Abstract
The X-linked lymphoproliferative syndrome (XLP) is an inherited immunodeficiency to Epstein-Barr virus infection that has been mapped to chromosome Xq25. Molecular analysis of XLP patients from ten different families identified a small interstitial constitutional deletion in 1 patient (XLP-D). This deletion, initially defined by a single marker, DF83, known to map to interval Xq24–q26.1, is nested within a previously reported and much larger deletion in another XLP patient (XLP-739). A cosmid minilibrary was constructed from a single mega-YAC and used to establish a contig encompassing the whole XLP-D deletion and a portion of the XLP-739 deletion. Based on this contig, the size of the XLP-D deletion can be estimated at 130 kb. The identification of this minimal deletion, within which at least a portion of the XLP gene is likely to reside, should greatly facilitate efforts in isolating the gene.
Similar content being viewed by others
Introduction
X-linked lymphoproliferative syndrome (XLP; MIM 308240), originally called Duncan disease [1, 2], is a rare primary immunodeficiency characterized by selective susceptibility to Epstein-Barr virus (EBV). EBV infection of affected boys results in severe or fatal infectious mononucleosis, with survivors developing hypo- or agammaglobulinemia [1]. Malignant B cell non-Hodgkin lymphoma arises in more than 25% of the survivors [3], with the disease usually fatal by age 40 [4]. Males carrying a defective XLP gene are healthy until they become infected with EBV, suggesting that their immunodeficiency is specific to this virus [5]. EBV is an ubiquitous herpesvirus, with more than 98% of the adult population showing immunological evidence of past infection. In addition to self-limited infectious mononucleosis, EBV has been directly implicated in the development of Burkitt lymphoma and nasopharyngeal carcinoma [6]. EBV is also associated with lymphoproliferative syndromes in individuals with genetic or acquired immunodeficiency, including patients with AIDS or organ transplant recipients who have undergone immunosuppressive regimens [7]. The XLP syndrome, therefore, provides a genetic model for the interaction between an infectious agent and a congenital immunodeficiency, and the isolation of the responsible gene should provide insights into the normal control of virally induced lymphoproliferation.
The XLP locus, named LYP [8], has been assigned by linkage studies to the interval Xq24–q26.1, proximal to HPRT [9]. This locus shows 1–2% recombination with DXS42 and 0% with DXS37, both of which are centromeric to it [9–12]. The telomeric border of the XLP locus is defined by DXS100. Multipoint analysis of this region has suggested a genetic map with the following order of markers [10, 11]: cen − DXS11 − 10 cM − DXS37 − 2 cM − DXS42 − DXS739 − DXS100 − 7 cM − HPRT − tel. This order is consistent with the physical map reported by Wu et al. [13]: cen − DXS42, DXS12 − (2,140 kb) − DXS6 − (700 kb) − DXS982 − (340 kb) − DXS739 − (1,120 kb) − DXS75 − (900 kb) − DXS100 − (1,760 kb) − DXS10 − DXS177 − tel. Cytogenetic studies of individuals from XLP families led to the identification of a male with a large constitutional deletion in the Xq25 region [14]. Genomic sequences corresponding to DXS6, DXS739, and DXS100 were absent in this patient. Two additional patients who had deletions of markers in the Xq25 region were also reported [15, 16]: 1 had deletions of DXS739 and DXS100 and the 2nd a deletion of DXS739. In both cases, high-resolution cytogenetics of lymphoblastoid cell lines revealed cytogenetically visible deletions, estimated at 6,000 and 2,000–3,000 kb, respectively. Nonetheless, the large size of this candidate region has complicated positional cloning efforts. Here, we report a novel constitutional deletion of only 130 kb identified in an XLP patient. This new minimal deletion is spanned by a cosmid contig which should greatly facilitate efforts at cloning the XLP gene.
Patients and Methods
Patients
The patients XLP-D, XLP-M, XLP-S, and XLP-H have been described elsewhere [17]. Patients LB8001, LB8002, and LB8005 are previously undescribed XLP patients, while individuals XLP-192, XLP-724, XLP-739, and XLP-978 are XLP patients studied in our laboratory:
XLP-192 (corresponding to the IARC192 cell line) was the 3rd son of a French family in which 2 boys had died from lymphoproliferative disease after EBV infection. At the age of 2, he developed fatal infectious mononucleosis with strong lymphoproliferative syndrome. There was serological evidence of EBV infection in this child.
