Abstract
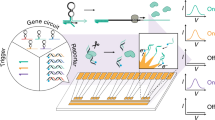
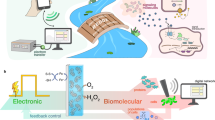
Although there has been considerable progress in the development of engineering principles for synthetic biology, a substantial challenge is the construction of robust circuits in a noisy cellular environment. Such an environment leads to considerable intercellular variability in circuit behaviour, which can hinder functionality at the colony level. Here we engineer the synchronization of thousands of oscillating colony ‘biopixels’ over centimetre-length scales through the use of synergistic intercellular coupling involving quorum sensing within a colony and gas-phase redox signalling between colonies. We use this platform to construct a liquid crystal display (LCD)-like macroscopic clock that can be used to sense arsenic via modulation of the oscillatory period. Given the repertoire of sensing capabilities of bacteria such as Escherichia coli, the ability to coordinate their behaviour over large length scales sets the stage for the construction of low cost genetic biosensors that are capable of detecting heavy metals and pathogens in the field.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Gibson, D. et al. Creation of a bacterial cell controlled by a chemically synthesized genome. Science 329, 52–56 (2010)
Hasty, J., McMillen, D. & Collins, J. J. Engineered gene circuits. Nature 420, 224–230 (2002)
Sprinzak, D. & Elowitz, M. B. Reconstruction of genetic circuits. Nature 438, 443–448 (2005)
Endy, D. Foundations for engineering biology. Nature 438, 449–453 (2005)
Ellis, T., Wang, X. & Collins, J. Diversity-based, model-guided construction of synthetic gene networks with predicted functions. Nature Biotechnol. 27, 465–471 (2009)
Kobayashi, H. et al. Programmable cells: interfacing natural and engineered gene networks. Proc. Natl Acad. Sci. USA 101, 8414–8419 (2004)
You, L., Cox, R. S., III, Weiss, R. & Arnold, F. H. Programmed population control by cell–cell communication and regulated killing. Nature 428, 868–871 (2004)
Basu, S., Gerchman, Y., Collins, C. H., Arnold, F. H. & Weiss, R. A synthetic multicellular system for programmed pattern formation. Nature 434, 1130–1134 (2005)
Mukherji, S. & Van Oudenaarden, A. Synthetic biology: understanding biological design from synthetic circuits. Nature Rev. Genet. 10, 859–871 (2009)
Grilly, C., Stricker, J., Pang, W., Bennett, M. & Hasty, J. A synthetic gene network for tuning protein degradation in Saccharomyces cerevisiae . Mol. Syst. Biol. 3, 127 (2007)
Gardner, T. S., Cantor, C. R. & Collins, J. J. Construction of a genetic toggle switch in Escherichia coli . Nature 403, 339–342 (2000)
Elowitz, M. B. & Leibler, S. A synthetic oscillatory network of transcriptional regulators. Nature 403, 335–338 (2000)
Lu, T. & Collins, J. Dispersing biofilms with engineered enzymatic bacteriophage. Proc. Natl Acad. Sci. USA 104, 11197–11202 (2007)
Friedland, A. et al. Synthetic gene networks that count. Science 324, 1199–1202 (2009)
Danino, T., Mondragon-Palomino, O., Tsimring, L. & Hasty, J. A synchronized quorum of genetic clocks. Nature 463, 326–330 (2010)
Tamsir, A., Tabor, J. & Voigt, C. Robust multicellular computing using genetically encoded NOR gates and chemical ‘wires’. Nature 469, 212–215 (2011)
Tabor, J. et al. A synthetic genetic edge detection program. Cell 137, 1272–1281 (2009)
Stricker, J. et al. A fast, robust and tunable synthetic gene oscillator. Nature 456, 516–519 (2008)
Mondragon-Palomino, O., Danino, T., Selimkhanov, J., Tsimring, L. & Hasty, J. Entrainment of a population of synthetic genetic oscillators. Science 333, 1315–1319 (2011)
Tigges, M., Marquez-Lago, T., Stelling, J. & Fussenegger, M. A tunable synthetic mammalian oscillator. Nature 457, 309–312 (2009)
Westinghouse, G. System of electrical distribution. US patent 373. 035 (1887)
Lewandowski, W., Azoubib, J. & Klepczynski, W. GPS: primary tool for time transfer. Proc. IEEE 87, 163–172 (1999)
Vladimirov, A., Kozyreff, G. & Mandel, P. Synchronization of weakly stable oscillators and semiconductor laser arrays. Europhys. Lett. 61, 613 (2003)
Gast, T. Sensors with oscillating elements. J. Phys. E: Sci. Instrum. 18, 783 (1985)
