Systems biologists combine biochemical mapping, multi-omics data and mathematical modelling to reveal the network designs underpinning health and disease.Credit: Tek Image/ Getty Images
The medicines we need to treat complex diseases may already exist; we just haven’t found the right targets yet. That’s the view of Matteo Barberis, professor of systems biology at the University of Surrey, UK, who integrates biochemical maps with multi-omics data and mathematical modelling to uncover the network designs underpinning healthy and disease states.
The goal is not just to find one therapeutic target, but to understand the design principles of cellular organization and reveal key players and compensatory processes that are deployed by the cell to relieve the disease state. Barberis’s approach is partly inspired by his mother’s experience of cancer treatment. “She receives four different drugs, each specific to a separate target,” says Barberis. “But nobody knows whether these targets are actually interacting with one another. If we knew this, we could potentially disrupt the network with fewer drugs.”
From multi-scale maps to mathematical models
Barberis takes a multi-pronged approach to systems biology, starting with layering multi-omics data on to a biochemical map. Next, insights from the experimental literature are included to build a mechanistic map of disease, creating multi-scale molecular maps that inform different types of mathematical model.
“Traditional bioinformatics approaches can tell us which molecules are up- or down-regulated in disease versus health, but they cannot tell us about the connections between those molecules,” says Barberis. “We biochemically analyse multi-omics datasets, for instance checking all possible metabolic reactions, and place information about differentially expressed pathways into the broader context of the known biochemical map.”
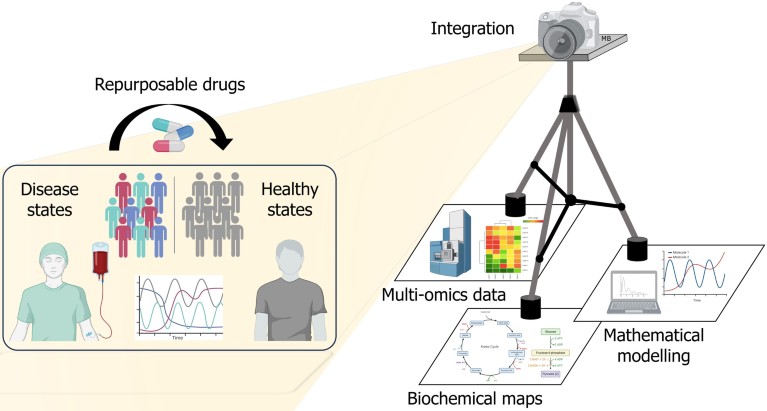
A 'tripod' of information – multi-omics data, biochemical maps and mathematical modelling – is integrated in a systems biology 'camera' to enhance researchers' view of how new molecular targets of disease networks can be identified for treatment, possibly repurposing existing drugs against different disease states.Credit: Matteo Barberis/ University of Surrey
The next challenge is to envision how these differentially expressed targets link together. “We use metabolomics data and metabolic information1 from the mechanistic map to reconstruct and design minimal yet functional cells through engineering-biology principles, using just enough metabolites and connectors for the metabolic modelling of pathways to run as it would in the disease or healthy states,” explains Barberis. “This allows us to propose mechanisms underlying a disease that are testable experimentally.”
Matteo Barberis, expert in biochemistry of cell cycle control, metabolism and the immune system, is a professor of systems biology at the University of Surrey in Guildford, United Kingdom. He is vice-chairman of the International Society of Systems Biology (ISSB) and editor-in-chief of Current Opinion in Systems Biology (Elsevier). Credit: University of Surrey
Clinical implications
Insights generated using Barberis’s strategy have already revealed key network motifs in the cell cycle3, and targetable genes in autoimmune disorders such as rheumatoid arthritis4. Targets for immune disorders and cancer are being identified by the Barberis lab. These data can be used to guide biochemical functional testing, tailored drug development and vaccine design, where the temporal dynamics of immune networks could be used to optimize response.
Drug discovery tends to focus on single targets or phenotypes, without full knowledge of the downstream consequences. Barberis believes systems biology offers a better way — data-informed targeting of the molecules and motifs that disease networks rely on. “If we understand the designs and their rewiring, we can find drugs to move the cell from one state to another,” he argues. “And because we are targeting networks that may have been explored for other diseases, many of these drugs may already exist and could be repurposed.”