XLP-724 (corresponding to the IARC724 cell line) was a 15-year-old Italian boy who presented with agammaglobulinemia, with no proof of EBV infection. Two of his brothers died at 1 year of age from lymphoproliferative disease.
XLP-739 (corresponding to the IARC739 cell line) was a 3-year-old Italian boy who died from lymphoproliferative disease after EBV infection.
XLP-978 (corresponding to the IARC978 cell line) was a 2-year-old French boy with typical XLP symptoms, including polyclonal B cell lymphoproliferative disease, hypogammaglobulinemia, and hepatomegaly with abnormal liver functions tests. Immunofluorescence detected that the vast majority of blood cells expressed EBNA (nuclear antigen) antibodies. The boy died 3 weeks after his symptoms appeared.
Markers Used
The markers used for physical mapping are described in table 1 [see also refs. 12, 13, 18–20]. DF83, PF12, and CF1a were isolated from a library of human brain cDNAs after screening with sequences generated by laser microdissection of the terminal portion of the human Xq. These markers were assigned to Xq24–q26.1 [19]. They are considered to be untranslated cDNA and have several stop codons in all reading frames [19].
For Southern blot analysis, a DF83 subfragment of 400 bp was obtained by polymerase chain reaction (PCR) using the following primers under standard conditions:
-
DF83a: AGGAATGCACCTACATATGTGGCA
-
DF83b: CAGACTGCTTATTTATGGAGATCC
Amplification of probes 1D10T3 and 1D10T7 was performed under standard conditions using the following primers:
-
For 1D10T3:
-
1D10T3s: 5′-TTCAAATATAAATCACCTAATAA-3′
-
1D10T3as: 5′-CCAATTGTCTCCTTTTCCAAA-3′
-
For 1D10T7:
-
1D10T7s: 5′-GCAAGGGAATCTATGGAACTG-3′
-
1D10T7as: 5′-GGTGCAACTAATTAGTTGGGC-3′
H3-4 (DXS739) is a 1.5-kb HindIII fragment from phage bA6. [Nelson, personal commun.; 13]. The preparation of riboprobes used for cosmid walking is described in the section Cosmid Isolation and Construction of Contigs.
Screening of the YAC (Yeast Artificial Chromosome) Library
YAC 916D1 was isolated by PCR screening of pooled DNAs of the CEPH mega-YAC library using oligos from X2-183 (DXS982). YAC yWXD592 was obtained from the Center for Genetics in Medicine (St. Louis, Mo., USA) and was localized within large XLP deletions previously described [14]. Yeast cell culture and DNA isolation were performed as described [21].
PCR Conditions and Primers. PCR was performed in 50 µl of the reaction mixture for 40 cycles using a one-step cycle program of 94°C for 4 min, 55 for 2, and 72°C for 2 min and an additional extension of 10 min at 72°C. The X2-183 primers (GDB ID: G00-216-816) used were:
-
TA12T7:
-
5′-CCGTCACATATAGAGATAGTATTACTGAACC-3′
-
TA12SP6:
-
5′-GGTCATTATGAATGGTCAAAGGAATTTGACTGC-3′
The amplified products (1.1 kb) were electrophoresed in 1.1% agarose.
YAC 916D1 Cosmid Minilibrary and Gridded X Chromosome Library
For construction of cosmid libraries from YAC 916D1 in the supercos vector, the following procedure was used: 2 µg of genomic DNA from a yeast strain carrying YAC 916D1 was prepared in agarose blocks, partially digested with 0.1 U of MboI (New England Biolabs, Beverly, Mass., USA), and dephosphorylated with 0.1 U calf intestinal phosphatase (New Englands Biolabs). Yeast DNA was then ligated into the BAMHI site of Supercos vector (Stratagene, La Jolla, Calif., USA), previously linearized with XbaI and dephosphorylated, and packaged (Stratagene). XLI Blue MR cells (Stratagene) grown in 0.2% maltose and 10 mM MgSO4 were infected with packaged DNA and selected for growth in ampicillin. Colonies containing human-derived genomic DNA were identified by hybridization to 32P-labeled Cot-1 DNA and arrayed in 96-well plates.