Ozbudak, E. M., Thattai, M., Kurtser, I., Grossman, A. D. & van Oudenaarden, A. Regulation of noise in the expression of a single gene. Nature Genet. 31, 69–73 (2002)
Elowitz, M. B., Levine, A. J., Siggia, E. D. & Swain, P. S. Stochastic gene expression in a single cell. Science 297, 1183–1186 (2002)
Golding, I., Paulsson, J., Zawilski, S. & Cox, E. Real-time kinetics of gene activity in individual bacteria. Cell 123, 1025–1036 (2005)
Blake, W. et al. Phenotypic consequences of promoter-mediated transcriptional noise. Mol. Cell 24, 853–865 (2006)
Austin, D. et al. Gene network shaping of inherent noise spectra. Nature 439, 608–611 (2006)
Waters, C. & Bassler, B. Quorum sensing: cell-to-cell communication in bacteria. Annu. Rev. Cell Dev. Biol. 21, 319–346 (2005)
Ferry, M., Razinkov, I. & Hasty, J. Microfluidics for synthetic biology from design to execution. Methods Enzymol. 497, 295 (2011)
Messner, K. & Imlay, J. The identification of primary sites of superoxide and hydrogen peroxide formation in the aerobic respiratory chain and sulfite reductase complex of Escherichia coli . J. Biol. Chem. 274, 10119–10128 (1999)
Bose, J. L. et al. Bioluminescence in Vibrio fischeri is controlled by the redox-responsive regulator arca. Mol. Microbiol. 65, 538–553 (2007)
Georgellis, D., Kwon, O. & Lin, E. Quinones as the redox signal for the arc two-component system of bacteria. Science 292, 2314–2316 (2001)
Seaver, L. & Imlay, J. Hydrogen peroxide fluxes and compartmentalization inside growing Escherichia coli . J. Bacteriol. 183, 7182–7189 (2001)
Fridovich, I. The biology of oxygen radicals. Science 201, 875–880 (1978)
McCord, J. & Fridovich, I. Superoxide dismutase. J. Biol. Chem. 244, 6049–6055 (1969)
Berg, J., Tymoczko, J. L. & Stryer, L. Biochemistry (W.H. Freeman, 2006)
Remington, S. Fluorescent proteins: maturation, photochemistry and photophysics. Curr. Opin. Struct. Biol. 16, 714–721 (2006)
Kelner, M., Bagnell, R. & Welch, K. Thioureas react with superoxide radicals to yield a sulfhydryl compound. explanation for protective effect against paraquat. J. Biol. Chem. 265, 1306–1311 (1990)
Touati, D., Jacques, M., Tardat, B., Bouchard, L. & Despied, S. Lethal oxidative damage and mutagenesis are generated by iron in delta fur mutants of Escherichia coli: protective role of superoxide dismutase. J. Bacteriol. 177, 2305–2314 (1995)
Kohanski, M. A., DePristo, M. A. & Collins, J. J. Sublethal antibiotic treatment leads to multidrug resistance via radical-induced mutagenesis. Mol. Cell 37, 311–320 (2010)
Nordstrom, D. Worldwide occurrences of arsenic in ground water. Science 296, 2143 (2002)
van der Meer, J. & Belkin, S. Where microbiology meets microengineering: design and applications of reporter bacteria. Nature Rev. Microbiol. 8, 511–522 (2010)
Daunert, S. et al. Genetically engineered whole-cell sensing systems: coupling biological recognition with reporter genes. Chem. Rev. 100, 2705–2738 (2000)
Leveau, J. & Lindow, S. Bioreporters in microbial ecology. Curr. Opin. Microbiol. 5, 259–265 (2002)
Mather, W., Bennett, M., Hasty, J. & Tsimring, L. Delay-induced degrade-and-fire oscillations in small genetic circuits. Phys. Rev. Lett. 102, 068105 (2009)
Quan, J. & Tian, J. Circular polymerase extension cloning of complex gene libraries and pathways. PLoS ONE 4, e6441 (2009)
Stocker, J. et al. Development of a set of simple bacterial biosensors for quantitative and rapid measurements of arsenite and arsenate in potable water. Environ. Sci. Technol. 37, 4743–4750 (2003)
Keiler, K., Waller, P. & Sauer, R. Role of a peptide tagging system in degradation of proteins synthesized from damaged messenger RNA. Science 271, 990–993 (1996)
Acknowledgements
This work was supported by the National Institutes of Health and General Medicine (R01GM69811), the San Diego Center for Systems Biology (P50GM085764), the DoD NDSEG (A.P.) and NSF Graduate Research (P.S.) Fellowship Programs. L.T. was supported, in part, by the Office of Naval Research (MURI N00014-07-0741). We would like to thank J. Imlay, S. Ali and J. Collins for helpful discussions and J. Hickman, B. Taylor and K. Lomax for their help with illustrations.
