
-
An upstream regulatory element confers orientation-independent enhancement of the TSG101 promoter activity in transformed cells
Molecular Biology Reports (2012)
-
A small-molecule inhibitor shows that pirin regulates migration of melanoma cells
Nature Chemical Biology (2010)
-
DNA methylation of polycomb group target genes in cores taken from breast cancer centre and periphery
Breast Cancer Research and Treatment (2010)