The ICRF-gridded X chromosome flow-sorted cosmid library is a reference library in Lawrist 4 vector [22]. The sources of cosmids used for physical mapping are listed below.
Cosmids from ICRF were (the official reference name is in parentheses):
-
1818 (ICRFc104E1818), 64 (ICRFc104J201),
-
E079 (ICRFc104E079), B0920 (ICRFc104B0920),
-
P1213 (ICRFc104P1213Q), L0129 (ICRFc104L0129),
-
L2015 (ICRFc104L015QD), N1335 (ICRFc104N1335),
-
D1785 (ICRFcl04N1785), D8112 (ICRFc104D8112),
-
4 (ICRFc104D196), 31 (ICRFc104D1912), and
-
100 (ICRFc104H07100).
Cosmids from the YAC 916D1 minilibrary were: 1D10, E8, C8, 1N5, 1U8, A8, 1Z4, F11, 1X6, 1J11, A12, and 1H11.
Cosmid Isolation and Construction of Contigs
DNAs from individual cosmids of the YAC 916D1 cosmid library were prepared by classical alkaline lysis DNA extraction [23] and spotted onto nylon filters by manual dot blot. Dot blot filters were hybridized with probes mapped in the Xq25 region. ICRF-gridded X chromosome cosmid filters were hybridized with the probes as described by Nizetic et al. [22]. DNA was extracted from hybridized clones by alkaline lysis as previously described [23]. Cosmid clones were used as a starting point for chromosome walking.
For walking, terminal-end riboprobes of cosmid and P1 clones were synthesized and hybridized to cosmid dot blots. Positive clones were selected, and the digested DNA from these clones was compared with that of flanking cosmids, blotted, and probed with the riboprobe originally used to select the cosmid from dot blots.
Riboprobes were synthesized as follows: 3 µg of cosmid DNA was digested by RsaI (Promega, Madison, Wisc., USA), extracted with phenol-chloroform, and ethanol precipitated. The reactions were performed in 25 µl final volume with 1 mM nucleotide (ACG), 30 mM dithiothreitol, and 5 µCi 32P-UTP using 10 U of RNA polymerase at 37 ° C for 30 min. T3 and T7 RNA polymerases were used for YAC 916D1 cosmid library (in supercos vector), SP6, and T7 RNA pol. For ICRF cosmid library (in Lawrist 4) 10 U of RNAse-free DNAse was then added to the reaction mixture and incubated for 15 min at 37°C. Finally, riboprobes were phenol-chloroform extracted and ethanol precipitated before hybridization. This process of terminal-end riboprobes and filter hybridization of cosmid DNAs was repeated at each stage of the walk.
Cosmid overlaps were verified by the DNA fingerprinting. Cosmid DNAs were HinfI or TaqI digested, blotted, and probed with total human placental DNA. Cosmids were subcloned into pBluescript (Stratagene) using standard techniques [23].
Restriction Map
A restriction map with BamHI was generated by partial digestion of cosmid DNA and subsequently confirmed by complete digestion, while the map of a rare cutter (ClaI) was generated only by complete digestion. For the partial digestion approach, cosmids cloned in Lawrist 4 vector were linearized with EagI (position 4272 of the vector). Partially digested DNA fragments were visualized by hybridization with two end probes, Law22 (nucleotides 5027–5281 of the vector) and Law33 (nucleotides 3870–4212), respectively, and amplified from the vector by PCR in standard conditions. The primer sequences for amplifying these two probes were:
-
Law22a: TTCTGAGCGGGACTCTGG
-
Law22b: TGCACTGCCGGTAGAACTC
-
Law33a: CAAGCTAGCTTCACGCTGC
-
Law33b: TGATCAGATCTTGATCCCCTG
Cosmid clones in Supercos vector were linearized with BssHII (position 4256 of the vector). The two end probes were Sup 43-290 (nucleotides 4395–4684 of the vector) and Sup 38-270 (nucleotides 3899–4168). Primer sequences were:
-
Sup 43-290a: CGTTGGCTACCCGTGATATT
-
Sup 43-290b: ACTCCAGCATGAGATCCCC
-
Sup 38-270b: CTGAATGAACTGCAGGACGA
-
Sup 38-270b: AGTACGTGCTCGCTCGATG
Southern Blot Analysis
Isolation of high-molecular-weight DNA, restriction endonuclease digestion, DNA separation and blotting, and dotting of DNA fragments were performed as described previously [23].