Author information
Authors and Affiliations
Contributions
All authors contributed extensively to the work presented in this paper. A.P. and P.S. are equally contributing first authors.
Corresponding author
Supplementary information
Supplementary Information
The file contains Supplementary Text and Data, Supplementary Figures 1-12 with legends and additional references. (PDF 1384 kb)
Supplementary Movie 1
The movie shows timelapse fluorescence microscopy of a 200 trap sensor array displaying NDH-2 engineered synchronization. An EMCCD camera was used to keep exposure times extremely low (4X magnification, 20ms, 95% attenuation) to ensure no fluorescence interaction, hence the appearance of lower signal. (MOV 11321 kb)
Supplementary Movie 2
The movie shows timelapse fluorescence microscopy of the 500 trap biosensor array showing the onset of synchronization from disparate initial conditions using period modulator circuit. Flashes indicate changes in arsenite concentration which result in changes in the oscillatory period. (MOV 9624 kb)
Supplementary Movie 3
The movie shows timelapse fluorescence microscopy of a sensor array containing thresholding circuit. Red color indicates addition of 0.25 mM arsenite that initiates oscillations in blue. (MOV 1671 kb)
Supplementary Movie 4
The movie shows timelapse fluorescence microscopy of a modified 500 trap sensor array in which traps are farther apart. This increased separation results in anti phase oscillations, where a biopixel and its nearest neighbors alternate bursts. (MOV 2678 kb)
Supplementary Movie 5
The movie shows timelapse fluorescence microscopy of the 12,000 trap scaled up array showing oscillation and synchronization maintained over a maximum distance of 27 mm. (MOV 7711 kb)
Supplementary Movie 6
The movie shows Real time microscopy depicting the loading of our microfluidic device. Cells flow in from the cell port and fill the trapping regions. (MOV 5442 kb)
Supplementary Movie 7
The movie shows Timelapse fluorescence microscopy of a modified 500 trap sensor array in which traps of 2 sizes are present. This results in 2:1 resonant oscillations where larger traps oscillate at twice the frequency of smaller traps. (MOV 2456 kb)
Supplementary Movie 8
The movie shows timelapse fluorescence microscopy of a 500 trap sensor array showing unsynchronized oscillations when NDH-2 is not present and high-intensity fluorescence bursts are not used. (MOV 10904 kb)
Rights and permissions
About this article
Cite this article
Prindle, A., Samayoa, P., Razinkov, I. et al. A sensing array of radically coupled genetic ‘biopixels’. Nature 481, 39–44 (2012). https://doi.org/10.1038/nature10722
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature10722
This article is cited by
-
Engineered bacterial swarm patterns as spatial records of environmental inputs
Nature Chemical Biology (2023)
-
Competition and evolutionary selection among core regulatory motifs in gene expression control
Nature Communications (2023)
-
The Application Potential of Synthetic Biology in Microbial Communication
Current Clinical Microbiology Reports (2023)
-
Synthetic Micrographs of Bacteria (SyMBac) allows accurate segmentation of bacterial cells using deep neural networks
BMC Biology (2022)
-
Probing patterning in microbial consortia with a cellular automaton for spatial organisation
Scientific Reports (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.