Cosmid inserts were hybridized in the presence of excess human competitor DNA as previously described [24]. The filters were hybridized in Church solution − 0.5 M phosphate buffer, pH 7.2, 7% sodium dodecyl sulfate (SDS), 1% bovine serum albumin — at 65°C and then washed once stringently in 2 × SSC and 0.1% SDS at 65°C before washing in 0.1 × SSC and 0.1% SDS at 65°C (1 × SSC = 0.15 M NaCl, 1.5 M sodium citrate).
DNA probes were 32P-labeled by random priming (Amersham) and used at a specific activity of 1 million cpm/ml of hybridization solution. Riboprobes were hybridized under the same conditions in Church buffer.
Results
Identification of an XLP Patient Carrying a Small Deletion
To define a minimal interval containing the XLP gene, EBV-immortalized cell lines were generated from 10 unrelated male patients. These were screened for hemizygous deletion or rearrangement of Xq25 markers (table 2). Markers known to be deleted in previous patients [14–16], X2-183 (DXS982), H3-4 (DXS739), and p45-h (DXS100), were tested. In addition, three new markers, recently mapped to Xq25-26, CF1a, DF83, and PF12 [19], were found to be deleted in patient XLP-739 who has a large deletion, spanning at least 2,000 kb, and, therefore, localized in the XLP region locus. Screening patients by both Southern blot [25] and/or PCR amplification identified a patient, XLP-D, lacking only one marker: DF83 (fig. 1). This patient was, therefore, presumed to carry a small interstitial deletion that might define the smallest region of overlap with the deletion of patient XLP-739.
Southern blot analysis of EcoRI-digested genomic DNAs from XLP patients. DNAs from XLP patients were digested with EcoRI and analyzed by Southern blot hybridization. A p86-RP1, and EcoRI subfragment of cosmid ckT1-1286 [25] which had been isolated from a cosmid library prepared from the X3000 hybrid cell line, was used. ckT1-1286 was mapped to Xq25; however, no deletion was seen when this cosmid was used as a probe on DNA from XLP patients. B The DF83 PCR subfragment, obtained using primers described in Patients and Methods, was used. 1202 is a lymphoblastoid cell line containing four X chromosomes per cell, obtained from NIGMS Human Genetic Mutant Cell Repository, Institute for Medical Research (Camden, N.J., USA). X3000 is a human hamster cell hybrid containing human Xq25-qter, obtained from D. Nelson (Houston, Tex., USA). The sizes of the bands are indicated on the right.
Isolation of YAC Clones from the Deleted Region
The CEPH megayac library was screened with probes X2-183 (DXS982) and DF83. The YAC 916D1 (1.74 Mbp) was found to contain X2-183 and DF83. Another YAC previously identified in the Xq25 region (yWXD592) also contained the same two markers. YAC916D1 was characterized and compared with yWXD592 using additional markers — PF12, H3-4 (DXS739), CFla, and p45h (DXS100) — with the following results: (1) YAC 916D1 was found to contain probes PF12, CFla, and H3-4 (DXS739), but not p45h (DXS100), and (2) YAC yWXD592 was found to be negative for these four markers.
The pattern of overlap among these markers indicates that DF83, X2-183 (DXS982), and PF12 lie relatively close to each other. YAC 916D1 and yWXD592 contain both DF83 which is deleted in patient XLP-D and X2-183 which is not deleted in this patient, indicating that these YACs contain at least one breaking point of the XLP-D deletion. Since DXS982 maps centromeric to DXS739 [13], the minimal deletion of XLP-D lies centromeric to DXS982 and DXS739 (fig. 2).
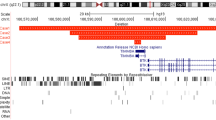
Physical map and cosmid contigs encompassing the XLP-D deletion in Xq25. A The large horizontal line indicates the relative position of markers in a normal individual. For description of markers, see Patients and Methods. B The two horizontal lines represent YAC916D1 and YAC yWXD592, respectively, localized according to the probes they contain. Note that the location of the two extremities of YAC 916D1 and the centromeric end of YAC yWXD592 are not precisely known. C The size of the cosmids and lengths of the overlapping regions are represented to scale. The size of YAC is not represented to scale because of the gap in the cosmid contig. 1818, 64, E079, B0920, P1213, L0129, L2015, N1335, D1785, D8112, 4, 31, and 100 are cosmids from the ICRF X-Chromosome Library (see Patients and Methods for official reference names). Other cosmids come from the YAC 916D1 minilibrary. D Patient XLP-D. E Patient XLP-739. Deletions are represented by heavy dotted lines. The extent of the XLP-D deletion is given to scale. The orientation of the cosmid contig encompassing the XLP-D deletion is unknown.
Southern blot analysis of HindIII-digested genomic DNA from a normal individual, patient XLP-739, and from patient XLP-D. When a 7-kb HindIII fragment from 1D10 was used as probe, it revealed a 7-kb HindIII fragment in normal controls and a 10-kb fragment in patient XLP-D. This probe is deleted in patient XLP-739.
Construction of Cosmid Contig
A cosmid library was generated from YAC 916D1 which appeared to span the XLP-D deletion. Whole-yeast DNA was partially digested with Sau3A and cloned into the SuperCos vector (Stratagene). Recombinants were screened for the presence of human repetitive sequences (Cotl), and 400 cosmids were arrayed, representing an estimated threefold coverage of YAC 916D1. Additional clones were obtained from the Imperial Cancer Research Fund gridded human X chromosome cosmid library [22]. Four cosmids positive for DF83, one cosmid containing X2-183 (DXS982), two cosmids containing PF12, and one cosmid containing H3-4 (DXS739) were isolated and used as a starting point for chromosome walking. End probes from the T3 or T7 ends of cosmids were generated either as riboprobes or by sequencing followed by PCR amplification of end fragments. At each step of the chromosome walk, cosmids fragments were tested by Southern blotting to confirm their presence or absence within the deleted region.
Four initial contigs around markers DF83, X2-183 (DXS982), PF12, and H3-4 (DXS739) were linked to form a contig with a gap in it partially encompassing the deletion of patient XLP-739 and completely covering the deletion of patient XLP-D. Since cosmids from the contig linking DXS739 and DXS982 do not contain probes deleted in patient XLP-D, we can localize the XLP-D deletion centromeric to DXS982. A gap was not filled in between the XLP-D deletion and marker X2-183 (DXS982; fig. 2), even after several screening efforts of the X chromosome cosmid library. The size of this gap remains, therefore, unknown. Overlap between cosmids in the contig was verified by DNA fingerprinting using repetitive sequence probes. A restriction map based on one frequent-cutter enzyme (BamHI) and one rare-cutter enzyme (ClaI) was constructed for the centromeric high-density part of the contig covering the smaller deletion (figure 3). This restriction map makes it possible to estimate precisely the length of overlap between different cosmids.
Characterization of Breaking Points
To characterize the two breaking points of the XLP-D deletion, we hybridized EcoRI and BamHI subfragments of three cosmids containing deleted and nondeleted sequences (1D10, E079, and B0920) on genomic DNA from patient XLP-D and normal individuals digested with several restriction enzymes (EcoRI, HindIII-BamHI, Pst1).
By Southern blot analysis, a nondeleted probe from cosmid ID 10 identified in patient XLP-D a larger fragment with respect to normal controls after HindIII digestion (fig. 4). Sixty normal individuals were tested to exclude the possibility of polymorphisms. The breaking point was finally localized in a 7-kb HindIII-BamHI fragment of 1D10. The other breaking point was mapped in a 10-kb region covered by cosmids E079 and B0920 and delimited by probes JL17 (deleted in XLP-D) and B0920.7 (nondeleted in XLP-D; see figure 4). It was not possible to identify in this 10-kb region, which is rich in repetitive sequences, a single-copy probe which could be used to localize more precisely the breaking point. The restriction map of this region (not shown) was found to be identical in both cosmids, excluding the possibility of chimerism. Based on the data just summarized, the length of the XLP-D deletion was estimated to be approximately 130 kb.
Discussion
We identified a new XLP patient harboring the smallest deletion in the XLP region reported so far (estimated at approximately 130 kb). A cosmid contig covering 500 kb of genomic DNA and encompassing completely this new deletion was constructed. A physical linkage between several markers in the region was obtained, and new markers were localized precisely with respect to previously known markers in Xq25. In particular, markers that had been assigned to Xq24–q26.1 (DF83, PF12 and CF1a) by use of somatic cell hybrid panels [19] were physically linked in a 1.5-Mbp region of Xq25 contained in YAC 916D1. Moreover, DF83 was mapped as centromeric to PF12 [19]. The position of CF1a with respect to the contig is unknown, as no other YAC, with the exception of YAC 916D1, nor cosmid was found to contain this probe.
The physical map presented here adds useful information about polymorphic markers of this region. DXS982 is physically linked to the XLP-D deletion as indicated by YAC yWXD592. DXS739 and PF12 are very close (50 kb), and the two markers DXS982 and DXS739 are physically linked. As DXS982 is centromeric to DXS739, the following sequence of markers can be drawn:
-
cen − del XLP-D − DXS982 − PF12 − DXS739 − tel.
In their high-resolution map of the region, Wu et al. [13] studied the relative position of a set of markers from Xq24–q26.1 using fluorescence in situ hybridization. These authors estimated the distance between DXS982 and DXS739 at 340 kb with an uncertainty of 70 kb. This estimate is in agreement with the distance of 300 kb we obtained using our cosmid contig.
A gap, covered by YACs yWXD592 and 916D1, is not represented in our cosmids. Efforts to isolate further cosmids between cosmids P1213 and L0129 were unsuccessful. Because of this gap, we are unable to give an orientation towards the centromeric cosmid contig covering the XLP-D deletion.
Because of the limited number and the small size of XLP families, it was very difficult to restrict the critical XLP region by genetic analysis, and the XLP gene could be localized only in a 2-cM interval between DXS42 and DXS100. The identification of XLP patients harboring interstitial deletions represented an important step in reducing this interval. Studies using fluorescence in situ hybridization [13] and physical mapping with YACs [26] assigned the XLP gene to a about a 2-Mb interval between DXS6 and DXS100, around DXS739, which is deleted in 3 XLP patients [15]. The identification of the new interstitial deletion in XLP-D, therefore, reduces considerably the candidate region for the XLP gene.
The XLP-D deletion which maps in the previously defined large candidate region is physically linked to DXS982 and is centromeric to DXS739. The restriction map of the region deleted in patient XLP-D will be instrumental in the positional cloning of the gene. Single-copy fragments from cosmids within the minimal XLP deletion contig were used as probes to search for small genomic rearrangements in our panel of XLP patients. No new abnormal fragments were seen by Southern blotting, indicating that the XLP-D deletion defines the smallest genomic interval within which a fragment of the XLP gene must reside.
The new markers that we have assigned to the XLP locus, DF83 and PF12, are potentially interesting, since they were derived from expressed sequences [19]. However, they do not contain an open reading frame, nor is their expression detectable other than in minimal amounts by reverse-transcriptase-PCR in all the tissues we have screened. Screening of cDNA libraries with these markers has not identified a further transcribed sequence. Thus, the relationship, if any, of these transcribed sequences to the XLP gene remains unknown.
In the cosmid contig presented here, no clear CpG island was found (data not shown). This finding is in agreement with the low density of genes in this portion of the X chromosome and also with the possibility that the XLP gene is spread over a large genomic region.
The search for conserved and transcribed sequences in the contig by zoo blot and Northern blot analysis has so far been unsuccessful. As it is not known which cell types are involved in XLP, it is important to study the expression of candidate sequences in a wide range of tissues. Cells of the immune system, like B and T cells and macrophages, are good candidates for tissue-specific expression of the XLP gene.
Finally, the characterization of the function of the XLP gene should help in understanding the fundamental aspects of EBV biology and the role of the XLP gene in controlling this viral infection.
References
Purtilo DT, Yang JP, Cassel CK, Harper P, Stephenson SR, Landin BH, Vawter GF: X-linked recessive progressive combined variable immunodeficiency (Duncan’s disease). Lancet 1975;i:935–940
On-Line Mendelian Inheritance in Man: OMIM (TM), Immunodeficiency X-Linked Progressive Combined Variable [Lymphoproliferative Disease, X-linked, XLP], MIM No. 312060. Baltimore, Johns Hopkins University, 1988.
Harrington DS, Weisenburger DD, Purtilo DT: Malignant lymphoma in the X-linked lymphoproliferative Syndrome. Cancer 1987;59:1419–1429
Purtilo DT, Sakamoto K, Bernabei V, Seeley J, Bechtold T, Rogers G, XLP Collaborators: Epstein-Barr virus induced in boys with the X-linked lymphoproliferative syndrome (XLP). Update on studies of the registry. Am J Med 1982;73:49–56
Grierson H, Skare J, Hawk J, Panza M, Purtilo D: Immunoglobulin class and subclass deficiencies prior to EBV infection in males with X-linked lymphoproliferative disease (XLP). Am J Hum Genet 1991;40:294–297
Miller G: Epstein-Barr virus: Biology, pathogenesis and medical aspects, 1921–1958; in Fields BN, Knipe DM (eds): Virology. New York, Raven Press, 1990.
Klein G: Epstein-Barr virus strategy in normal and neoplastic B cells. Cell 1994;77:791–793
Mandel JL, Willard HF, Nussbaum RL, Romeo G, Puck JM, Davies KE: Report of the Committee on the Genetic Constitution of the X Chromosome. Cytogenet Cell Genet 1989;51:384–437
Skare JC, Grierson HL, Sullivan JL, Nussbaum RL, Purtilo DT, Sylla BS, Lenoir GM, Reilly DS, White BN, Milunsky A: Linkage analysis of seven kindred with the X-linked lymphoproliferative syndrome (XLP) confirms that XLP locus in near DXS42 and DXS37. Hum Genet 1989;82:354–358
Sylla BS, Wang Q, Hayoz D, Lathrop GM, Lenoir GM: Multipoint linkage mapping of the Xq25–26 region in a family affected by the X-linked lymphoproliferative syndrome. Clin Genet 1989;36:459–462
Skare JC, Grierson H, Wyandt H, Sanger W, Milunsky J, Purtilo D, Sullivan J, Milunsky A: Genetic of the X-linked lymphoproliferative syndrome (abstract 630). Am J Hum Genet 1989;45:A161.
Purtilo DT, Grierson HL: Methods of detection of new families with X linked lymphoproliferative disease (XLP). Cancer Genet Cytogenet 1991;51:143–153
Wu BL, Milunsky A, Nelson D, Schmeckpeper BJ, Porta G, Schlessinger D, Skare J: High-resolution mapping of probes near the X-linked lymphoproliferative disease (XLP) locus. Genomics 1993;17:163–170
Wyandt H, Grierson H, Sanger W, Skare J, Milunsky A, Purtilo D: Chromosome deletion of Xq25 in an individual with X-linked lymphoproliferative disease. Am J Med Genet 1989;33:426–430
Skare J, Wu BL, Madan S, Pulijaal V, Purtilo D, Haber D, Nelson D, Sylla B, Grierson H, Nitowsky H, Glaser J, Wissink J, White B, Holden J, Housman D, Lenoir G, Wyandt H: Characterization of three overlapping deletions causing X-linked lymphoproliferative disease. Genomics 1993;16:254–255
Sylla BS, Lamartine J, Wang Q, Pauly S, Lenoir GM, Haber D, Skare J, Nelson DL, Romeo G, Tsuji O: Characterization of an XLP patient carrying an interstitial deletion at Xq25. Cytogenet Cell Genet 1993;64:189.
Purtilo DT, Sakamoto K, Barnabei V, Seeley J, Bechtold T, Rogers G, Yetz J, Harada S: Epstein-Barr virus-induced diseases in boys with the X-linked lymphoproliferative syndrome (XLP): Update on studies of the registry. Am J Med 1982;73:49–56
Aldrige J, Kunkel L, Bruns G, Tantravahi U, Lalande M, Brewster T, Moreau E, Wilson M, Bromley W, Roderick T, Latt SA: A strategy to reveal high frequency RFLPs along the human X chromosome. Am J Hum Genet 1984;36:546–564
Yokoi H, Hadano S, Kogi M, Kang X, Wakasa K, Ikeda J: Isolation of expressed sequences encoded by the human Xq terminal portion using microclone probes generated by laser microdissection. Genomics 1994;20:404–411
Mulligan L, Phillips M, Foster-Gibson C, Beckett J, Partington M, Simpson N, Holden JJ, White BN: Genetic mapping of DNA segments relative to the locus for the fragile-X syndrome at Xq27-3. Am J Hum Genet 1985;37:463–472
Green ED, Olson MV: Systematic screening of yeast artificial chromosome libraries by use of the polymerase chain reaction. Proc Natl Acad Sci USA 1990;87:1213–1217
Nizetic D, Zehetner G, Monaco AP, Gellen L, Young BD, Lehrach H: Construction, arraying, and high density screening of large inserts libraries of human chromosome X and 21: Their potential use as reference libraries. Proc Natl Acad Sci USA 1991;88:3233–3237
Sambrook J, Fritsch EF, Maniatis T: Molecular Cloning. Cold Spring Harbor, Cold Spring Harbor Laboratory, 1989.
Blonden LAJ, den Dunnen JT, van Paassen HMB, Wapenaar MC, Grootscholten PM, Ginjaar HB, Bakker E, Pearson PL, van Ommen GJB: High resolution deletion breakpoint mapping in the DMD gene by whole cosmid hybridization. Nucleic Acids Res 1989;17:5611–5621
Kaneko K, Myatake T, Mijeon BR, Warren ST, Tsuji S: Isolation of cosmid clones containing CpG island at Xq24-qter region. Am J Hum Genet 1992;49:2069.
Wang Q, Ishikawa-Brush Y, Monaco AP, Nelson DL, Caskey CT, Pauly SP, Lenoir GM, Sylla BS: Physical mapping of Xq24–q25 around loci closely linked to the X-linked lymphoproliferative syndrome locus: An overlapping YAC map and linkage between DXS12, DXS42 and DXS37. Eur J Hum Genet 1993;1:64–71
Acknowledgements
We thank Dr. A. Harris for providing cell lines. We thank also Dr. D. LePallier for making accessible to us the CEPH mega-YAC library. We thank the ICRF for providing us filters for X chromosome cosmid library. The technical help of S. Pauly, M.-F. Lavoué, and C. Bonnardel is gratefully acknowledged. We also thank Dr. S. Jones for editing the manuscript. This work was supported in part by the Ligue Nationale Française contre le Cancer, Comité Départemental du Rhône. F. Heitzmann was supported by a fellowship from the Ligue Nationale Française contre le Cancer, Comité Départemental de l’Yonne. D. Haber and K. Nichols were supported by grants from NIH (CA4498 for D.H. and NIH CA4509 for K.N.). M. Krainer was supported by a grant from Erwin-Schrödinger-Stipendium, Fonds zur Förderung der Wissenschaftlichen Forschung. G. Porta was funded by the Associazione Italiana Ricerca sul Cancro and the Telethon e 294.
Author information
Authors and Affiliations
Corresponding author
Additional information
The first two authors contributed equally to this work.
Rights and permissions
About this article
Cite this article
Lamartine, J., Nichols, K.E., Yin, L. et al. Physical Map and Cosmid Contig Encompassing a New Interstitial Deletion of the X-Linked Lymphoproliferative Syndrome Region. Eur J Hum Genet 4, 342–351 (1996). https://doi.org/10.1159/000472230
Received:
Revised:
Accepted:
Issue Date:
DOI: https://doi.org/10.1159/000472230
Key Words
This article is cited by
-
Why commonplace encounters turn to fatal attraction
Nature Genetics (1998)
-
The X-linked lymphoproliferative-disease gene product SAP regulates signals induced through the co-receptor SLAM
Nature (1998)
-
Host response to EBV infection in X-linked lymphoproliferative disease results from mutations in an SH2-domain encoding gene
Nature Genetics (1998)